

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0034a.6
(1532 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
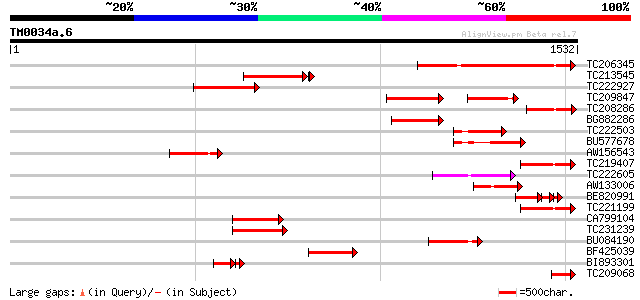
Score E
Sequences producing significant alignments: (bits) Value
TC206345 similar to UP|O24518 (O24518) Unconventional myosin, pa... 424 e-118
TC213545 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, part... 288 7e-78
TC222927 homologue to UP|Q9M5A6 (Q9M5A6) Unconventional myosin X... 261 2e-69
TC209847 257 3e-68
TC208286 weakly similar to UP|Q39160 (Q39160) Myosin, partial (7%) 246 6e-65
BG882286 216 6e-56
TC222503 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, part... 209 6e-54
BU577678 195 1e-49
AW156543 similar to PIR|T14275|T1 myosin-like protein my1 - comm... 164 4e-40
TC219407 similar to UP|O24517 (O24517) Unconventional myosin, pa... 155 1e-37
TC222605 similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain class ... 149 9e-36
AW133006 similar to GP|29170491|d myosin XI {Nicotiana tabacum},... 145 1e-34
BE820991 138 2e-32
TC221199 weakly similar to UP|Q9XEI4 (Q9XEI4) Unconventional myo... 134 4e-31
CA799104 125 2e-28
TC231239 similar to GB|BAB03161.1|11994771|AP002050 myosin-like ... 124 3e-28
BU084190 similar to GP|29170491|d myosin XI {Nicotiana tabacum},... 120 3e-27
BF425039 similar to PIR|T14279|T1 myosin-like protein my5 - comm... 114 4e-25
BI893301 similar to GP|7243765|gb| unconventional myosin XI {Val... 82 5e-24
TC209068 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, part... 103 6e-22
>TC206345 similar to UP|O24518 (O24518) Unconventional myosin, partial (27%)
Length = 1867
Score = 424 bits (1091), Expect = e-118
Identities = 221/433 (51%), Positives = 296/433 (68%), Gaps = 7/433 (1%)
Frame = +1
Query: 1102 SDSRRSKLTAERHQDNCEFLSRCIKENLG-FKNGKPLAAPIIYKCLLHWHAFESERTAIF 1160
S+ + K E+ Q+N + L +CI + G F P+AA +IYKCLLHW +FE ERT++F
Sbjct: 16 SEGKPQKSLNEKQQENQDLLIKCITQXFGIFXGAXPVAACVIYKCLLHWRSFEVERTSVF 195
Query: 1161 DYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGSAGLASR 1220
D II+ I ++A+D+ DVL YWLSNTS LL LLQR L+++G + T QR R
Sbjct: 196 DRIIQTIASAVEAQDNTDVLAYWLSNTSTLLLLLQRTLKASGAASLTPQR--------RR 351
Query: 1221 TVHVSYLVSTSINHSCGPKSP-LKFIGYD-----DGVSHVEARYPAILFKQQLTACVEKI 1274
T S S P+S L F+ D + VEA+YPA+LFKQQLTA +EKI
Sbjct: 352 TASSSLFGRMSQGLRASPQSAGLSFLNGRGLNRLDDLRQVEAKYPALLFKQQLTAFLEKI 531
Query: 1275 FGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPSGGQWANIVNFLDSLM 1334
+G++RD+LKKE+SPLLGLCIQAP+ RQ K + + QQ W +IV L++ +
Sbjct: 532 YGMIRDNLKKEISPLLGLCIQAPRNSRQSLVKGRAQANAVAQQALIAHWQSIVKSLNNYL 711
Query: 1335 SKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMKSGLAELEKWIVN 1394
+ N+ P F +RK+ TQ+FSFIN+ LFNSLLLRRECC+FSNGEY+K+GLAELE+W +
Sbjct: 712 KIMKANYAPPFLVRKVFTQIFSFINVQLFNSLLLRRECCSFSNGEYVKTGLAELEQWCIE 891
Query: 1395 AKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDK 1454
A EEY G++W EL +IRQAVGFLVIHQK KKSL+EI ++LCP L+++Q+YRISTMYWDDK
Sbjct: 892 ATEEYTGSAWEELKHIRQAVGFLVIHQKPKKSLNEITKELCPVLSIQQLYRISTMYWDDK 1071
Query: 1455 YGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDL 1514
YGT SVS +V++ MR ++ S+D+ N S SFLLDDD SIPFS +DI ++ ++ ++D
Sbjct: 1072 YGTHSVSTDVITNMRAMM-SEDSNNAVSTSFLLDDDSSIPFSVDDISKSMQQVEVADVDP 1248
Query: 1515 PAFVPEYSCAQFL 1527
P + E S FL
Sbjct: 1249 PPLIRENSGFGFL 1287
>TC213545 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, partial (13%)
Length = 608
Score = 288 bits (738), Expect(2) = 7e-78
Identities = 148/173 (85%), Positives = 159/173 (91%)
Frame = +1
Query: 633 NSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYD 692
NSLNRPQ FEN SVIHQLRCGGVLEAVRISLAGYPTRR YSEFVDRFGLIA EFMDGSYD
Sbjct: 1 NSLNRPQKFENTSVIHQLRCGGVLEAVRISLAGYPTRRIYSEFVDRFGLIAPEFMDGSYD 180
Query: 693 DKAVAEKILQKLKLENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARR 752
DKAV KILQKLKLENFQLGRTKVFLRAGQI ILDSRRAEVLDNAA+CIQR+LRTFIARR
Sbjct: 181 DKAVTLKILQKLKLENFQLGRTKVFLRAGQICILDSRRAEVLDNAAKCIQRRLRTFIARR 360
Query: 753 VFIAVRAAAVCLQACCRGYIGQKMYASKRETAAAISVQKYIRMWLRRKTYMKL 805
FI+++AAA+ +QACCRG IG+K+YASKRETAAAIS+QKYIRM L R Y+KL
Sbjct: 361 DFISIQAAALSIQACCRGCIGRKIYASKRETAAAISIQKYIRMCLMRHAYVKL 519
Score = 22.7 bits (47), Expect(2) = 7e-78
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = +2
Query: 807 SSATIIQSCVRGFMTRQR 824
+SA I QS RGF TRQ+
Sbjct: 524 NSAIIXQSNGRGFTTRQK 577
>TC222927 homologue to UP|Q9M5A6 (Q9M5A6) Unconventional myosin XI, partial
(12%)
Length = 718
Score = 261 bits (667), Expect = 2e-69
Identities = 123/179 (68%), Positives = 151/179 (83%)
Frame = +1
Query: 496 KKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISHYAGKVTYH 555
+KP GI+ALLDEACMFP+STHETF+ KL+Q F +H R K K +++DF I HYAG VTY
Sbjct: 184 QKPGGIIALLDEACMFPRSTHETFAQKLYQTFKNHKRFSKPKLARSDFTICHYAGDVTYQ 363
Query: 556 TDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVASRFKQQLQAL 615
T+ FLDKN+DYVV EH LL +SKCPFVSGLFP PEESS+ S +FSS+ASRFKQQLQAL
Sbjct: 364 TELFLDKNKDYVVAEHQELLYASKCPFVSGLFPPSPEESSKQS-KFSSIASRFKQQLQAL 540
Query: 616 METLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYPTRRTYSE 674
+ETL++TEPHY+RCVKPN+L +P +FEN +V+ QLRCGGV+EA+RIS AGYPTR+T+ E
Sbjct: 541 LETLSATEPHYIRCVKPNNLLKPAIFENKNVLQQLRCGGVMEAIRISCAGYPTRKTFDE 717
>TC209847
Length = 1275
Score = 257 bits (656), Expect = 3e-68
Identities = 130/157 (82%), Positives = 141/157 (89%), Gaps = 1/157 (0%)
Frame = +2
Query: 1017 REFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPL-KSNRQGFAKSLSERYSNAV 1075
RE EQKCSQLEQNVK LEEK+LSLEDENHVLRQKALS PL KSNR FAKS+SE+YS+A+
Sbjct: 8 RESEQKCSQLEQNVKRLEEKLLSLEDENHVLRQKALSTPLLKSNRPSFAKSISEKYSSAI 187
Query: 1076 ASRTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGK 1135
ASRTERK IFESPTPTKLI FT G+SDSRRSKLTAER QDN EFLS+CIKENLGFKNGK
Sbjct: 188 ASRTERKTIFESPTPTKLIAPFTLGLSDSRRSKLTAERQQDNYEFLSKCIKENLGFKNGK 367
Query: 1136 PLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLK 1172
P+AA IIYKCLLHWH+FES+RT IFD II+G NEVLK
Sbjct: 368 PIAARIIYKCLLHWHSFESDRTTIFDSIIKGTNEVLK 478
Score = 185 bits (469), Expect = 2e-46
Identities = 97/138 (70%), Positives = 109/138 (78%), Gaps = 1/138 (0%)
Frame = +2
Query: 1237 GPKSPLKFIGYDDGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQA 1296
GPKSPLKFIGYDDGV+HVEA+YPAILFKQQLTACVEKIFGLLRD+LKK+LSPLLG CIQA
Sbjct: 479 GPKSPLKFIGYDDGVAHVEAKYPAILFKQQLTACVEKIFGLLRDNLKKDLSPLLGSCIQA 658
Query: 1297 PKTGR-QHGGKLSRSPSGLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVF 1355
PKTGR HGGK SR P G+PQQ S GQW+NIV FLDSLMSKL NHV S + +
Sbjct: 659 PKTGRGVHGGKSSRFPGGIPQQSSSGQWSNIVKFLDSLMSKLRQNHVCSVLFQ------W 820
Query: 1356 SFINITLFNSLLLRRECC 1373
S + ++L N L + CC
Sbjct: 821 SLL-LSLVN*LYILSICC 871
>TC208286 weakly similar to UP|Q39160 (Q39160) Myosin, partial (7%)
Length = 867
Score = 246 bits (628), Expect = 6e-65
Identities = 122/135 (90%), Positives = 128/135 (94%)
Frame = +3
Query: 1397 EEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDKYG 1456
EEYAGTSWHELNYIRQA+GFLVIHQKRKKSL+EIRQDLCP LTVRQIYRISTMYWDDKYG
Sbjct: 3 EEYAGTSWHELNYIRQAIGFLVIHQKRKKSLEEIRQDLCPVLTVRQIYRISTMYWDDKYG 182
Query: 1457 TQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPA 1516
TQSVSNEVVSEMREIV SKDNQN+TSNSFLLDDD+SIPFSAEDIDMAIPAID DEIDLP
Sbjct: 183 TQSVSNEVVSEMREIV-SKDNQNLTSNSFLLDDDLSIPFSAEDIDMAIPAIDVDEIDLPE 359
Query: 1517 FVPEYSCAQFLNPIQ 1531
F+ EYSCAQFL+ Q
Sbjct: 360 FMSEYSCAQFLSSHQ 404
>BG882286
Length = 419
Score = 216 bits (550), Expect = 6e-56
Identities = 108/139 (77%), Positives = 120/139 (85%), Gaps = 1/139 (0%)
Frame = +2
Query: 1033 LEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTK 1092
LE K+ SLEDENHVLRQKALS KSN +G KSLSE+YS+A+A RTE+KP FESPTPTK
Sbjct: 2 LEGKLSSLEDENHVLRQKALSVSPKSNHRGLTKSLSEKYSSAIAPRTEQKPTFESPTPTK 181
Query: 1093 LIPTFTPG-MSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHA 1151
LIP T G +SDS RSKLTA+RHQDN E LSRCIKE+LGFKNGKPLAA IIYKCL HWHA
Sbjct: 182 LIPHITRGGLSDSHRSKLTADRHQDNYELLSRCIKEDLGFKNGKPLAASIIYKCLHHWHA 361
Query: 1152 FESERTAIFDYIIEGINEV 1170
FESERTAIFDYI++GIN+V
Sbjct: 362 FESERTAIFDYIVDGINDV 418
>TC222503 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, partial (5%)
Length = 541
Score = 209 bits (533), Expect = 6e-54
Identities = 109/144 (75%), Positives = 114/144 (78%), Gaps = 1/144 (0%)
Frame = +3
Query: 1200 SNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYP 1259
SNGFLTTT QRY GS+ L SR H GPKSPLKFIGYDDGV HVEARYP
Sbjct: 3 SNGFLTTTAQRYPGSSALTSRAGH-------------GPKSPLKFIGYDDGVLHVEARYP 143
Query: 1260 AILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGR-QHGGKLSRSPSGLPQQP 1318
AILFKQQLTACVEKIFGLLRD+LKK+LSPLLG CIQAPKTGR HGGK SRSP G+PQQ
Sbjct: 144 AILFKQQLTACVEKIFGLLRDNLKKDLSPLLGSCIQAPKTGRGLHGGKSSRSPGGIPQQS 323
Query: 1319 SGGQWANIVNFLDSLMSKLHGNHV 1342
S GQW+NIV FLDSLMSKL NHV
Sbjct: 324 SSGQWSNIVKFLDSLMSKLRQNHV 395
>BU577678
Length = 423
Score = 195 bits (495), Expect = 1e-49
Identities = 111/193 (57%), Positives = 118/193 (60%)
Frame = +1
Query: 1200 SNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYP 1259
SN FLTTT Q YT S+GL SR + G +SPLK +GYDD SHVEA
Sbjct: 1 SNVFLTTTAQLYTRSSGLTSRIGN-------------GMRSPLKLLGYDDSASHVEA--- 132
Query: 1260 AILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPS 1319
PKTGR GGK SRSP GLPQQ
Sbjct: 133 -------------------------------------PKTGRVQGGKSSRSPGGLPQQSP 201
Query: 1320 GGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGE 1379
QW NI+NFLDSLMS+L NHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGE
Sbjct: 202 VAQWDNIINFLDSLMSRLCANHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGE 381
Query: 1380 YMKSGLAELEKWI 1392
Y+KSGLAELEKWI
Sbjct: 382 YVKSGLAELEKWI 420
>AW156543 similar to PIR|T14275|T1 myosin-like protein my1 - common
sunflower, partial (12%)
Length = 431
Score = 164 bits (414), Expect = 4e-40
Identities = 79/144 (54%), Positives = 102/144 (69%)
Frame = +3
Query: 432 DIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEFVDNQDVL 491
DIYGFE F NSFEQFCIN+ANE+LQQHFN H+FK+EQEEY ++ I+W+ +EF DNQD L
Sbjct: 6 DIYGFESFNRNSFEQFCINYANERLQQHFNRHLFKLEQEEYIQDGIDWAKVEFEDNQDCL 185
Query: 492 ELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISHYAGK 551
L EK+P+G+++LLDE FP T TF+ KL QH S+ E+ F + HYAG+
Sbjct: 186 NLFEKRPLGLLSLLDEESTFPNGTDLTFANKLKQHLNSNSCFKGER--DQAFTVHHYAGQ 359
Query: 552 VTYHTDTFLDKNRDYVVVEHCNLL 575
VTY T FL+KNRD + ++ LL
Sbjct: 360 VTYDTTGFLEKNRDLLHLDSIQLL 431
>TC219407 similar to UP|O24517 (O24517) Unconventional myosin, partial (10%)
Length = 906
Score = 155 bits (393), Expect = 1e-37
Identities = 79/149 (53%), Positives = 105/149 (70%)
Frame = +1
Query: 1380 YMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALT 1439
Y+K+GLAELE W AKEEYAG+SW EL +IRQAVGFLVIHQK + S DEI DLCP ++
Sbjct: 1 YVKAGLAELELWCCQAKEEYAGSSWDELKHIRQAVGFLVIHQKYRISYDEIINDLCPIMS 180
Query: 1440 VRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAED 1499
V+Q+YRI T+YWD Y T+SVS +V+S MR ++ ++D+ N S+SFLLDD SIPFS +D
Sbjct: 181 VQQLYRICTLYWDANYNTRSVSPDVLSSMR-VLMAEDSNNAQSDSFLLDDSSSIPFSVDD 357
Query: 1500 IDMAIPAIDPDEIDLPAFVPEYSCAQFLN 1528
++ D ++ + E +FLN
Sbjct: 358 FSTSLQEKDFSDMKPADELLENPAFRFLN 444
>TC222605 similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain class XI E3 protein,
partial (11%)
Length = 687
Score = 149 bits (376), Expect = 9e-36
Identities = 90/227 (39%), Positives = 123/227 (53%), Gaps = 4/227 (1%)
Frame = +2
Query: 1143 YKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNG 1202
YK LLHW + E+E+T IFD I +++++ L YWLS TS LL LQ ++++
Sbjct: 14 YKSLLHWRSLEAEKTHIFDKITHAFRSSIESQEGIHDLAYWLSTTSTLLFYLQCTMKASN 193
Query: 1203 FLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYPAIL 1262
+ A L + S + S G + + S VEA+YPAIL
Sbjct: 194 TTKAVSRNRNSPATLFGKMAQGLRSSSLGLGISSGYSG---MVDKTNDQSKVEAKYPAIL 364
Query: 1263 FKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLS----RSPSGLPQQP 1318
FKQ LTA VEKI+G++RD LKKE+SP L LCIQAP++ R + S S QQ
Sbjct: 365 FKQHLTAYVEKIYGMIRDSLKKEISPFLNLCIQAPRSIRTRSIRGSSRNIHSNIVAKQQA 544
Query: 1319 SGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNS 1365
W IV+ LD+ + L N+VP RK+ +QVFSF+N+ LFNS
Sbjct: 545 LHMYWKGIVDKLDTALRILSDNYVPPIITRKIFSQVFSFMNVQLFNS 685
>AW133006 similar to GP|29170491|d myosin XI {Nicotiana tabacum}, partial (9%)
Length = 409
Score = 145 bits (367), Expect = 1e-34
Identities = 73/133 (54%), Positives = 97/133 (72%)
Frame = +1
Query: 1254 VEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSG 1313
+ A+YPA+LFKQQLTA VEKI+ +LRD+LKKEL+ +L LCIQAP+T + + RS
Sbjct: 22 IYAKYPALLFKQQLTAYVEKIYCILRDNLKKELASMLSLCIQAPRTSKG----VLRSGRS 189
Query: 1314 LPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECC 1373
+ G W +I+ L++L+ L N VP I+K+ TQ FS+IN+ LFNSLLLRR+CC
Sbjct: 190 FGKDSPMGHWQSIIESLNTLLCTLKENFVPPVLIQKIFTQTFSYINVQLFNSLLLRRDCC 369
Query: 1374 TFSNGEYMKSGLA 1386
TFSNGEY+K+GLA
Sbjct: 370 TFSNGEYVKAGLA 408
>BE820991
Length = 732
Score = 138 bits (347), Expect = 2e-32
Identities = 64/72 (88%), Positives = 69/72 (94%)
Frame = -3
Query: 1367 LLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKS 1426
LLRRECCTFSNGEY+KSG+AELEKWIVNA EEYAGTSWHELNYI QA+GFLVIHQKRKKS
Sbjct: 331 LLRRECCTFSNGEYVKSGVAELEKWIVNATEEYAGTSWHELNYI*QAIGFLVIHQKRKKS 152
Query: 1427 LDEIRQDLCPAL 1438
L+EIR DLCPA+
Sbjct: 151 LEEIRLDLCPAV 116
Score = 65.1 bits (157), Expect(2) = 9e-17
Identities = 31/33 (93%), Positives = 31/33 (93%)
Frame = -1
Query: 1438 LTVRQIYRISTMYWDDKYGTQSVSNEVVSEMRE 1470
LTV IYRISTMYWDDKYGTQSVSNEVVSEMRE
Sbjct: 492 LTVSLIYRISTMYWDDKYGTQSVSNEVVSEMRE 394
Score = 41.6 bits (96), Expect(2) = 9e-17
Identities = 18/24 (75%), Positives = 22/24 (91%)
Frame = -3
Query: 1470 EIVSSKDNQNITSNSFLLDDDMSI 1493
E + SKDNQN+TSNSFLLDDD+S+
Sbjct: 400 ERILSKDNQNLTSNSFLLDDDLSL 329
>TC221199 weakly similar to UP|Q9XEI4 (Q9XEI4) Unconventional myosin heavy
chain, partial (7%)
Length = 763
Score = 134 bits (336), Expect = 4e-31
Identities = 69/149 (46%), Positives = 99/149 (66%), Gaps = 2/149 (1%)
Frame = +3
Query: 1381 MKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTV 1440
+ GL EL W + ++ AG+SW E +IRQAVGFLV HQK +KSL+EI +LCP L++
Sbjct: 12 LXXGLHELXXWCLXXXDQXAGSSWAEXKHIRQAVGFLVXHQKTQKSLEEITNELCPVLSI 191
Query: 1441 RQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDI 1500
QIY I TM+WDDKYG +S EV+S MR ++ ++D+ NI ++SFLL+ D SIPF E++
Sbjct: 192 PQIYCIGTMFWDDKYGAHGLSAEVISRMR-VIMTEDSINIHNSSFLLEVDSSIPFLMEEM 368
Query: 1501 --DMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
M+ + ++D P + + S QFL
Sbjct: 369 FQSMSDIRLSDMDVDPPPILRQRSDFQFL 455
>CA799104
Length = 460
Score = 125 bits (313), Expect = 2e-28
Identities = 64/139 (46%), Positives = 91/139 (65%), Gaps = 2/139 (1%)
Frame = +3
Query: 603 SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRIS 662
SVA++FK QL LM+ L ST PH++RC+KPN+L P+ +E G V+ QLRC GVLE VRIS
Sbjct: 21 SVATKFKGQLFRLMQQLESTTPHFIRCIKPNNLQSPESYEQGLVLQQLRCCGVLEVVRIS 200
Query: 663 LAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL--ENFQLGRTKVFLRA 720
+G+PTR + +F R+G + L+ + S D +V+ IL + + E +Q+G TK+F R
Sbjct: 201 RSGFPTRMFHQKFARRYGFLLLDHV-ASQDPLSVSVVILHQFNILPEMYQVGYTKLFFRT 377
Query: 721 GQIGILDSRRAEVLDNAAR 739
GQIG+L+ R L R
Sbjct: 378 GQIGVLEDTRNRTLHGILR 434
>TC231239 similar to GB|BAB03161.1|11994771|AP002050 myosin-like protein
{Arabidopsis thaliana;} , partial (13%)
Length = 655
Score = 124 bits (311), Expect = 3e-28
Identities = 68/151 (45%), Positives = 96/151 (63%), Gaps = 2/151 (1%)
Frame = +3
Query: 603 SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRIS 662
SVA++FK QL LM+ L ST PH++RC+KPN+L P +E V+ QLRC GVLE VRIS
Sbjct: 186 SVATKFKGQLFQLMQRLESTTPHFIRCIKPNNLQSPGSYEQSLVLQQLRCCGVLEVVRIS 365
Query: 663 LAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL--ENFQLGRTKVFLRA 720
+G+PTR ++ +F R+G + LE + S D +V+ IL + + E +Q+G TK+F R
Sbjct: 366 RSGFPTRVSHQKFARRYGFLLLENV-ASQDPLSVSVAILHQFNILPEMYQVGYTKLFFRT 542
Query: 721 GQIGILDSRRAEVLDNAARCIQRQLRTFIAR 751
GQIG+L+ R L R +Q R + AR
Sbjct: 543 GQIGVLEDTRNRTLHGVLR-VQSCFRGYRAR 632
>BU084190 similar to GP|29170491|d myosin XI {Nicotiana tabacum}, partial (9%)
Length = 412
Score = 120 bits (302), Expect = 3e-27
Identities = 66/144 (45%), Positives = 92/144 (63%)
Frame = +3
Query: 1133 NGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLC 1192
+GKP+AA IYKCLLHW +FE+ERT++FD +I+ I ++ +DD+D + YWLSNTSALL
Sbjct: 3 HGKPIAAFTIYKCLLHWKSFEAERTSVFDRLIQMIGSEIENQDDNDHMAYWLSNTSALLF 182
Query: 1193 LLQRNLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVS 1252
LL+++L+S T R + + +S+L S S + P + D V
Sbjct: 183 LLEQSLKSGSSANATPARKPPNPTSLFGRMTMSFLSSPSSANLAAPPA--------DVVR 338
Query: 1253 HVEARYPAILFKQQLTACVEKIFG 1276
VEA+YPA+LFKQQLTA EKI+G
Sbjct: 339 KVEAKYPALLFKQQLTAYFEKIYG 410
>BF425039 similar to PIR|T14279|T1 myosin-like protein my5 - common
sunflower, partial (8%)
Length = 414
Score = 114 bits (284), Expect = 4e-25
Identities = 62/133 (46%), Positives = 85/133 (63%)
Frame = +3
Query: 807 SSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRMYKFRSAFHRHQASIVAIQCLWRR 866
SSA I+Q+ +R R F K+ +AAT IQA R S + R Q + V QC WRR
Sbjct: 3 SSAIILQTGLRAMKARDEFRFRKQTKAATYIQAYLRRLIAYSYYKRLQKAAVVTQCGWRR 182
Query: 867 RQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQ 926
R A+RELR LK A ETGAL+ AK KLEK+++ELTWRL +EK++R E+ K E +KLQ
Sbjct: 183 RVARRELRMLKMAARETGALKEAKDKLEKRVEELTWRLQIEKRLRTDLEEEKAQETAKLQ 362
Query: 927 KMIEALNLELDAA 939
+ A+ ++++ A
Sbjct: 363 EAXHAMQIQVEEA 401
>BI893301 similar to GP|7243765|gb| unconventional myosin XI {Vallisneria
gigantea}, partial (5%)
Length = 428
Score = 81.6 bits (200), Expect(2) = 5e-24
Identities = 41/58 (70%), Positives = 46/58 (78%)
Frame = -1
Query: 552 VTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVASRFK 609
VTY T+ FLDKN+DYVV EH LL SKCPFVSGLFP PEESS+ S +FSS+ SRFK
Sbjct: 332 VTYQTELFLDKNKDYVVAEHQALLYVSKCPFVSGLFPPSPEESSKQS-KFSSIGSRFK 162
Score = 49.7 bits (117), Expect(2) = 5e-24
Identities = 20/25 (80%), Positives = 25/25 (100%)
Frame = -3
Query: 609 KQQLQALMETLNSTEPHYVRCVKPN 633
+QQLQAL+ETL++TEPHY+RCVKPN
Sbjct: 75 QQQLQALLETLSATEPHYIRCVKPN 1
>TC209068 similar to UP|Q9SMY9 (Q9SMY9) Myosin-like protein, partial (3%)
Length = 568
Score = 103 bits (257), Expect = 6e-22
Identities = 53/63 (84%), Positives = 58/63 (91%)
Frame = +3
Query: 1465 VSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCA 1524
VSEMREIVS KDNQ++TSNSFLLDDDMSIPFSAEDID AIPAI+ D+IDLPAF+ EY CA
Sbjct: 3 VSEMREIVS-KDNQSLTSNSFLLDDDMSIPFSAEDIDKAIPAINTDDIDLPAFLCEYPCA 179
Query: 1525 QFL 1527
QFL
Sbjct: 180 QFL 188
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,117,614
Number of Sequences: 63676
Number of extensions: 862082
Number of successful extensions: 4494
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 4420
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4469
length of query: 1532
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1422
effective length of database: 5,635,272
effective search space: 8013356784
effective search space used: 8013356784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0034a.6


