

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
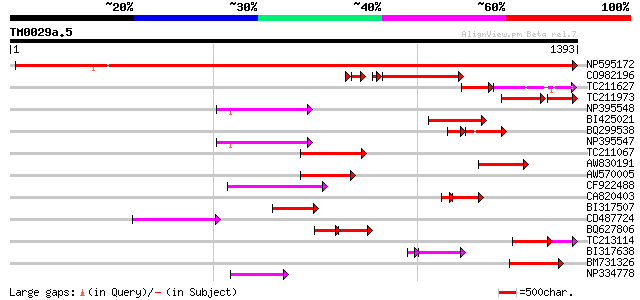
Query= TM0029a.5
(1393 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 1154 0.0
CO982196 245 1e-73
TC211627 116 7e-49
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 118 1e-43
NP395548 reverse transcriptase [Glycine max] 164 2e-40
BI425021 159 8e-39
BQ299538 108 4e-36
NP395547 reverse transcriptase [Glycine max] 150 5e-36
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 147 2e-35
AW830191 147 4e-35
AW570005 141 2e-33
CF922488 140 4e-33
CA820403 weakly similar to GP|13273463|gb| pol protein integrase... 108 5e-33
BI317507 124 3e-28
CD487724 120 4e-27
BQ627806 76 2e-26
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 115 1e-25
BI317638 weakly similar to GP|9294238|dbj| contains similarity t... 93 4e-22
BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza... 102 2e-21
NP334778 reverse transcriptase [Glycine max] 100 4e-21
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 1154 bits (2986), Expect = 0.0
Identities = 616/1406 (43%), Positives = 858/1406 (60%), Gaps = 27/1406 (1%)
Frame = +1
Query: 15 RVDIPMFNGNDAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLERA 74
++D P F+G + W+ K E+FF +A+++ + + ++ + W+Q ++ +
Sbjct: 295 KLDFPRFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQMLQKTEPFSS 474
Query: 75 WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLKGVFL 134
W+ F +AL F P+ P L + Q +V EY F L + E + F+
Sbjct: 475 WQAFTRALELDFGPSAYDCPRATLFKLNQSATVNEYYMQFTALVNRVDGLSAEAILDCFV 654
Query: 135 NGLQEEIKAEMKLYPADDLAELMDRALLLEEKNTAMRGGKPKEEDKRGWKD---LQNKGG 191
+GLQEEI ++K L + + A L EEK T+ K R + K
Sbjct: 655 SGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTSATQKYP 834
Query: 192 TGNQDTEGKQPE----------KKWN----GGQRLTQTELQERSRKGLCFKCGDKWGKEH 237
NQ + +P K +N ++++ E+Q R K LC+ C +K+ H
Sbjct: 835 PTNQKNDNPKPNLPPLLPTPSTKPFNLRNQNIKKISPAEIQLRREKNLCYFCDEKFSPAH 1014
Query: 238 ICSMKNYQLILMEVEEDEEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGK 297
C N Q++L+++EE +E++ + E ++ LSLN+ G + + G+
Sbjct: 1015 KCP--NRQVMLLQLEETDEDQTDEQVMVTEEANMDDDTHHLSLNAMRGSNGVGTIRFTGQ 1188
Query: 298 IGNREVLILIDCGATSNFISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEV 357
+G V IL+D G++ NFI + L++PV V VGNG G+ + L L +
Sbjct: 1189 VGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLVGNGQILSAEGIVQQLPLHI 1368
Query: 358 QGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCR 417
QG + ++L + G +V+LG WLA+LG A++ L +++ + + LQGE +
Sbjct: 1369 QGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALTLKFFQNDKFITLQGEGNSEA 1548
Query: 418 VTANWKSIK-ITEQQEAEGYYLSYEYQKE-EEKTEAEVPEGMRK----ILEEYPEVFQEP 471
A + + + E + QKE E T ++P + +L Y +VF P
Sbjct: 1549 TQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPTNIDPELAILLHTYAQVFAVP 1728
Query: 472 KGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSP 531
LPP+R DHAI L++G+ +RPYRYP QK++IEK+++EML GII+ S SPFS P
Sbjct: 1729 ASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSPFSLP 1908
Query: 532 AILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIR 591
+LVKKKDG WRFC DYRALN T+ D FP+P +DELLDE+ A FSKLDL+SGYHQI
Sbjct: 1909 ILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQIL 2088
Query: 592 MKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIY 651
++ ED KTAFRTH GHYE+LV+PFGLTNAP+TFQ LMN++ + LRKFVLVFFDDILIY
Sbjct: 2089 VQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIY 2268
Query: 652 SKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDML 711
S + + H HL VLQ LK++ L A KCSFG E+ YLGH +S GV+ + +K++ +L
Sbjct: 2269 SASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVL 2448
Query: 712 DWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQAFVKLKEV 771
DWP P VK LRGFLGLTGYYRRF+K+Y+ +A PL LL+K+SF W A AFVKLK+
Sbjct: 2449 DWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDSFLWNNEAEAAFVKLKKA 2628
Query: 772 MTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELM 831
MT PVL P+F +PFILETDASG G+GAVL Q G P+AY SK L+ R Q +S Y REL+
Sbjct: 2629 MTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNGHPIAYFSKKLAPRMQKQSAYTRELL 2808
Query: 832 AVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGI 891
A+ A+ K+RHYLLG+KF+I TDQRSL+ L DQ + EQQ W+ K +GYDF+I+YKPG
Sbjct: 2809 AITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIEYKPGK 2988
Query: 892 ENKAADALSRKLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLKRG 951
+N+AADALSR A S +L A ++ D + K L E QG A Y ++ G
Sbjct: 2989 DNQAADALSRMFML-AWSEPHSIFLEELRARLISDP-HLKQLMETYKQGADASHYTVREG 3162
Query: 952 RLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQ 1011
L +KDR+V+P + +L+E+H + +GGHAGI RT R+ A FYW M+ D++ Y+Q
Sbjct: 3163 LLYWKDRVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLARLKAQFYWPKMQEDVKAYIQ 3342
Query: 1012 KCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYA 1071
KC +CQ+ K PAG LQPLPIP Q W D++MDFI GLP + G I+VV+DR TKYA
Sbjct: 3343 KCLICQQAKSNNTLPAGLLQPLPIPQQVWEDVAMDFITGLPNSFGLSVIMVVIDRLTKYA 3522
Query: 1072 HFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSS 1131
HFI L YN+K +AE F+ +V+LHG P SIVSDRDRVF STFW +FKL GT L SS
Sbjct: 3523 HFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSS 3702
Query: 1132 AYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALY 1191
AYHPQ+DGQ+EV+N+C+E YLRC T PK W K L WAEFWYNT YH ++ TPF+ALY
Sbjct: 3703 AYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALY 3882
Query: 1192 GREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEV 1251
GREPP + + S+ EV + +R+ +L +LK NL +AQ M++QA+K R DV +++
Sbjct: 3883 GREPPTLTRQACSIDDPAEVREQLTDRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQI 4062
Query: 1252 GDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHIS 1311
GD V +K+QPY+ S R NQKLS RY+GP+ ++AKI AYKL+LP +++HPVFH+S
Sbjct: 4063 GDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHPVFHVS 4242
Query: 1312 LLKKAVNAGVQSQPLPAALT-EEWELKVEPEAIMDTR---ENRDGDLEVLIRWKDLPTFE 1367
L K N Q LP LT E ++P I+ +R + ++L++W++ E
Sbjct: 4243 QL-KPFNGTAQDPYLPLPLTVTEMGPVMQPVKILASRIIIRGHNQIEQILVQWENGLQDE 4419
Query: 1368 DSWEDFSKLLDQFPNHQLEDKLNLQG 1393
+WED + +P LEDK+ +G
Sbjct: 4420 ATWEDIEDIKASYPTFNLEDKVVFKG 4497
>CO982196
Length = 812
Score = 245 bits (626), Expect(4) = 1e-73
Identities = 119/201 (59%), Positives = 149/201 (73%)
Frame = +1
Query: 915 EWADLEAEILEDERYRKVLQELATQGNSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKE 974
E AD E EI ++ Q + T+ GY ++ G+L +KDR+VL K STKI +LKE
Sbjct: 208 ELADWEEEIQAYLELYEIYQGILTKTTKKPGYAIRGGKLYFKDRLVLSKNSTKIPLLLKE 387
Query: 975 FHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLP 1034
D+ LGGH+G FRT+KR++ + +W+GMK ++YV CE+C+RNK L+PAG L LP
Sbjct: 388 LQDSPLGGHSGFFRTFKRVANVVFWQGMKKTTRDYVAACEICRRNKTSTLSPAGLL*LLP 567
Query: 1035 IPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVV 1094
IP++ WTDISMDFIGGLPKA GKD ILVVVDR TKYAHF ALSHPY AKE+AE+FIKE+V
Sbjct: 568 IPTKVWTDISMDFIGGLPKAQGKDNILVVVDRLTKYAHFFALSHPYTAKEVAELFIKELV 747
Query: 1095 RLHGFPTSIVSDRDRVFLSTF 1115
RLHGFP SIVSD R+F+S F
Sbjct: 748 RLHGFPASIVSDXXRLFMSLF 810
Score = 46.2 bits (108), Expect(4) = 1e-73
Identities = 20/34 (58%), Positives = 26/34 (75%)
Frame = +2
Query: 841 RHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKW 874
RHY +G KF+I T+ RS +FL +QR+M EEQ KW
Sbjct: 53 RHYPVGKKFIIRTN*RSSKFLNEQRLMSEEQFKW 154
Score = 25.0 bits (53), Expect(4) = 1e-73
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +3
Query: 891 IENKAADALSRKLQFSAISSVQ 912
+ N ++ LSR+ FSAIS VQ
Sbjct: 135 VRNSSSGLLSRQFSFSAISMVQ 200
Score = 21.9 bits (45), Expect(4) = 1e-73
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = +3
Query: 825 VYERELMAVVLAV 837
+YERELM VVL V
Sbjct: 6 MYERELMDVVLPV 44
>TC211627
Length = 1034
Score = 116 bits (290), Expect(2) = 7e-49
Identities = 71/213 (33%), Positives = 119/213 (55%), Gaps = 10/213 (4%)
Frame = +3
Query: 1190 LYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQY 1249
+YG+ PP + + ++V+ V+ + I L L+K Q+ M++ A+ HRRD+ +
Sbjct: 243 MYGKPPPALPLYSAGTSTVEAVDAILHSLATIHHTLTCRLQKYQDSMKRIADSHRRDLTF 422
Query: 1250 EVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFH 1309
+GD VY+++ PY+ S+ + + KLS R+YGPY I A++ AY+LQLP S++HP+FH
Sbjct: 423 NIGDWVYVRL*PYRQTSI-QSTYTKLSKRFYGPYQIQARVGQVAYRLQLPPTSKIHPIFH 599
Query: 1310 ISLLKKAVNAGVQSQPLPAAL-------TEEWELKVEPEAIMDTRENRDGD---LEVLIR 1359
+SLLK V P+P L T L V+P +D + + +VL++
Sbjct: 600 VSLLK------VHHGPIPPELLALPPFSTTNHPL-VQPLQFLDWKMDESTTPPIPQVLVQ 758
Query: 1360 WKDLPTFEDSWEDFSKLLDQFPNHQLEDKLNLQ 1392
W +L + +WE +++L D + LEDK+ Q
Sbjct: 759 WTNLAPEDTTWESWTQLKDIY---DLEDKVCFQ 848
Score = 98.2 bits (243), Expect(2) = 7e-49
Identities = 44/79 (55%), Positives = 58/79 (72%)
Frame = +2
Query: 1110 VFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSW 1169
+F+S W E+F ++GTKL+FS+AYHPQTDGQTEV+NR +E YLR P+ W K+LS
Sbjct: 2 IFISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSL 181
Query: 1170 AEFWYNTNYHSAIKTTPFK 1188
AE YNT+ HS I +PF+
Sbjct: 182 AE*CYNTSVHSGIGFSPFE 238
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 118 bits (296), Expect(2) = 1e-43
Identities = 57/109 (52%), Positives = 78/109 (71%)
Frame = +1
Query: 1208 VDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSL 1267
++EV KL R+ +L L+ NL K+ + M NK RRD++Y VGD V+LK+QPY+ +SL
Sbjct: 13 LEEVNKLIIARDGLLATLRENLLKS*DIM*ANTNKRRRDIEYVVGD*VFLKMQPYRRRSL 192
Query: 1268 AKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHISLLKKA 1316
AKR N+KLSPR+Y P+ + K+ AYKL LP ++HPVFH+SLLKKA
Sbjct: 193 AKRINEKLSPRFYAPFQVFNKVGTIAYKLDLPSHIKIHPVFHVSLLKKA 339
Score = 78.2 bits (191), Expect(2) = 1e-43
Identities = 34/72 (47%), Positives = 55/72 (76%)
Frame = +3
Query: 1322 QSQPLPAALTEEWELKVEPEAIMDTRENRDGDLEVLIRWKDLPTFEDSWEDFSKLLDQFP 1381
QSQPLP L+E+W+L+ ++++D+RE + G+++VLI+WK+LP E+SWE +KL + F
Sbjct: 339 QSQPLPPMLSEDWKLQTYSDSVLDSRELQPGNVKVLIQWKNLPPSENSWESVAKLQEIFS 518
Query: 1382 NHQLEDKLNLQG 1393
+ LEDK++L G
Sbjct: 519 IYHLEDKVSLLG 554
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 164 bits (416), Expect = 2e-40
Identities = 95/254 (37%), Positives = 137/254 (53%), Gaps = 19/254 (7%)
Frame = +1
Query: 508 IEKLVKEMLNSGIIRH-STSPFSSPAILVKKKDG------------------GWRFCVDY 548
+ K V ++L G+I S S + SP ++V KK+G W+ C+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 549 RALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGH 608
R LN+AT D FP+P +D++L+ + + LD GY+QI + +D K AF G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 609 YEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQV 668
+ Y +PFGL NAP+TFQ M + + K + VF DD ++ + E L +VLQ
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 669 LKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGL 728
E NLV N +KC F E I LGH IS G+ D +KI + P P VKG+R FLG
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 729 TGYYRRFVKNYSKL 742
+YRRF+K+++K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>BI425021
Length = 426
Score = 159 bits (402), Expect = 8e-39
Identities = 74/142 (52%), Positives = 99/142 (69%)
Frame = -1
Query: 1030 LQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVF 1089
L PLP+P + W D+SMDFI GLP G TI VVV+RF+K H L + A +A +F
Sbjct: 426 LCPLPVPQRPWEDLSMDFIVGLPPYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASLF 247
Query: 1090 IKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVE 1149
+ V++LHGFP SIVSDRD +F+S FW ++F+L+GT L+ SSAYHPQTDGQTEV+NR +E
Sbjct: 246 LNIVIKLHGFPRSIVSDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIE 67
Query: 1150 TYLRCVTGSKPKQWPKWLSWAE 1171
YLR +P+ +++ W E
Sbjct: 66 QYLRAFVHGRPRNLGRFIPWVE 1
>BQ299538
Length = 426
Score = 108 bits (269), Expect(2) = 4e-36
Identities = 49/99 (49%), Positives = 68/99 (68%)
Frame = +3
Query: 1121 KLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHS 1180
KL GT LK S++YHP DGQT VN C+ET+LRC +PK +WLSWAE+WYNTN+H+
Sbjct: 135 KLPGTYLKMSTSYHP*IDGQT--VNHCLETFLRCFVADQPKM*VQWLSWAEYWYNTNFHA 308
Query: 1181 AIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERN 1219
+ TTPF+ +YGR+PPV+ + V+ V + +R+
Sbjct: 309 STGTTPFEVVYGRKPPVLNRFLPGEVRVEAVRRELQDRD 425
Score = 63.5 bits (153), Expect(2) = 4e-36
Identities = 25/44 (56%), Positives = 35/44 (78%)
Frame = +1
Query: 1076 LSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEM 1119
L HPY+A+ +AE+F KEVV LHG P S++SD D +F+S+FW E+
Sbjct: 1 LKHPYSARVLAEIFTKEVVHLHGVPASVLSDEDPIFVSSFWKEL 132
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 150 bits (378), Expect = 5e-36
Identities = 91/254 (35%), Positives = 134/254 (51%), Gaps = 19/254 (7%)
Frame = +1
Query: 508 IEKLVKEMLNSGIIRH-STSPFSSPAILVKKKDG------------------GWRFCVDY 548
+ K V ++L +G+I S S + SP +V KK G WR C+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 549 RALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGH 608
R LN+AT D +P+P +D++L + + LD SGY+QI + +D KTAF
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 609 YEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQV 668
+ Y +PFGL NA +TFQ M + + K + VF DD + + +L VLQ
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 669 LKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGL 728
+++NLV N +KC F E I LGH IS+ G+ K+ + P P VKG+ FLG
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 729 TGYYRRFVKNYSKL 742
G+YRRF+K+++K+
Sbjct: 721 VGFYRRFIKDFTKV 762
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 147 bits (372), Expect = 2e-35
Identities = 74/163 (45%), Positives = 103/163 (62%), Gaps = 1/163 (0%)
Frame = +1
Query: 714 PIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN-SFQWTEGATQAFVKLKEVM 772
P K V +R F GL +YRRFV N+S +A PLN+L+KKN +F W E QAF LKE +
Sbjct: 97 PTLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKL 276
Query: 773 TTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMA 832
T PVL P+F K F LE DASG G+ AVL+Q G P+AY S+ L Y++EL A
Sbjct: 277 TKAPVLALPDFSKTFELECDASGVGVRAVLLQGGHPIAYFSEKLHSATLNYPTYDKELYA 456
Query: 833 VVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWM 875
++ A Q W H+L+ +FVIH+D +SL+++ + + + KW+
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWV 585
Score = 33.1 bits (74), Expect = 0.88
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = +2
Query: 688 IIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRG 724
I + G V+ + GV DP KIK + +WP P +V + G
Sbjct: 17 IFFSGFVVGRNGVQMDPEKIKAIQEWPPP*KVWEILG 127
>AW830191
Length = 372
Score = 147 bits (370), Expect = 4e-35
Identities = 66/123 (53%), Positives = 96/123 (77%)
Frame = +3
Query: 1152 LRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREPPVIFKGNDSLTSVDEV 1211
LRC+TG+KP+QWPK LSWAEFW+NTNY++++K TPFK LYG +PP + KG ++ +EV
Sbjct: 6 LRCLTGTKPQQWPKRLSWAEFWFNTNYNNSLKLTPFKVLYGCDPPHLLKGAIISSTAEEV 185
Query: 1212 EKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRS 1271
+T +R+ +L +LK NL KAQN+M+ A++ RR + VGD VYLK+QPY+ +SLA+++
Sbjct: 186 NVMTNDRDQMLHDLKGNLAKAQNQMK-YADRSRRSIPLNVGDWVYLKLQPYRQRSLARKT 362
Query: 1272 NQK 1274
N+K
Sbjct: 363 NEK 371
>AW570005
Length = 413
Score = 141 bits (355), Expect = 2e-33
Identities = 70/137 (51%), Positives = 88/137 (64%)
Frame = -2
Query: 714 PIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQAFVKLKEVMT 773
P P+ + LRGFL LTG+YRRF+K Y+ +A PL+ LL K+SF W+ A AF LK V+T
Sbjct: 412 PPPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDSFVWSPEADVAFQALKNVVT 233
Query: 774 TVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAV 833
VL P+F KPF +ETDASG +GAVL QEG P+A+ SK + S Y EL A+
Sbjct: 232 NTLVLALPDFTKPFTVETDASGSDMGAVLSQEGHPIAFFSKEFCPKLVRSSTYVHELAAI 53
Query: 834 VLAVQKWRHYLLGSKFV 850
V+KWR YLLG V
Sbjct: 52 TNVVKKWRQYLLGHHLV 2
>CF922488
Length = 741
Score = 140 bits (353), Expect = 4e-33
Identities = 89/246 (36%), Positives = 132/246 (53%), Gaps = 1/246 (0%)
Frame = +3
Query: 535 VKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKE 594
V K+DG CVDYR LN A+ DKFP+P I+ L+D + FS +D SGY+QI++
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 595 EDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKN 654
ED+ KT F T G + Y + FGL N +T+Q M + + K + V+ DD+++ S+
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 655 EELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWP 714
EE H +LR + + L++ L N KC F L + S G+ D +K+K +L+
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 715 IPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSF-QWTEGATQAFVKLKEVMT 773
P K ++GFLG Y RF+ +PL LL KN F +W AF ++K+ +
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 774 TVPVLV 779
VLV
Sbjct: 723 NPHVLV 740
>CA820403 weakly similar to GP|13273463|gb| pol protein integrase region
{Ginkgo biloba}, partial (52%)
Length = 421
Score = 108 bits (269), Expect(2) = 5e-33
Identities = 52/77 (67%), Positives = 62/77 (79%), Gaps = 1/77 (1%)
Frame = -3
Query: 1089 FIKEVVRLHGFPTSIVSDRDRVFL-STFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRC 1147
FIKE V+LHG +SIVSD DR+FL S FW+E+FK+ GTKLKFS AYHPQ D T+VVNRC
Sbjct: 335 FIKEAVKLHGCSSSIVSDWDRLFLIS*FWTELFKMEGTKLKFSLAYHPQPDSHTKVVNRC 156
Query: 1148 VETYLRCVTGSKPKQWP 1164
+E L+C+T SK KQWP
Sbjct: 155 IEMNLQCLTTSKRKQWP 105
Score = 53.1 bits (126), Expect(2) = 5e-33
Identities = 22/34 (64%), Positives = 30/34 (87%)
Frame = -1
Query: 1060 ILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEV 1093
I+V+V RFTKYAHF+ LSHPY AKE++EV +K++
Sbjct: 421 IMVIVYRFTKYAHFVVLSHPYLAKEVSEVLLKKL 320
>BI317507
Length = 359
Score = 124 bits (311), Expect = 3e-28
Identities = 62/114 (54%), Positives = 81/114 (70%), Gaps = 1/114 (0%)
Frame = -1
Query: 645 FDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADP 704
F +ILIYS + + H HL VL VLK+ LVAN+KKC F Q I YLGHVIS+ VA D
Sbjct: 359 FYNILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDS 180
Query: 705 SKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLA-QPLNQLLKKNSFQW 757
+K+K +++WP+PK VK + FL LTGYYR+F+K+Y KLA +PL L K + F+W
Sbjct: 179 NKVKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLTKNDGFKW 18
>CD487724
Length = 676
Score = 120 bits (301), Expect = 4e-27
Identities = 67/216 (31%), Positives = 117/216 (54%), Gaps = 2/216 (0%)
Frame = +3
Query: 303 VLILIDCGATSNFISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEVQGISI 362
++ L+D G+T NF+ Q LV +L +P +T V VGNG + + +C+ + + +Q I
Sbjct: 24 LVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMVGNGHHLKCTTICEAIPISIQNIEF 203
Query: 363 MQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCRVTANW 422
+ H ++L + G +VLG+ WL +LG I ++ L +Q+ Q + + L+GE N
Sbjct: 204 LVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEAQLGLLNH 383
Query: 423 KSIKITEQ--QEAEGYYLSYEYQKEEEKTEAEVPEGMRKILEEYPEVFQEPKGLPPRRTT 480
++ Q + ++++ + + +P+ ++ +L+++ +FQ P+GLPP R T
Sbjct: 384 HQLRRLHQTHEPVTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQGLPPARET 563
Query: 481 DHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEML 516
DH I L + N+R Y YP Y NEIE V ML
Sbjct: 564 DHHIHLLP*SEPVNMRLY*YPHY*NNEIEHQVNLML 671
>BQ627806
Length = 435
Score = 75.9 bits (185), Expect(2) = 2e-26
Identities = 36/83 (43%), Positives = 52/83 (62%)
Frame = +1
Query: 808 PVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIM 867
P+ + + S Y REL A+ +AV+KWR YLLG FVI TD RSL+ L Q +
Sbjct: 181 PLPFSFMPFCSKLLRASTYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQ 360
Query: 868 GEEQQKWMSKLMGYDFEIKYKPG 890
EQQ ++++LMG+D+ I+Y+ G
Sbjct: 361 TPEQQIYLARLMGFDYTIQYRAG 429
Score = 63.5 bits (153), Expect(2) = 2e-26
Identities = 30/63 (47%), Positives = 42/63 (66%)
Frame = +3
Query: 749 LLKKNSFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRP 808
LL K+ F W E A +AF +LK + PVL P+F+ F++ETDASG G+GA+L Q P
Sbjct: 3 LLVKDQFHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHHP 182
Query: 809 VAY 811
+A+
Sbjct: 183 LAF 191
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 115 bits (288), Expect = 1e-25
Identities = 56/98 (57%), Positives = 71/98 (72%)
Frame = +3
Query: 1236 MRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYK 1295
M+ A++ RR V VGD VYLK+QPY+LKSLAK+ N+KLSPR+YGPY I +I A++
Sbjct: 3 MKAYADRSRRAVTLSVGDWVYLKLQPYRLKSLAKKRNEKLSPRFYGPYQIKKQIGLVAFE 182
Query: 1296 LQLPEGSQVHPVFHISLLKKAVNAGVQSQPLPAALTEE 1333
L LP ++HPVFH SLLKKAV A QPLP L+E+
Sbjct: 183 LDLPPARKIHPVFHASLLKKAVAATANPQPLPLMLSED 296
Score = 56.6 bits (135), Expect = 7e-08
Identities = 28/71 (39%), Positives = 41/71 (57%)
Frame = +2
Query: 1323 SQPLPAALTEEWELKVEPEAIMDTRENRDGDLEVLIRWKDLPTFEDSWEDFSKLLDQFPN 1382
S P+ + +EL+V P + N +G EVLI+ +DLP FE +WE + +QFP+
Sbjct: 263 SSTTPSDVI*RFELRVFPAEVKAVHNNSNGIAEVLIQLEDLPDFEATWESVEVIKEQFPS 442
Query: 1383 HQLEDKLNLQG 1393
LEDK+ L G
Sbjct: 443 FHLEDKVTLLG 475
>BI317638 weakly similar to GP|9294238|dbj| contains similarity to reverse
transcriptase~gene_id:K11J14.5 {Arabidopsis thaliana},
partial (5%)
Length = 420
Score = 92.8 bits (229), Expect(2) = 4e-22
Identities = 49/117 (41%), Positives = 64/117 (53%)
Frame = -2
Query: 1004 LDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVV 1063
L+ + C CQ KYE L PL +P + W D+S+DFI GL ILVV
Sbjct: 353 LECAQMLPNCLDCQHTKYETKRIVDLLCPLLVPHRPWEDLSLDFITGLLPYHVHTAILVV 174
Query: 1064 VDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMF 1120
VD F+K H L + A +A +FI V +LHG P S+VSD D +F+S FW E+F
Sbjct: 173 VDHFSKGIHLGMLPSSHTAHTVACLFIDSVAKLHGLPRSLVSDCDLLFVSHFWQELF 3
Score = 32.0 bits (71), Expect(2) = 4e-22
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = -1
Query: 978 TALGGHAGIFRTYKRISALFYWEGMKLDI 1006
T GGH GI +T +S YW GM+ D+
Sbjct: 420 TTTGGHTGIAKTLA*LSKNIYWFGMRTDV 334
>BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 424
Score = 102 bits (253), Expect = 2e-21
Identities = 53/133 (39%), Positives = 84/133 (62%), Gaps = 2/133 (1%)
Frame = +1
Query: 1229 LEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAK 1288
LE+AQ+ M + AN HRR VGD VYLKI+P++ S+ R + KL+ R+YGPY ++ +
Sbjct: 25 LERAQSLMVKHANNHRRPHDINVGDWVYLKIRPHRQGSMPPRLHPKLTARFYGPYLVMRQ 204
Query: 1289 INPAAYKLQLPEGSQVHPVFHISLLKKAVNAGVQSQPLPAALTEEWEL--KVEPEAIMDT 1346
+ A++LQLP +++HPVFH+S LK+A+ + LP L + EL V+ I +
Sbjct: 205 VGAVAFQLQLPSEARIHPVFHVSQLKRALGNHQAQEELPPDLEHQAELYFPVQILKIREV 384
Query: 1347 RENRDGDLEVLIR 1359
++ + + +VLIR
Sbjct: 385 QKQHEVERQVLIR 423
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 100 bits (249), Expect = 4e-21
Identities = 54/142 (38%), Positives = 84/142 (59%)
Frame = +3
Query: 543 RFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAF 602
R CVDYR LN+A+ D FP+P ID L+ + + +FS +D SGY+QI+M ED+ KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 603 RTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHL 662
T G + Y V+ FGL N +T+ M + + + K + + D+++ S+ EE H +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 663 RIVLQVLKENNLVANQKKCSFG 684
+ + L++ L N +KC FG
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFG 428
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 57,431,894
Number of Sequences: 63676
Number of extensions: 762284
Number of successful extensions: 4492
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 4317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4449
length of query: 1393
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1284
effective length of database: 5,698,948
effective search space: 7317449232
effective search space used: 7317449232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0029a.5


