

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.7
(715 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
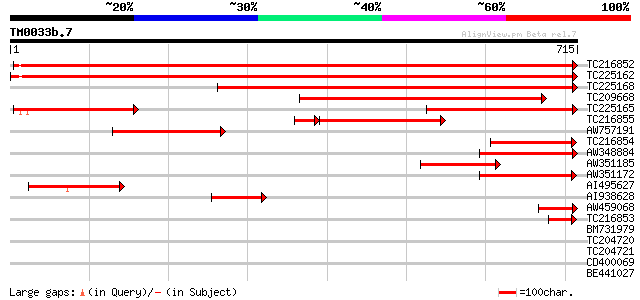
Score E
Sequences producing significant alignments: (bits) Value
TC216852 UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 ,... 1258 0.0
TC225162 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammon... 1230 0.0
TC225168 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammon... 820 0.0
TC209668 homologue to UP|PAL2_CICAR (Q9SMK9) Phenylalanine ammon... 525 e-149
TC225165 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammon... 320 2e-87
TC216855 homologue to PRF|1807329B.0|228615|1807329B Phe ammonia... 266 5e-82
AW757191 similar to SP|P19142|PAL2 Phenylalanine ammonia-lyase c... 202 3e-52
TC216854 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammon... 188 6e-48
AW348884 similar to PIR|JQ1070|JQ1 phenylalanine ammonia-lyase (... 185 7e-47
AW351185 homologue to PIR|JQ1070|JQ10 phenylalanine ammonia-lyas... 171 1e-42
AW351172 similar to SP|P19143|PAL3 Phenylalanine ammonia-lyase c... 168 7e-42
AI495627 similar to SP|P19143|PAL3 Phenylalanine ammonia-lyase c... 154 2e-37
AI938628 similar to SP|P19142|PAL2_ Phenylalanine ammonia-lyase ... 101 1e-21
AW459068 similar to SP|Q43052|PAL2_ Phenylalanine ammonia-lyase ... 89 9e-18
TC216853 UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 ,... 75 1e-13
BM731979 34 0.20
TC204720 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 34 0.20
TC204721 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 32 1.3
CD400069 similar to GP|17862860|gb| RE01153p {Drosophila melanog... 30 4.9
BE441027 29 8.3
>TC216852 UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 , complete
Length = 2311
Score = 1258 bits (3256), Expect = 0.0
Identities = 636/711 (89%), Positives = 670/711 (93%)
Frame = +1
Query: 5 TNATQNGSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTI 64
TN QNGS FC + DPL+WG AAE+MKGSHLDEVKRMVAEYRK VV+LGGETLTI
Sbjct: 10 TNGHQNGS-FCLSTAKGNNDPLNWGAAAEAMKGSHLDEVKRMVAEYRKPVVRLGGETLTI 186
Query: 65 AQVAAIAANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNG 124
AQVAA+A + GV+VEL ESAR GVKASS+WVMNSMNNGTDSYGVTTGFGATSHRRTK G
Sbjct: 187 AQVAAVAGHDHGVAVELSESAREGVKASSEWVMNSMNNGTDSYGVTTGFGATSHRRTKQG 366
Query: 125 NALQLELIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKL 184
ALQ ELIRFLNAGIFGNGTES+HTLP ATRAAMLVRINTLLQGYSGIRFEILEAITKL
Sbjct: 367 GALQKELIRFLNAGIFGNGTESSHTLPHTATRAAMLVRINTLLQGYSGIRFEILEAITKL 546
Query: 185 INNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDS 244
+NNN+TPCL LRGT+TASGDLVPLSYIAGLLTGRPNSKAVGPSGEV+NAK+AF+LA I+S
Sbjct: 547 LNNNVTPCLDLRGTITASGDLVPLSYIAGLLTGRPNSKAVGPSGEVLNAKEAFELASINS 726
Query: 245 GFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLT 304
FFELQPKEGLALVNGTAVGSGLAS+VLF+ANILA+L+EVLSAIFAEVMQGKPEFTDHLT
Sbjct: 727 EFFELQPKEGLALVNGTAVGSGLASMVLFEANILAVLSEVLSAIFAEVMQGKPEFTDHLT 906
Query: 305 HKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIE 364
HKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHE+DPLQKPKQDRYALRTSPQWLGPLIE
Sbjct: 907 HKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEIDPLQKPKQDRYALRTSPQWLGPLIE 1086
Query: 365 VIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFA 424
VIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALA+IGKLMFA
Sbjct: 1087VIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALASIGKLMFA 1266
Query: 425 QFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQ 484
QF+ELV+D YNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQ
Sbjct: 1267QFSELVNDFYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQ 1446
Query: 485 HNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAK 544
HNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLK SVK+TVSQV+K
Sbjct: 1447HNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKNSVKNTVSQVSK 1626
Query: 545 RTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNG 604
R LTTGVNGELHPSRFCEKDLLKVVDRE +F+YIDDPC ATYPLMQKLRQVLVDHALVN
Sbjct: 1627RILTTGVNGELHPSRFCEKDLLKVVDREYIFSYIDDPCSATYPLMQKLRQVLVDHALVNA 1806
Query: 605 EYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVR 664
E EKD +SIFQKIA FE+ELK+LLPKEVESARAAYESG A+PNKI ECRSYPLYKFVR
Sbjct: 1807ESEKDVNSSIFQKIAIFEEELKNLLPKEVESARAAYESGKAAIPNKIQECRSYPLYKFVR 1986
Query: 665 KELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
+ELGT LLTGEK RSPGEE DKLFTA+CQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 1987EELGTGLLTGEKVRSPGEEFDKLFTAMCQGKIIDPLLECLGEWNGAPLPIC 2139
>TC225162 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammonia-lyase
class II , partial (97%)
Length = 2519
Score = 1230 bits (3182), Expect = 0.0
Identities = 621/714 (86%), Positives = 664/714 (92%)
Frame = +2
Query: 2 APNTNATQNGSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGET 61
A NTN N S N +A DPL+WG AAE+M GSHLDEVKRM+ EYR+ VVKLGGET
Sbjct: 89 AANTNFCVNVS---NNGYISANDPLNWGAAAEAMAGSHLDEVKRMLEEYRRPVVKLGGET 259
Query: 62 LTIAQVAAIAANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRT 121
LTI+QVAAIAA+ QGV VEL ES+RAGVKASSDWVM SMN GTDSYGVTTGFGATSHRRT
Sbjct: 260 LTISQVAAIAAHDQGVKVELAESSRAGVKASSDWVMESMNKGTDSYGVTTGFGATSHRRT 439
Query: 122 KNGNALQLELIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAI 181
K G ALQ ELIRFLNAGIFGNGTES TLP ATRAAMLVRINTLLQGYSGIRFEILEAI
Sbjct: 440 KQGAALQKELIRFLNAGIFGNGTESNCTLPHTATRAAMLVRINTLLQGYSGIRFEILEAI 619
Query: 182 TKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAG 241
TKL+NNNITPCLPLRGT+TASGDLVPLSYIAGLLTGRPNSKAVGPSGE++NAK+AF+LA
Sbjct: 620 TKLLNNNITPCLPLRGTITASGDLVPLSYIAGLLTGRPNSKAVGPSGEILNAKEAFELAN 799
Query: 242 IDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTD 301
I + FFELQPKEGLALVNGTAVGSGLASIVLF+ANI+A+L+EV+SAIFAEVMQGKPEFTD
Sbjct: 800 IGAEFFELQPKEGLALVNGTAVGSGLASIVLFEANIIAVLSEVISAIFAEVMQGKPEFTD 979
Query: 302 HLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGP 361
HLTHKLKHHPGQIEAAAIMEHIL+GSSY+KAAKKLHE+DPLQKPKQDRYALRTSPQWLGP
Sbjct: 980 HLTHKLKHHPGQIEAAAIMEHILEGSSYVKAAKKLHEIDPLQKPKQDRYALRTSPQWLGP 1159
Query: 362 LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKL 421
LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALA+IGKL
Sbjct: 1160LIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALASIGKL 1339
Query: 422 MFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQS 481
MFAQF+ELV+D+YNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVT+HVQS
Sbjct: 1340MFAQFSELVNDYYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTSHVQS 1519
Query: 482 AEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQ 541
AEQHNQDVNSLGLISSRKT+EAIEILKLMSSTFL+ALCQAIDLRHLEENLK +VK+ VSQ
Sbjct: 1520AEQHNQDVNSLGLISSRKTHEAIEILKLMSSTFLVALCQAIDLRHLEENLKNTVKNVVSQ 1699
Query: 542 VAKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHAL 601
VAKRTLTTGVNGELHPSRFCEKDLLKVVDRE FAYIDDPC TYPLMQKLRQVLVD+AL
Sbjct: 1700VAKRTLTTGVNGELHPSRFCEKDLLKVVDREYTFAYIDDPCSGTYPLMQKLRQVLVDYAL 1879
Query: 602 VNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYK 661
NGE EK++ TSIFQKIATFE+ELK+LLPKEVE AR AYE+ A+PNKI ECRSYPLYK
Sbjct: 1880ANGENEKNTSTSIFQKIATFEEELKTLLPKEVEGARVAYENDQCAIPNKIKECRSYPLYK 2059
Query: 662 FVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
FVR+ELGT LLTGE+ SPGEECDK+FTA+CQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 2060FVREELGTALLTGERVISPGEECDKVFTALCQGKIIDPLLECLGEWNGAPLPIC 2221
>TC225168 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammonia-lyase
class II , partial (64%)
Length = 1627
Score = 820 bits (2118), Expect = 0.0
Identities = 407/453 (89%), Positives = 434/453 (94%)
Frame = +1
Query: 263 VGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEH 322
VGSGLASIVLF+ANI+A+L+EV+SAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEH
Sbjct: 1 VGSGLASIVLFEANIIAVLSEVISAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEH 180
Query: 323 ILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVND 382
IL+GSSY+KAAKKLHE+DPLQKPKQDRYALRTSPQWLGP IEVIRFSTKSIEREINSVND
Sbjct: 181 ILEGSSYIKAAKKLHEIDPLQKPKQDRYALRTSPQWLGPQIEVIRFSTKSIEREINSVND 360
Query: 383 NPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNL 442
NPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALA+IGKLMFAQF+ELV+D+YNNGLPSNL
Sbjct: 361 NPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALASIGKLMFAQFSELVNDYYNNGLPSNL 540
Query: 443 TASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNE 502
TASRNPSLDYGFKGAEIAMASYCSELQYLANPVT+HVQSAEQHNQDVNSLGLISSRKT+E
Sbjct: 541 TASRNPSLDYGFKGAEIAMASYCSELQYLANPVTSHVQSAEQHNQDVNSLGLISSRKTHE 720
Query: 503 AIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCE 562
AIEILKLMSSTFLIALCQAIDLRHLEENLK +VK+ VSQVAKRTLTTGVNGELHPSRFCE
Sbjct: 721 AIEILKLMSSTFLIALCQAIDLRHLEENLKNTVKNVVSQVAKRTLTTGVNGELHPSRFCE 900
Query: 563 KDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIATFE 622
KDLLKVVDRE FAYIDDPC TYPLMQKLRQVLVD+AL NGE EK++ TSIFQKIATFE
Sbjct: 901 KDLLKVVDREYTFAYIDDPCSGTYPLMQKLRQVLVDYALANGENEKNTNTSIFQKIATFE 1080
Query: 623 DELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGE 682
+ELK+LLPKEVE AR AYE+ A+PNKI ECRSYPLYKFVR+ELGT LLTGE+ SPGE
Sbjct: 1081EELKTLLPKEVEGARVAYENDQCAIPNKIKECRSYPLYKFVREELGTALLTGERVVSPGE 1260
Query: 683 ECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
ECDK+FTA+CQGKIIDPLLECLGEWNGAPLPIC
Sbjct: 1261ECDKVFTAMCQGKIIDPLLECLGEWNGAPLPIC 1359
>TC209668 homologue to UP|PAL2_CICAR (Q9SMK9) Phenylalanine ammonia-lyase 2
, partial (43%)
Length = 940
Score = 525 bits (1352), Expect = e-149
Identities = 261/311 (83%), Positives = 284/311 (90%)
Frame = +3
Query: 366 IRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQ 425
IR +TK IEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLA+A+IGKLMFAQ
Sbjct: 3 IRHATKMIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLAIASIGKLMFAQ 182
Query: 426 FTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQH 485
F+ELV+D YNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVT HVQSAEQH
Sbjct: 183 FSELVNDFYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTNHVQSAEQH 362
Query: 486 NQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKR 545
NQDVNSLGL+SSRKT E I+ILKL +STFL+ALCQAIDLRHLEENLK +V++TVSQVAKR
Sbjct: 363 NQDVNSLGLVSSRKTAEKIDILKLRASTFLVALCQAIDLRHLEENLKNTVRNTVSQVAKR 542
Query: 546 TLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGE 605
LT G+NGELHPSRFCEKDLLK+VDRE ++AYIDDPC ATYPLMQKLR VLVDHAL NG+
Sbjct: 543 VLTVGINGELHPSRFCEKDLLKIVDREYVYAYIDDPCSATYPLMQKLRLVLVDHALQNGD 722
Query: 606 YEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRK 665
E S TSIFQKI FE+EL +LLPKEVESAR E+GNPA+PN+I ECRSYPLYKFVR+
Sbjct: 723 KEASSSTSIFQKIGAFEEELNALLPKEVESARIEVENGNPAIPNRIKECRSYPLYKFVRE 902
Query: 666 ELGTELLTGEK 676
LGT LLTGEK
Sbjct: 903 NLGTTLLTGEK 935
>TC225165 homologue to UP|PAL2_PHAVU (P19142) Phenylalanine ammonia-lyase
class II , partial (46%)
Length = 1221
Score = 320 bits (819), Expect = 2e-87
Identities = 155/190 (81%), Positives = 169/190 (88%)
Frame = +1
Query: 526 HLEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLAT 585
+ EN K +VK+ VSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRE FAYIDDPC T
Sbjct: 532 YASENXKNTVKNVVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDREYTFAYIDDPCSGT 711
Query: 586 YPLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNP 645
YPLMQKLRQVLVD+AL NGE EK++ TSIFQKIATFE+ELK+LLPKEVE AR AYE+
Sbjct: 712 YPLMQKLRQVLVDYALANGENEKNTNTSIFQKIATFEEELKTLLPKEVEGARVAYENDQC 891
Query: 646 AMPNKINECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLG 705
A+PNKI ECRSYPLYKFVR+ELGT LLTGE+ SPGEECDK+FTA+CQGKIIDPLLECLG
Sbjct: 892 AIPNKIKECRSYPLYKFVREELGTALLTGERVISPGEECDKVFTALCQGKIIDPLLECLG 1071
Query: 706 EWNGAPLPIC 715
EWNGAPLPIC
Sbjct: 1072EWNGAPLPIC 1101
Score = 230 bits (587), Expect = 1e-60
Identities = 123/164 (75%), Positives = 133/164 (81%), Gaps = 7/164 (4%)
Frame = +2
Query: 6 NATQNGS---IFCNTVKT----AATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLG 58
N TQ G+ FC +V +A DPL+WG AAE+M GSHLDEVKRMV EYR+ VVKLG
Sbjct: 53 NMTQEGNGNTNFCMSVNNNGYISANDPLNWGAAAEAMAGSHLDEVKRMVEEYRRPVVKLG 232
Query: 59 GETLTIAQVAAIAANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSH 118
GETLTI+QVAAIAA+ QGV VEL ES+RAGVKASSDWVM SM+ GTDSYGVTTGFGATSH
Sbjct: 233 GETLTISQVAAIAAHDQGVKVELAESSRAGVKASSDWVMESMDKGTDSYGVTTGFGATSH 412
Query: 119 RRTKNGNALQLELIRFLNAGIFGNGTESTHTLPQPATRAAMLVR 162
RRTK G AL ELIRFLNAGIFGNGTES TLP ATRAAMLVR
Sbjct: 413 RRTKQGGALHNELIRFLNAGIFGNGTESNCTLPHTATRAAMLVR 544
>TC216855 homologue to PRF|1807329B.0|228615|1807329B Phe ammonia lyase.
{Phaseolus vulgaris;} , partial (27%)
Length = 581
Score = 266 bits (679), Expect(2) = 5e-82
Identities = 131/159 (82%), Positives = 150/159 (93%)
Frame = +3
Query: 391 NKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSL 450
NKAL+GGNFQGTPIGVSMDN RLA+A+IGKL+FAQFTELV+D YNNGLPSNL+A RNPSL
Sbjct: 102 NKALNGGNFQGTPIGVSMDNARLAVASIGKLIFAQFTELVNDLYNNGLPSNLSAGRNPSL 281
Query: 451 DYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLM 510
DYGFK +E+AMA+YCSELQYLANPVT+HVQSAEQHNQDVNSLGLIS+ KT EA+EILKLM
Sbjct: 282 DYGFKASEVAMAAYCSELQYLANPVTSHVQSAEQHNQDVNSLGLISALKTVEAVEILKLM 461
Query: 511 SSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTT 549
SST+L+ALCQAIDLRHLEEN K +VK+TVS+VA++TL T
Sbjct: 462 SSTYLVALCQAIDLRHLEENFKSTVKNTVSRVAQKTLIT 578
Score = 58.2 bits (139), Expect(2) = 5e-82
Identities = 27/31 (87%), Positives = 30/31 (96%)
Frame = +2
Query: 360 GPLIEVIRFSTKSIEREINSVNDNPLIDVSR 390
GP IE+IR+STKSIEREINSVNDNPLIDV+R
Sbjct: 8 GPQIEIIRYSTKSIEREINSVNDNPLIDVTR 100
>AW757191 similar to SP|P19142|PAL2 Phenylalanine ammonia-lyase class II (EC
4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (19%)
Length = 430
Score = 202 bits (515), Expect = 3e-52
Identities = 106/143 (74%), Positives = 119/143 (83%)
Frame = +2
Query: 130 ELIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNI 189
ELIRFL AGIFG GTES TLP AT A ML+RIN LLQ YSGIRFEILEAITKL+NNNI
Sbjct: 2 ELIRFLKAGIFGIGTESNCTLPHTATIATMLLRINKLLQCYSGIRFEILEAITKLLNNNI 181
Query: 190 TPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFEL 249
PC+PLRGT+T+SGDLVPLSYIAGLLTG PNSKAVGPSG+++ A AF+L I + F L
Sbjct: 182 APCMPLRGTITSSGDLVPLSYIAGLLTG*PNSKAVGPSGDILYAI*AFELTNIGAEFCVL 361
Query: 250 QPKEGLALVNGTAVGSGLASIVL 272
QPKEGL LVNGT++GSG SIV+
Sbjct: 362 QPKEGLVLVNGTSIGSG*YSIVI 430
>TC216854 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1
, partial (15%)
Length = 512
Score = 188 bits (478), Expect = 6e-48
Identities = 91/108 (84%), Positives = 98/108 (90%)
Frame = +2
Query: 607 EKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKE 666
EKD +SIFQKIA FE+ELK+LLPKEVESARAAYESG A+PNKI ECR YPLYKFVR+E
Sbjct: 2 EKDVNSSIFQKIAIFEEELKNLLPKEVESARAAYESGKAAIPNKIQECRFYPLYKFVREE 181
Query: 667 LGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI 714
LGT LLTGEK RSPGEE DKLFTA+CQGKIIDPL+ECLGEWNGAPLPI
Sbjct: 182 LGTGLLTGEKVRSPGEEFDKLFTAMCQGKIIDPLMECLGEWNGAPLPI 325
>AW348884 similar to PIR|JQ1070|JQ1 phenylalanine ammonia-lyase (EC 4.3.1.5)
- soybean (fragment), partial (29%)
Length = 455
Score = 185 bits (469), Expect = 7e-47
Identities = 89/123 (72%), Positives = 102/123 (82%)
Frame = -2
Query: 593 RQVLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKIN 652
RQVLV++AL E K++ TSIFQKIATFE+ L +L KEVE AR AYE+ A+PNKI
Sbjct: 454 RQVLVNYALAXXEXXKNTNTSIFQKIATFEE*LXTLXXKEVEGARVAYENDQCAIPNKIK 275
Query: 653 ECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPL 712
ECRSYPLYKFVR+ELGT LLTGE+ SPGEECDK+FT +CQGKIIDPLLECLG+WNGAPL
Sbjct: 274 ECRSYPLYKFVREELGTALLTGERVVSPGEECDKVFTVMCQGKIIDPLLECLGDWNGAPL 95
Query: 713 PIC 715
P C
Sbjct: 94 PTC 86
>AW351185 homologue to PIR|JQ1070|JQ10 phenylalanine ammonia-lyase (EC
4.3.1.5) - soybean (fragment), partial (23%)
Length = 448
Score = 171 bits (433), Expect = 1e-42
Identities = 84/101 (83%), Positives = 91/101 (89%)
Frame = -2
Query: 519 CQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRETLFAYI 578
CQAIDLRHLEENLK +VK+ VSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDRE FAYI
Sbjct: 447 CQAIDLRHLEENLKNTVKNVVSQVAKRTLTTGVNGELHPSRFCEKDLLKVVDREYTFAYI 268
Query: 579 DDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQKIA 619
DDPC TYPLMQKLRQVLVD+AL NGE EK++ TSIFQ ++
Sbjct: 267 DDPCSGTYPLMQKLRQVLVDYALANGENEKNTNTSIFQYVS 145
>AW351172 similar to SP|P19143|PAL3 Phenylalanine ammonia-lyase class III (EC
4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (17%)
Length = 508
Score = 168 bits (426), Expect = 7e-42
Identities = 78/122 (63%), Positives = 99/122 (80%)
Frame = -3
Query: 593 RQVLVDHALVNGEYEKDSKTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKIN 652
+QVL + A ++ +K+ IF+KI FEDELKSLLPKEVE+AR AYE+GNPA+PN+I
Sbjct: 506 KQVLYEKAHISAINDKNVSLLIFEKIGAFEDELKSLLPKEVENARVAYENGNPAIPNRIK 327
Query: 653 ECRSYPLYKFVRKELGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPL 712
ECRSYPLYKFVR+EL LLTGEK SP EE +K++TA+CQ KI+DP+LECLG+W G+P+
Sbjct: 326 ECRSYPLYKFVREELEIGLLTGEKNLSPDEEFEKVYTAMCQAKIVDPILECLGDWKGSPI 147
Query: 713 PI 714
PI
Sbjct: 146 PI 141
>AI495627 similar to SP|P19143|PAL3 Phenylalanine ammonia-lyase class III (EC
4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (17%)
Length = 412
Score = 154 bits (388), Expect = 2e-37
Identities = 81/127 (63%), Positives = 97/127 (75%), Gaps = 5/127 (3%)
Frame = +1
Query: 24 DPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGG-ETLTIAQVAAIA---ANHQ-GVS 78
DPL+W AA+S+KGSH +EVKRMVAEYRK ++ LGG ETLTI+QVAA+A ANH
Sbjct: 28 DPLNWSHAADSLKGSHFEEVKRMVAEYRKPLISLGGGETLTISQVAAVAVANANHNLQAK 207
Query: 79 VELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAG 138
V+L ESARAGV AS DW+ ++N GT YGVTTGFGA SHR+T+ G ALQ E++RFLN
Sbjct: 208 VDLSESARAGVDASCDWITQNINKGTPIYGVTTGFGAASHRQTQQGLALQKEMVRFLNCA 387
Query: 139 IFGNGTE 145
IFG TE
Sbjct: 388 IFGYQTE 408
>AI938628 similar to SP|P19142|PAL2_ Phenylalanine ammonia-lyase class II (EC
4.3.1.5). [Kidney bean French bean] {Phaseolus
vulgaris}, partial (9%)
Length = 208
Score = 101 bits (251), Expect = 1e-21
Identities = 52/69 (75%), Positives = 58/69 (83%)
Frame = +2
Query: 255 LALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQI 314
LALVNGTA GSGLASI LF+ANI+A L+E +S IFAE+MQG+PE TDHL H LKHHPGQ
Sbjct: 2 LALVNGTAGGSGLASIGLFEANIIAGLSEGISTIFAEMMQGEPEVTDHLNH*LKHHPGQN 181
Query: 315 EAAAIMEHI 323
A AIMEHI
Sbjct: 182 GACAIMEHI 208
>AW459068 similar to SP|Q43052|PAL2_ Phenylalanine ammonia-lyase G2B (EC
4.3.1.5). [Aspen] {Populus kitakamiensis}, partial (6%)
Length = 391
Score = 88.6 bits (218), Expect = 9e-18
Identities = 38/49 (77%), Positives = 44/49 (89%)
Frame = -3
Query: 667 LGTELLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
LGT LLTGEK +SPGEE DK+FTA+C+GK+IDPLL+CL EW GAPLPIC
Sbjct: 386 LGTTLLTGEKVKSPGEEFDKVFTALCEGKLIDPLLDCLKEWKGAPLPIC 240
>TC216853 UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1 , partial
(5%)
Length = 278
Score = 74.7 bits (182), Expect = 1e-13
Identities = 32/35 (91%), Positives = 34/35 (96%)
Frame = +3
Query: 680 PGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPI 714
PGEE DKLFTA+CQGKIIDPL+ECLGEWNGAPLPI
Sbjct: 3 PGEEFDKLFTAMCQGKIIDPLMECLGEWNGAPLPI 107
>BM731979
Length = 418
Score = 34.3 bits (77), Expect = 0.20
Identities = 27/83 (32%), Positives = 38/83 (45%), Gaps = 5/83 (6%)
Frame = +3
Query: 10 NGSIFCNTVKTAATDPLSWGVAAESMKGSHL---DEVKRMVAEYRKAVVKLGGETL--TI 64
NGSI T +T+ TDPLS V A + + L M ++ R LG L ++
Sbjct: 27 NGSIMVGTEQTSHTDPLSGSVVAVLLPHNSLVTGTSNSAMASDDRSGASSLGALVLQPSL 206
Query: 65 AQVAAIAANHQGVSVELCESARA 87
A +A H G + E E+A A
Sbjct: 207 PVAAGVAERHDGEAAEG*EAAGA 275
>TC204720 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (31%)
Length = 1936
Score = 34.3 bits (77), Expect = 0.20
Identities = 17/66 (25%), Positives = 33/66 (49%)
Frame = -3
Query: 141 GNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLRGTVT 200
GN TH L P++R + + I + L + ++ ++L+ +T N+N P + +
Sbjct: 302 GNSIRRTHKLNSPSSRIRISIHITSNLNFGTRLQLKVLDGLTTFSNDNTNPPIGDINLLA 123
Query: 201 ASGDLV 206
+ DLV
Sbjct: 122 RTADLV 105
>TC204721 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (30%)
Length = 1762
Score = 31.6 bits (70), Expect = 1.3
Identities = 15/53 (28%), Positives = 28/53 (52%)
Frame = -1
Query: 141 GNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCL 193
GN S H L +P+ R + + I + L + ++ EIL +++ NN+ P +
Sbjct: 172 GNCFRSAHKLNRPSGRIRVRINITSNLNFGTRLQLEILNSLSAFTNNHTNPAI 14
>CD400069 similar to GP|17862860|gb| RE01153p {Drosophila melanogaster},
partial (14%)
Length = 485
Score = 29.6 bits (65), Expect = 4.9
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +3
Query: 421 LMFAQFTELVDDHYNNGLPSNLTASRNPSL 450
L F + +V H NNG P NL S++P++
Sbjct: 42 LFFYKKNTMVSKHPNNGFPQNLRGSKHPNV 131
>BE441027
Length = 472
Score = 28.9 bits (63), Expect = 8.3
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = -1
Query: 468 LQYLANPVTTHVQSAEQHNQDVNSLGLISS 497
L YL +PVTT A HNQ+V + L S+
Sbjct: 151 LSYLKSPVTTRALHALDHNQEVKVIVLSSA 62
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,470,757
Number of Sequences: 63676
Number of extensions: 313254
Number of successful extensions: 1554
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1546
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1552
length of query: 715
length of database: 12,639,632
effective HSP length: 104
effective length of query: 611
effective length of database: 6,017,328
effective search space: 3676587408
effective search space used: 3676587408
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0033b.7


