

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.5
(88 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
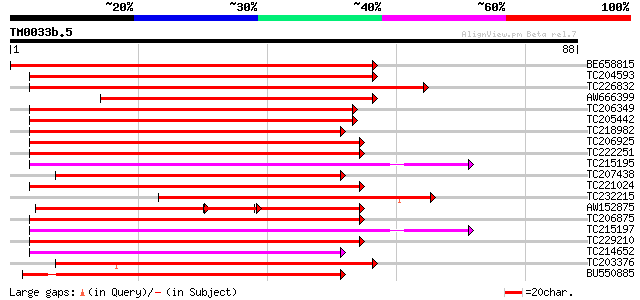
Score E
Sequences producing significant alignments: (bits) Value
BE658815 similar to GP|12311680|em GTP binding protein {Cichoriu... 92 3e-20
TC204593 homologue to UP|Q9AW99 (Q9AW99) GTP binding protein, co... 73 3e-14
TC226832 homologue to UP|Q40210 (Q40210) RAB5B, partial (98%) 70 2e-13
AW666399 homologue to SP|P29687|RAB5_ Ras-related protein Rab5. ... 59 5e-10
TC206349 UP|P92963 (P92963) Rab2-like protein (GTP-binding RAB2A... 54 1e-08
TC205442 UP|Q40208 (Q40208) RAB2A, complete 54 1e-08
TC218982 homologue to UP|Q38922 (Q38922) ATGB2 (GTP-binding prot... 53 2e-08
TC206925 homologue to UP|Q39844 (Q39844) Small GTP-binding prote... 50 1e-07
TC222251 similar to UP|Q6ZXT1 (Q6ZXT1) Small GTPase, partial (94%) 50 1e-07
TC215195 similar to UP|Q9LFT9 (Q9LFT9) GTP-binding protein, comp... 49 3e-07
TC207438 homologue to UP|Q08145 (Q08145) GTP-binding protein, pa... 49 4e-07
TC221024 similar to UP|Q40196 (Q40196) RAB11F, partial (87%) 49 4e-07
TC232215 similar to UP|Q40209 (Q40209) RAB5A, partial (68%) 49 6e-07
AW152875 34 1e-06
TC206875 UP|Q39844 (Q39844) Small GTP-binding protein (Fragment)... 47 1e-06
TC215197 homologue to PIR|T01588|T01588 GTP-binding protein At2g... 46 3e-06
TC229210 similar to UP|Q08149 (Q08149) GTP-binding protein, part... 46 4e-06
TC214652 homologue to UP|Q39860 (Q39860) GTPase, complete 46 4e-06
TC203376 homologue to UP|Q41023 (Q41023) Small GTP-binding prote... 45 6e-06
BU550885 weakly similar to SP|P25766|RGP1 Ras-related protein RG... 45 6e-06
>BE658815 similar to GP|12311680|em GTP binding protein {Cichorium intybus x
Cichorium endivia}, partial (68%)
Length = 561
Score = 92.4 bits (228), Expect = 3e-20
Identities = 46/57 (80%), Positives = 50/57 (87%)
Frame = -1
Query: 1 SQILVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+Q VMALVANKSDLEPKREVE E G QFAQENGMFYMETSAKT ENIN+LF+EI +
Sbjct: 336 NQKSVMALVANKSDLEPKREVEAEVGEQFAQENGMFYMETSAKTAENINELFYEIAK 166
>TC204593 homologue to UP|Q9AW99 (Q9AW99) GTP binding protein, complete
Length = 1072
Score = 72.8 bits (177), Expect = 3e-14
Identities = 32/54 (59%), Positives = 43/54 (79%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DLE KR+V EE +A+ENG+F+METSAKT N+ND+F+EI +
Sbjct: 577 MVMALAGNKADLEDKRKVTAEEARVYAEENGLFFMETSAKTASNVNDIFYEIAK 738
>TC226832 homologue to UP|Q40210 (Q40210) RAB5B, partial (98%)
Length = 1214
Score = 70.1 bits (170), Expect = 2e-13
Identities = 33/62 (53%), Positives = 46/62 (73%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+VMALV NK+DL KREV +++G +A++N MF++ETSAKT +NIN+LF EI + P
Sbjct: 663 IVMALVGNKADLLEKREVAVQDGTDYAEKNDMFFIETSAKTADNINELFEEIAKRLPRPS 842
Query: 64 RS 65
S
Sbjct: 843 VS 848
>AW666399 homologue to SP|P29687|RAB5_ Ras-related protein Rab5. [Common
tobacco] {Nicotiana tabacum}, partial (23%)
Length = 424
Score = 58.5 bits (140), Expect = 5e-10
Identities = 25/43 (58%), Positives = 34/43 (78%)
Frame = +3
Query: 15 LEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LE KR+V EE +A+ENG+F+METSAKT N+ND+F+EI +
Sbjct: 282 LEDKRKVTAEEARVYAEENGLFFMETSAKTASNVNDIFYEIAK 410
>TC206349 UP|P92963 (P92963) Rab2-like protein (GTP-binding RAB2A like
protein) (At4g17170), partial (90%)
Length = 728
Score = 54.3 bits (129), Expect = 1e-08
Identities = 23/51 (45%), Positives = 35/51 (68%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
+ + L+ NK DL +R V EEG QFA+E+G+ +ME SAKT +N+ + F +
Sbjct: 493 MTIMLIGNKCDLAHRRAVSTEEGEQFAKEHGLIFMEASAKTAQNVEEAFIK 645
>TC205442 UP|Q40208 (Q40208) RAB2A, complete
Length = 1159
Score = 54.3 bits (129), Expect = 1e-08
Identities = 23/51 (45%), Positives = 35/51 (68%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
+ + L+ NK DL +R V EEG QFA+E+G+ +ME SAKT +N+ + F +
Sbjct: 431 MTIMLIGNKCDLAHRRAVSTEEGEQFAKEHGLIFMEASAKTAQNVEEAFIK 583
>TC218982 homologue to UP|Q38922 (Q38922) ATGB2 (GTP-binding protein GB2),
complete
Length = 939
Score = 53.1 bits (126), Expect = 2e-08
Identities = 22/49 (44%), Positives = 34/49 (68%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+ + L+ NK DL +R V EEG QFA+ENG+ ++E SA+T +N+ + F
Sbjct: 462 MTIMLIGNKCDLSHRRAVSKEEGEQFAKENGLLFLEASARTAQNVEEAF 608
>TC206925 homologue to UP|Q39844 (Q39844) Small GTP-binding protein
(Fragment), partial (95%)
Length = 1111
Score = 50.4 bits (119), Expect = 1e-07
Identities = 21/52 (40%), Positives = 35/52 (66%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L+ NKSDLE +R+V E+ +FA++ G+F++ETSA N+ F +
Sbjct: 619 IVIILIGNKSDLENQRQVPTEDAKEFAEKEGLFFLETSALEATNVETAFMTV 774
>TC222251 similar to UP|Q6ZXT1 (Q6ZXT1) Small GTPase, partial (94%)
Length = 911
Score = 50.4 bits (119), Expect = 1e-07
Identities = 24/52 (46%), Positives = 35/52 (67%)
Frame = +1
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NK DL+ REVE EEG FA+ G+ +METSA N+ ++F ++
Sbjct: 526 MVVVLVGNKCDLDESREVEKEEGKGFAETEGLCFMETSALKNLNVEEVFLQM 681
>TC215195 similar to UP|Q9LFT9 (Q9LFT9) GTP-binding protein, complete
Length = 1022
Score = 49.3 bits (116), Expect = 3e-07
Identities = 28/69 (40%), Positives = 41/69 (58%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V IEEG A+E + ++ETSAK NI LF +I PG
Sbjct: 456 VIIVLVGNKTDLVEKRQVSIEEGEAKARELNVMFIETSAKAGFNIKALFRKIAAAL--PG 629
Query: 64 RSSYAPTKE 72
+ + K+
Sbjct: 630 METLSSAKQ 656
>TC207438 homologue to UP|Q08145 (Q08145) GTP-binding protein, partial (95%)
Length = 853
Score = 48.9 bits (115), Expect = 4e-07
Identities = 24/45 (53%), Positives = 28/45 (61%)
Frame = +2
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
LV NK DLE REV EEG A+E G+F+METSA N+ F
Sbjct: 479 LVGNKCDLENIREVSTEEGKSLAEEEGLFFMETSALDATNVQTAF 613
>TC221024 similar to UP|Q40196 (Q40196) RAB11F, partial (87%)
Length = 769
Score = 48.9 bits (115), Expect = 4e-07
Identities = 24/52 (46%), Positives = 35/52 (67%)
Frame = +2
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NKSDL+ R+VE EEG FA+ + +METSA N+++ F E+
Sbjct: 290 MVVVLVGNKSDLDQSRQVEREEGKVFAETEELCFMETSALQNLNVDEAFLEM 445
>TC232215 similar to UP|Q40209 (Q40209) RAB5A, partial (68%)
Length = 718
Score = 48.5 bits (114), Expect = 6e-07
Identities = 21/45 (46%), Positives = 34/45 (74%), Gaps = 2/45 (4%)
Frame = +2
Query: 24 EEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGF--ESPGRSS 66
+EG ++A+ENG+ ++ETSAKT +N+N+LF+EI + +P R S
Sbjct: 296 KEGEEYAKENGLSFLETSAKTAQNVNELFYEIAKRLAKANPSRQS 430
>AW152875
Length = 411
Score = 33.9 bits (76), Expect(3) = 1e-06
Identities = 18/27 (66%), Positives = 20/27 (73%)
Frame = +1
Query: 5 VMALVANKSDLEPKREVEIEEGGQFAQ 31
VMAL ANKS LEPKR+VE + FAQ
Sbjct: 229 VMALQANKSYLEPKRQVETQV*QHFAQ 309
Score = 30.8 bits (68), Expect(3) = 1e-06
Identities = 13/17 (76%), Positives = 16/17 (93%)
Frame = +3
Query: 39 ETSAKTVENINDLFFEI 55
+TSAKT E+INDLF+EI
Sbjct: 333 KTSAKTAEDINDLFYEI 383
Score = 21.2 bits (43), Expect(3) = 1e-06
Identities = 7/9 (77%), Positives = 9/9 (99%)
Frame = +2
Query: 31 QENGMFYME 39
++NGMFYME
Sbjct: 308 KQNGMFYME 334
>TC206875 UP|Q39844 (Q39844) Small GTP-binding protein (Fragment), complete
Length = 871
Score = 47.4 bits (111), Expect = 1e-06
Identities = 20/52 (38%), Positives = 33/52 (63%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ L NK DLE +R+V E+ +FA++ G+F++ETSA N+ F +
Sbjct: 393 IVIILTGNKCDLENQRDVPTEDAKEFAEKEGLFFLETSALEATNVETAFITV 548
>TC215197 homologue to PIR|T01588|T01588 GTP-binding protein At2g44610 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(97%)
Length = 981
Score = 46.2 bits (108), Expect = 3e-06
Identities = 26/69 (37%), Positives = 40/69 (57%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V EEG ++E + ++E SAK NI LF +I PG
Sbjct: 480 VIVVLVGNKTDLVDKRQVSTEEGEAKSRELNVMFIEASAKAGFNIKALFRKIAAAL--PG 653
Query: 64 RSSYAPTKE 72
+ + TK+
Sbjct: 654 METLSTTKQ 680
>TC229210 similar to UP|Q08149 (Q08149) GTP-binding protein, partial (90%)
Length = 962
Score = 45.8 bits (107), Expect = 4e-06
Identities = 21/52 (40%), Positives = 34/52 (65%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEI 55
+V+ LV NK+DL R V EE +FA++ +++METSA N+++ F E+
Sbjct: 471 VVVMLVGNKADLRHLRAVSTEEATEFAEKESIYFMETSALESLNVDNAFIEV 626
>TC214652 homologue to UP|Q39860 (Q39860) GTPase, complete
Length = 1004
Score = 45.8 bits (107), Expect = 4e-06
Identities = 22/49 (44%), Positives = 29/49 (58%)
Frame = +3
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+ M LV NK DLE R V ++EG A+ G+F+METSA N+ F
Sbjct: 432 VAMMLVGNKCDLENIRAVSVDEGKSLAEAEGLFFMETSALDSTNVKTAF 578
>TC203376 homologue to UP|Q41023 (Q41023) Small GTP-binding protein, complete
Length = 1011
Score = 45.1 bits (105), Expect = 6e-06
Identities = 24/51 (47%), Positives = 32/51 (62%), Gaps = 1/51 (1%)
Frame = +2
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF I R
Sbjct: 482 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNMNVEEVFFSIAR 634
>BU550885 weakly similar to SP|P25766|RGP1 Ras-related protein RGP1
(GTP-binding regulatory protein RGP1). [Rice] {Oryza
sativa}, partial (47%)
Length = 694
Score = 45.1 bits (105), Expect = 6e-06
Identities = 25/50 (50%), Positives = 30/50 (60%)
Frame = -2
Query: 3 ILVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
ILVM LV NKSDL R V E FAQ+ G+F++ETSA N+ F
Sbjct: 558 ILVM-LVGNKSDLSSLRAVPTEVARDFAQQEGLFFLETSALDSSNVESAF 412
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,351,495
Number of Sequences: 63676
Number of extensions: 33443
Number of successful extensions: 184
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 184
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 184
length of query: 88
length of database: 12,639,632
effective HSP length: 64
effective length of query: 24
effective length of database: 8,564,368
effective search space: 205544832
effective search space used: 205544832
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0033b.5


