

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
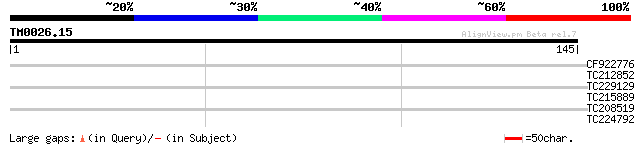
Query= TM0026.15
(145 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CF922776 28 2.0
TC212852 homologue to UP|Q9SXF7 (Q9SXF7) Amino acid transporter-... 27 3.4
TC229129 similar to UP|Q6NM52 (Q6NM52) At4g15770, complete 27 4.5
TC215889 homologue to PIR|T09261|T09261 JUN kinase-activation-do... 26 5.9
TC208519 similar to UP|Q99NH6 (Q99NH6) BZIP protein ATF7, partia... 26 5.9
TC224792 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich gly... 26 7.7
>CF922776
Length = 615
Score = 27.7 bits (60), Expect = 2.0
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = -3
Query: 121 LEGELWNWEDPPDDELRARSFV 142
+ G LWNW +P EL + FV
Sbjct: 79 INGALWNWAEPNSGELFSSKFV 14
>TC212852 homologue to UP|Q9SXF7 (Q9SXF7) Amino acid transporter-like protein
1, partial (19%)
Length = 672
Score = 26.9 bits (58), Expect = 3.4
Identities = 21/85 (24%), Positives = 35/85 (40%), Gaps = 5/85 (5%)
Frame = -1
Query: 47 RGRWCETYLMNG-----RVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENE 101
R + C Y++ RV LPY+ +F PI+ S L K +P D + + +
Sbjct: 405 RKKTCTNYIIRPLGNA*RVALPYSACAFAWPIIKNSIGLEEFKFKPIIDNAPNTSNGKGQ 226
Query: 102 HYVDSKLMSKPMEEEELPPLEGELW 126
+ + P+E P+E W
Sbjct: 225 AKSNPQPTQNPIE----VPIEAIFW 163
>TC229129 similar to UP|Q6NM52 (Q6NM52) At4g15770, complete
Length = 661
Score = 26.6 bits (57), Expect = 4.5
Identities = 7/35 (20%), Positives = 18/35 (51%)
Frame = -1
Query: 24 WVNDEHQVRRIYVGSAAVRSFDQRGRWCETYLMNG 58
W+ ++ R+++ S+++ + WC T + G
Sbjct: 628 WIKTNYKTRKMHQSSSSILKYSPTSAWCRTTIPFG 524
>TC215889 homologue to PIR|T09261|T09261 JUN kinase-activation-domain-binding
protein homolog - alfalfa {Medicago sativa;} , partial
(98%)
Length = 1420
Score = 26.2 bits (56), Expect = 5.9
Identities = 8/22 (36%), Positives = 14/22 (63%)
Frame = +1
Query: 20 RDRAWVNDEHQVRRIYVGSAAV 41
RD+ W ND H +R+ + + A+
Sbjct: 211 RDKPWANDPHYFKRVKISALAL 276
>TC208519 similar to UP|Q99NH6 (Q99NH6) BZIP protein ATF7, partial (6%)
Length = 783
Score = 26.2 bits (56), Expect = 5.9
Identities = 16/87 (18%), Positives = 36/87 (40%), Gaps = 8/87 (9%)
Frame = -3
Query: 50 WCETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEP--------EKDPLELVPDDENE 101
WC+T + + +PY + ++ ++ ++N K P D L+P +
Sbjct: 748 WCQTLITGTKFSVPYHT*NLDIHNYNNNNNIINHKILPLCICV*NDLNDTSLLLPSSQAN 569
Query: 102 HYVDSKLMSKPMEEEELPPLEGELWNW 128
H ++ +++KP + + W W
Sbjct: 568 HQ-NASVLNKPTKSGNNMVAKLRRWRW 491
>TC224792 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich glycoprotein
(Fragment), partial (86%)
Length = 688
Score = 25.8 bits (55), Expect = 7.7
Identities = 16/60 (26%), Positives = 25/60 (41%), Gaps = 5/60 (8%)
Frame = -3
Query: 6 RTGHSWEIILHDEDRDRAWVN-----DEHQVRRIYVGSAAVRSFDQRGRWCETYLMNGRV 60
R WE++ D D + V +E + + VGS + R ++RG E N V
Sbjct: 359 REDEEWEMVEEDFDSSKEVVENGSSREEEEREMVVVGSGSSRGEEERGMVVEEIYSNREV 180
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,414,890
Number of Sequences: 63676
Number of extensions: 101857
Number of successful extensions: 471
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 468
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 471
length of query: 145
length of database: 12,639,632
effective HSP length: 88
effective length of query: 57
effective length of database: 7,036,144
effective search space: 401060208
effective search space used: 401060208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0026.15


