

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.12
(125 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
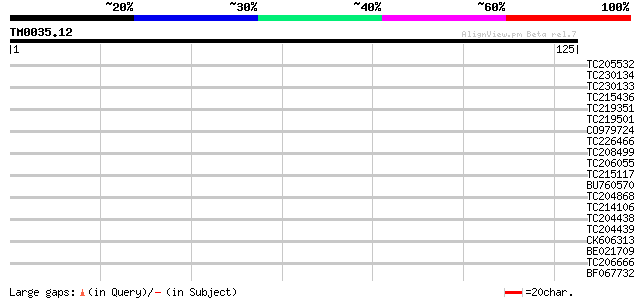
Score E
Sequences producing significant alignments: (bits) Value
TC205532 weakly similar to UP|O04559 (O04559) T7N9.16, partial (... 33 0.033
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 30 0.28
TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 28 0.82
TC215436 similar to UP|Q9LZY8 (Q9LZY8) Transketolase-like protei... 28 1.1
TC219351 27 1.8
TC219501 weakly similar to UP|Q9XE35 (Q9XE35) ESTD24835(R2634) c... 27 1.8
CO979724 27 1.8
TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {... 27 2.4
TC208499 similar to GB|AAP68331.1|31711950|BT008892 At5g14170 {A... 27 2.4
TC206055 similar to PRF|1604369A.0|226743|1604369A sulfated surf... 27 3.1
TC215117 homologue to PIR|T02313|T02313 endoplasmic reticulum in... 27 3.1
BU760570 similar to GP|23170343|gb| CG31556-PA {Drosophila melan... 27 3.1
TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, par... 27 3.1
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 26 4.1
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 26 4.1
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 26 4.1
CK606313 26 5.3
BE021709 similar to SP|Q43064|PYB3 Aspartate carbamoyltransferas... 26 5.3
TC206666 26 5.3
BF067732 similar to GP|2707600|gb|A triple LIM domain protein {H... 25 9.1
>TC205532 weakly similar to UP|O04559 (O04559) T7N9.16, partial (54%)
Length = 1350
Score = 33.1 bits (74), Expect = 0.033
Identities = 31/98 (31%), Positives = 46/98 (46%), Gaps = 5/98 (5%)
Frame = +1
Query: 10 HSHRRSFFTH--INWAPD---QHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEK 64
H S TH I WA + + P ++++F S F S D AS EE+
Sbjct: 409 HDDPASSVTHTWILWASEPVQEFPEKMVSFAESDFSSLDSEDELLAS----------EEE 558
Query: 65 ASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAAVR 102
+S +K+ +KT ++T AT +RSG +F AVR
Sbjct: 559 SSRSHKSMLKT--TSSNTSGATSIRSGMELFHRAKAVR 666
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 30.0 bits (66), Expect = 0.28
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = -1
Query: 7 QHHHSHRRSFFTHINWAPDQHPHEVL 32
+HHH HRR F +W D H +++L
Sbjct: 452 RHHHPHRRHRFHLHHWPRDHHQNQLL 375
>TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 850
Score = 28.5 bits (62), Expect = 0.82
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = -2
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVL 32
H+P +HHHSH W D H +++L
Sbjct: 447 HHPHRHHHSHLH------RWPQDHHQNQLL 376
>TC215436 similar to UP|Q9LZY8 (Q9LZY8) Transketolase-like protein, partial
(65%)
Length = 1625
Score = 28.1 bits (61), Expect = 1.1
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +3
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFK 40
H+P HHS S H P H VLA P+LF+
Sbjct: 129 HSPS--HHSLASSHILHAKQQPRPHVEGVLALPPALFE 236
>TC219351
Length = 758
Score = 27.3 bits (59), Expect = 1.8
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = -1
Query: 56 SLNKTQEEKASIWNKTQIKTPLNKASTFEAT 86
SLNKT ++ A IWN I +NK S+ +A+
Sbjct: 641 SLNKTLKK*ADIWNCKPIHIWINKLSSKDAS 549
>TC219501 weakly similar to UP|Q9XE35 (Q9XE35) ESTD24835(R2634) corresponds
to a region of the predicted gene, partial (16%)
Length = 544
Score = 27.3 bits (59), Expect = 1.8
Identities = 17/53 (32%), Positives = 26/53 (48%), Gaps = 3/53 (5%)
Frame = +2
Query: 5 PGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLA---RRKRASDGD 54
P +HH + + F H QH H ++A L ++ LA RR++ DGD
Sbjct: 188 PFRHHGARQDRFRRH------QHGHVIVAKTRRLLRAESLARLSRRRQVIDGD 328
>CO979724
Length = 788
Score = 27.3 bits (59), Expect = 1.8
Identities = 13/37 (35%), Positives = 18/37 (48%), Gaps = 1/37 (2%)
Frame = +3
Query: 3 HNPGQHHHSHRRS-FFTHINWAPDQHPHEVLAFMPSL 38
+NP HH H RS FF H + + + +PSL
Sbjct: 513 YNPNTEHHYHSRSYFFKHNQFLTKHNTSSTHSIIPSL 623
>TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {Arabidopsis
thaliana;} , partial (22%)
Length = 978
Score = 26.9 bits (58), Expect = 2.4
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +2
Query: 4 NPGQHHHSHRRSFFTHINWAPDQHPH 29
+P HHH+H++ H++ P HPH
Sbjct: 278 HPKLHHHNHQK---YHLSQLPHCHPH 346
>TC208499 similar to GB|AAP68331.1|31711950|BT008892 At5g14170 {Arabidopsis
thaliana;} , partial (18%)
Length = 703
Score = 26.9 bits (58), Expect = 2.4
Identities = 16/67 (23%), Positives = 33/67 (48%)
Frame = +1
Query: 8 HHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASI 67
H H RR+FF + +P + F+ +L +S +R + G+ S N +E ++
Sbjct: 61 HEHRRRRAFFLGFSQSP-------VEFINALIESQ--SRDLKLVSGEPSRNAEKERRSDF 213
Query: 68 WNKTQIK 74
+N+ ++
Sbjct: 214 FNQPWVE 234
>TC206055 similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (6%)
Length = 824
Score = 26.6 bits (57), Expect = 3.1
Identities = 11/26 (42%), Positives = 12/26 (45%)
Frame = +1
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHP 28
H HHH R S TH N +HP
Sbjct: 73 HQAQTHHHHRRHSHTTHNNLHRRRHP 150
>TC215117 homologue to PIR|T02313|T02313 endoplasmic reticulum insertion
protein F13P17.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (13%)
Length = 424
Score = 26.6 bits (57), Expect = 3.1
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +3
Query: 10 HSHRRSFFTHINWAPD 25
HSH RS + H+ W PD
Sbjct: 9 HSHCRSIWRHMYWCPD 56
>BU760570 similar to GP|23170343|gb| CG31556-PA {Drosophila melanogaster},
partial (4%)
Length = 373
Score = 26.6 bits (57), Expect = 3.1
Identities = 10/26 (38%), Positives = 12/26 (45%)
Frame = +3
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHP 28
H+P HHSH + W P HP
Sbjct: 303 HHP---HHSHNDKNLIFLEWCPQTHP 371
>TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, partial (91%)
Length = 865
Score = 26.6 bits (57), Expect = 3.1
Identities = 12/31 (38%), Positives = 14/31 (44%), Gaps = 1/31 (3%)
Frame = +3
Query: 3 HNPGQH-HHSHRRSFFTHINWAPDQHPHEVL 32
H+P H HH H R H+ HPH L
Sbjct: 258 HHPHPHPHHPHPRLVVLHLLLRSPHHPHHQL 350
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 26.2 bits (56), Expect = 4.1
Identities = 13/32 (40%), Positives = 14/32 (43%)
Frame = +2
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVLAF 34
H HHH HRR F P H H +L F
Sbjct: 242 HIRPHHHHLHRRHLF-----IPIYHHHHLLQF 322
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 26.2 bits (56), Expect = 4.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Frame = -3
Query: 12 HRRSFFTHINWAPDQHPHEVLAFMPSLFKSND 43
H++ FF H+++ P+QH + P+L +ND
Sbjct: 4324 HQKCFFCHLHFQPNQHHNT----QPALNLNND 4241
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 26.2 bits (56), Expect = 4.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Frame = -3
Query: 12 HRRSFFTHINWAPDQHPHEVLAFMPSLFKSND 43
H++ FF H+++ P+QH + P+L +ND
Sbjct: 4321 HQKCFFCHLHFQPNQHHNT----QPALDLNND 4238
>CK606313
Length = 617
Score = 25.8 bits (55), Expect = 5.3
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = -3
Query: 56 SLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSG 98
S+ +++ EKAS+W+ STF +L+ CI R G
Sbjct: 312 SVKESRHEKASLWSSWSSPCARRVTSTF---RLQKKCCIDRGG 193
>BE021709 similar to SP|Q43064|PYB3 Aspartate carbamoyltransferase 3
chloroplast precursor (EC 2.1.3.2) (Aspartate
transcarbamylase 3), partial (25%)
Length = 475
Score = 25.8 bits (55), Expect = 5.3
Identities = 14/35 (40%), Positives = 17/35 (48%), Gaps = 1/35 (2%)
Frame = +2
Query: 68 WNKTQIKTPLNKASTFEATKL-RSGTCIFRSGAAV 101
WN + KTP + FE KL G I R AA+
Sbjct: 101 WNHLKSKTPFGHSGVFEKCKLSHKGDEILRRAAAL 205
>TC206666
Length = 1493
Score = 25.8 bits (55), Expect = 5.3
Identities = 11/44 (25%), Positives = 20/44 (45%)
Frame = +3
Query: 8 HHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRAS 51
HH F + +++APD H + +F P + +L R +
Sbjct: 558 HHRREGYPFGSLVDFAPDSTGHPIFSFSPLAIHTRNLLADPRCT 689
>BF067732 similar to GP|2707600|gb|A triple LIM domain protein {Homo
sapiens}, partial (6%)
Length = 284
Score = 25.0 bits (53), Expect = 9.1
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -3
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVLAF 34
HN HH SHR H++ HPH +L +
Sbjct: 114 HN--HHHXSHRHDLHHHLH----LHPHPLLQY 37
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,876,284
Number of Sequences: 63676
Number of extensions: 78200
Number of successful extensions: 879
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 824
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 865
length of query: 125
length of database: 12,639,632
effective HSP length: 86
effective length of query: 39
effective length of database: 7,163,496
effective search space: 279376344
effective search space used: 279376344
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0035.12


