

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.14
(597 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
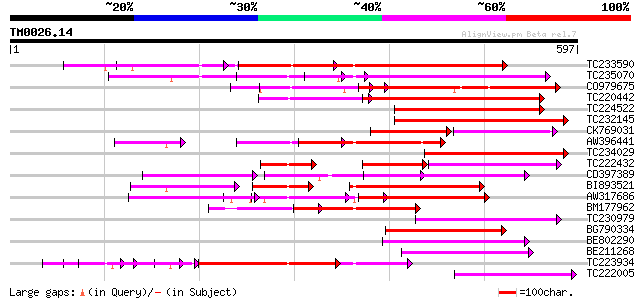
Score E
Sequences producing significant alignments: (bits) Value
TC233590 similar to UP|Q6K9T2 (Q6K9T2) Pentatricopeptide repeat-... 205 5e-53
TC235070 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding p... 193 2e-49
CO979675 171 8e-43
TC220442 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding p... 169 2e-42
TC224522 weakly similar to UP|Q9LUL5 (Q9LUL5) Emb|CAB66100.1, pa... 157 2e-38
TC232145 155 4e-38
CK769031 86 2e-32
AW396441 133 2e-31
TC234029 similar to UP|Q9LN01 (Q9LN01) T6D22.15, partial (32%) 133 2e-31
TC222432 91 1e-30
CD397389 similar to GP|6691205|gb| F1K23.11 {Arabidopsis thalian... 128 6e-30
BI893521 127 1e-29
AW317686 124 9e-29
BM177962 119 5e-27
TC230979 weakly similar to UP|Q6Z8F8 (Q6Z8F8) Selenium-binding p... 116 3e-26
BG790334 weakly similar to GP|2160154|gb| F19K23.18 gene product... 111 1e-24
BE802290 111 1e-24
BE211268 109 3e-24
TC223934 109 3e-24
TC222005 weakly similar to UP|Q9FIB2 (Q9FIB2) Selenium-binding p... 108 5e-24
>TC233590 similar to UP|Q6K9T2 (Q6K9T2) Pentatricopeptide repeat-containing
protein-like, partial (5%)
Length = 1094
Score = 205 bits (521), Expect = 5e-53
Identities = 104/283 (36%), Positives = 172/283 (60%)
Frame = +2
Query: 242 LTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPN 301
++G+ G++ +LF+++P+ D +WTS++ GYA++G+ EAL +F +M + ++P+
Sbjct: 2 VSGYAVWGDMESGAELFSQMPKSDSCSWTSLIRGYARNGMGYEALGVFKQMIKHQ-VRPD 178
Query: 302 NGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFD 361
T T L AC+ +ASL G+QIH + + NT VV A++NMYSKCG L AR++F+
Sbjct: 179 QFTLSTCLFACATIASLKHGRQIHAFLVLNNIKPNTIVVCAIVNMYSKCGSLETARRVFN 358
Query: 362 DGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLV 421
+ ++D++ WN MI A AH+GYG EAI + M ++G + N T+V +L AC H+GLV
Sbjct: 359 F-IGNKQDVVLWNTMILALAHYGYGIEAIMMLYNMLKIGVKPNKGTFVGILNACCHSGLV 535
Query: 422 DEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLA 481
EG+Q F + + ++HY L +L G+A E+ ++ + K V +
Sbjct: 536 QEGLQLFKSMTSEHGVVPDQEHYTRLANLLGQARCFNESVKDLQMMDCKPGDHVCNSSIG 715
Query: 482 GCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWK 524
C +HGN D G VA ++K++ +++ Y LLS YA++GKW+
Sbjct: 716 VCRMHGNIDHGAEVAAFLIKLQPQSSAAYELLSRTYAALGKWE 844
Score = 75.9 bits (185), Expect = 5e-14
Identities = 62/236 (26%), Positives = 100/236 (42%), Gaps = 2/236 (0%)
Frame = +2
Query: 113 VNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWN 172
V+GY +E LF +MP+ + SW +I GY +NG +AL +F++M K +
Sbjct: 2 VSGYAVWGDMESGAELFSQMPKSDSCSWTSLIRGYARNGMGYEALGVFKQMIKHQV---- 169
Query: 173 IIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPE 232
R L + A I R++ L L P
Sbjct: 170 ----------------------RPDQFTLSTCLFACATIASLKHGRQIHAFLVLNNIKPN 283
Query: 233 RDMASWNAMLTGFFQNGELNRAEKLFAELPQK-DVITWTSMMTGYAQHGLSEEALKMFTK 291
+ A++ + + G L A ++F + K DV+ W +M+ A +G EA+ M
Sbjct: 284 TIVVC--AIVNMYSKCGSLETARRVFNFIGNKQDVVLWNTMILALAHYGYGIEAIMMLYN 457
Query: 292 MQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQ-LISKTGFQENTRVVSALINM 346
M G+KPN GTFV +L AC + EG Q+ + + S+ G + + L N+
Sbjct: 458 M-LKIGVKPNKGTFVGILNACCHSGLVQEGLQLFKSMTSEHGVVPDQEHYTRLANL 622
Score = 46.6 bits (109), Expect = 3e-05
Identities = 46/183 (25%), Positives = 81/183 (44%), Gaps = 9/183 (4%)
Frame = +2
Query: 57 GRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ---PDAKKSVVTWTTMV 113
G + +LF QMP+ D WT++I GY + G EA +F Q + T +T +
Sbjct: 23 GDMESGAELFSQMPKSDSCSWTSLIRGYARNGMGYEALGVFKQMIKHQVRPDQFTLSTCL 202
Query: 114 NGYVKLNQIEEAES----LFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM-PKRNL 168
+ ++ L + N + ++ Y + G +E A +F + K+++
Sbjct: 203 FACATIASLKHGRQIHAFLVLNNIKPNTIVVCAIVNMYSKCGSLETARRVFNFIGNKQDV 382
Query: 169 VSWNIIIKALARCGRIEDAQWHLIRCRK-GMMPLRNVVSWNAMITGYAQNRRLDEALELF 227
V WN +I ALA G +A L K G+ P N ++ ++ + + E L+LF
Sbjct: 383 VLWNTMILALAHYGYGIEAIMMLYNMLKIGVKP--NKGTFVGILNACCHSGLVQEGLQLF 556
Query: 228 ERM 230
+ M
Sbjct: 557 KSM 565
Score = 40.8 bits (94), Expect = 0.002
Identities = 28/113 (24%), Positives = 57/113 (49%), Gaps = 4/113 (3%)
Frame = +2
Query: 55 KEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVN 114
K GR HA + + + + + + +++ Y KCG+++ AR++F+ K+ VV W TM+
Sbjct: 230 KHGRQIHAFLVLNNI-KPNTIVVCAIVNMYSKCGSLETARRVFNFIGNKQDVVLWNTMIL 406
Query: 115 GYVKLNQIEEAESLFYEM----PERNELSWNIMIGGYGQNGQIEKALDLFRRM 163
EA + Y M + N+ ++ ++ +G +++ L LF+ M
Sbjct: 407 ALAHYGYGIEAIMMLYNMLKIGVKPNKGTFVGILNACCHSGLVQEGLQLFKSM 565
>TC235070 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding protein-like,
partial (22%)
Length = 875
Score = 193 bits (490), Expect = 2e-49
Identities = 98/259 (37%), Positives = 154/259 (58%)
Frame = +2
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDL 370
ACS L S +GQ +H + KT FQ N V +AL++ YSKCG L A++ F + ++
Sbjct: 2 ACSCLCSFRQGQLLHAHLIKTPFQVNVYVGTALVDFYSKCGHLAEAQRSFIS--IFSPNV 175
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDK 430
+W +I YA+HG G+EAI LF M G N T+V +L+AC+HAGLV EG++ F
Sbjct: 176 AAWTALINGYAYHGLGSEAILLFRSMLHQGIVPNAATFVGVLSACNHAGLVCEGLRIFHS 355
Query: 431 LLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNAD 490
+ + + +HY C+VDL GR+G LKEA I + ++ +WG LL + +
Sbjct: 356 MQRCYGVTPTIEHYTCVVDLLGRSGHLKEAEEFIIKMPIEADGIIWGALLNASWFWKDME 535
Query: 491 IGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVG 550
+G+ A+K+ ++ + +LSNMYA +G+W + +R +++ L+K PGCSWIE+
Sbjct: 536 VGERAAEKLFSLDPNPIFAFVVLSNMYAILGRWGQKTKLRKRLQSLELRKDPGCSWIELN 715
Query: 551 NTVQVFVVGDKSHSQSEML 569
N + +F V DK+H S+++
Sbjct: 716 NKIHLFSVEDKTHLYSDVI 772
Score = 63.9 bits (154), Expect = 2e-10
Identities = 41/141 (29%), Positives = 71/141 (50%), Gaps = 1/141 (0%)
Frame = +2
Query: 240 AMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLK 299
A++ + + G L A++ F + +V WT+++ GYA HGL EA+ +F M + G+
Sbjct: 95 ALVDFYSKCGHLAEAQRSFISIFSPNVAAWTALINGYAYHGLGSEAILLFRSM-LHQGIV 271
Query: 300 PNNGTFVTVLGACSGLASLTEGQQI-HQLISKTGFQENTRVVSALINMYSKCGELHIARK 358
PN TFV VL AC+ + EG +I H + G + ++++ + G L A +
Sbjct: 272 PNAATFVGVLSACNHAGLVCEGLRIFHSMQRCYGVTPTIEHYTCVVDLLGRSGHLKEAEE 451
Query: 359 IFDDGLLRQRDLISWNGMIAA 379
F + + D I W ++ A
Sbjct: 452 -FIIKMPIEADGIIWGALLNA 511
Score = 51.6 bits (122), Expect = 1e-06
Identities = 61/264 (23%), Positives = 118/264 (44%), Gaps = 14/264 (5%)
Frame = +2
Query: 105 SVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMP 164
+V T +V+ Y K + EA+ F + N +W +I GY +G +A+ LFR M
Sbjct: 77 NVYVGTALVDFYSKCGHLAEAQRSFISIFSPNVAAWTALINGYAYHGLGSEAILLFRSML 256
Query: 165 KRNLV----SWNIIIKALARCGRIEDAQ--WHLIRCRKGMMPLRNVVSWNAMITGYAQNR 218
+ +V ++ ++ A G + + +H ++ G+ P + + ++ ++
Sbjct: 257 HQGIVPNAATFVGVLSACNHAGLVCEGLRIFHSMQRCYGVTP--TIEHYTCVVDLLGRSG 430
Query: 219 RLDEALELFERMP-ERDMASWNAMLTG--FFQNGELNR--AEKLFAELPQKDVITWTSMM 273
L EA E +MP E D W A+L F+++ E+ AEKLF+ L + + +
Sbjct: 431 HLKEAEEFIIKMPIEADGIIWGALLNASWFWKDMEVGERAAEKLFS-LDPNPIFAFVVLS 607
Query: 274 TGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGF 333
YA G + K+ ++Q+ L+ + G L L S+ + ++ +
Sbjct: 608 NMYAILGRWGQKTKLRKRLQSL-ELRKDPGCSWIELNNKIHLFSVEDKTHLYSDVIYATV 784
Query: 334 QENTRVVSALI---NMYSKCGELH 354
+ T ++++I +YS GE H
Sbjct: 785 EHITATINSIIPSNYLYSSHGE*H 856
>CO979675
Length = 732
Score = 171 bits (433), Expect = 8e-43
Identities = 84/219 (38%), Positives = 137/219 (62%), Gaps = 6/219 (2%)
Frame = -2
Query: 368 RDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQY 427
RD++ WN MI+ A HG G A+ +F++M++ G + +D+T++ + TACS++G+ EG+Q
Sbjct: 731 RDIVCWNAMISGLAMHGDGASALKMFSEMEKTGIKPDDITFIAVFTACSYSGMAHEGLQL 552
Query: 428 FDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGL------GVKLSLSVWGPLLA 481
DK+ I+ K +HY CLVDL RAG EA +I + G + +L+ W L+
Sbjct: 551 LDKMSSLYEIEPKSEHYGCLVDLLSRAGLFGEAMVMIRRITSTSWNGSEETLA-WRAFLS 375
Query: 482 GCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQ 541
C HG A + + AK++L++E+ ++G Y LLSN+YA+ GK +A VR M++KG+ K
Sbjct: 374 ACCNHGQAQLAERAAKRLLRLEN-HSGVYVLLSNLYAASGKHSDARRVRNMMRNKGVDKA 198
Query: 542 PGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKM 580
PGCS +E+ V F+ G+++H Q E + +L LH ++
Sbjct: 197 PGCSSVEIDGVVSEFIAGEETHPQMEEIHSVLEILHMQL 81
Score = 63.2 bits (152), Expect = 3e-10
Identities = 35/126 (27%), Positives = 66/126 (51%), Gaps = 5/126 (3%)
Frame = -2
Query: 264 KDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEG-Q 322
+D++ W +M++G A HG ALKMF++M+ G+KP++ TF+ V ACS EG Q
Sbjct: 731 RDIVCWNAMISGLAMHGDGASALKMFSEMEKT-GIKPDDITFIAVFTACSYSGMAHEGLQ 555
Query: 323 QIHQLISKTGFQENTRVVSALINMYSKCG----ELHIARKIFDDGLLRQRDLISWNGMIA 378
+ ++ S + + L+++ S+ G + + R+I + ++W ++
Sbjct: 554 LLDKMSSLYEIEPKSEHYGCLVDLLSRAGLFGEAMVMIRRITSTSWNGSEETLAWRAFLS 375
Query: 379 AYAHHG 384
A +HG
Sbjct: 374 ACCNHG 357
Score = 47.8 bits (112), Expect = 1e-05
Identities = 39/174 (22%), Positives = 79/174 (44%), Gaps = 6/174 (3%)
Frame = -2
Query: 233 RDMASWNAMLTGFFQNGELNRAEKLFAELP----QKDVITWTSMMTGYAQHGLSEEALKM 288
RD+ WNAM++G +G+ A K+F+E+ + D IT+ ++ T + G++ E L++
Sbjct: 731 RDIVCWNAMISGLAMHGDGASALKMFSEMEKTGIKPDDITFIAVFTACSYSGMAHEGLQL 552
Query: 289 FTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGF--QENTRVVSALINM 346
KM + ++P + + ++ S E + + I+ T + E T A ++
Sbjct: 551 LDKMSSLYEIEPKSEHYGCLVDLLSRAGLFGEAMVMIRRITSTSWNGSEETLAWRAFLSA 372
Query: 347 YSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELG 400
G+ +A + L + + + YA G ++A + N M+ G
Sbjct: 371 CCNHGQAQLAERAAKRLLRLENHSGVYVLLSNLYAASGKHSDARRVRNMMRNKG 210
>TC220442 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding protein-like,
partial (25%)
Length = 706
Score = 169 bits (429), Expect = 2e-42
Identities = 77/192 (40%), Positives = 120/192 (62%)
Frame = +1
Query: 372 SWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL 431
+W +I A HG G EA++ F +MQ+ G N +T+ +LTACSHAGL +EG F+ +
Sbjct: 16 AWTAIIGGLAIHGKGREALDWFTQMQKAGINPNSITFTAILTACSHAGLTEEGKSLFESM 195
Query: 432 LKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADI 491
+I+ +HY C+VDL GRAG LKEA IE + VK + ++WG LL C +H + ++
Sbjct: 196 SSVYNIKPSMEHYGCMVDLMGRAGLLKEAREFIESMPVKPNAAIWGALLNACQLHKHFEL 375
Query: 492 GKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGN 551
GK + K +++++ +++G Y L+++YA+ G+W + VR ++K +GL PGCS I +
Sbjct: 376 GKEIGKILIELDPDHSGRYIHLASIYAAAGEWNQVVRVRSQIKHRGLLNHPGCSSITLNG 555
Query: 552 TVQVFVVGDKSH 563
V F GD SH
Sbjct: 556 VVHEFFAGDGSH 591
Score = 57.0 bits (136), Expect = 2e-08
Identities = 35/122 (28%), Positives = 60/122 (48%), Gaps = 1/122 (0%)
Frame = +1
Query: 263 QKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQ 322
+K V WT+++ G A HG EAL FT+MQ G+ PN+ TF +L ACS EG+
Sbjct: 1 KKCVCAWTAIIGGLAIHGKGREALDWFTQMQ-KAGINPNSITFTAILTACSHAGLTEEGK 177
Query: 323 QIHQLISKT-GFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYA 381
+ + +S + + ++++ + G L AR+ + ++ I W ++ A
Sbjct: 178 SLFESMSSVYNIKPSMEHYGCMVDLMGRAGLLKEAREFIESMPVKPNAAI-WGALLNACQ 354
Query: 382 HH 383
H
Sbjct: 355 LH 360
Score = 40.8 bits (94), Expect = 0.002
Identities = 45/217 (20%), Positives = 94/217 (42%), Gaps = 9/217 (4%)
Frame = +1
Query: 139 SWNIMIGGYGQNGQIEKALDLFRRMPKR----NLVSWNIIIKALARCGRIEDAQWHLIRC 194
+W +IGG +G+ +ALD F +M K N +++ I+ A + G E+ + L
Sbjct: 16 AWTAIIGGLAIHGKGREALDWFTQMQKAGINPNSITFTAILTACSHAGLTEEGK-SLFES 192
Query: 195 RKGMMPLR-NVVSWNAMITGYAQNRRLDEALELFERMPER-DMASWNAMLTG--FFQNGE 250
+ ++ ++ + M+ + L EA E E MP + + A W A+L ++ E
Sbjct: 193 MSSVYNIKPSMEHYGCMVDLMGRAGLLKEAREFIESMPVKPNAAIWGALLNACQLHKHFE 372
Query: 251 LNR-AEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVL 309
L + K+ EL + + + YA G + +++ ++++ G L
Sbjct: 373 LGKEIGKILIELDPDHSGRYIHLASIYAAAGEWNQVVRVRSQIKHRGLLNH--------- 525
Query: 310 GACSGLASLTEGQQIHQLISKTGFQENTRVVSALINM 346
G +S+T +H+ + G + + + + N+
Sbjct: 526 ---PGCSSITLNGVVHEFFAGDGSHPHIQEIYGMPNL 627
Score = 36.2 bits (82), Expect = 0.042
Identities = 43/189 (22%), Positives = 83/189 (43%), Gaps = 15/189 (7%)
Frame = +1
Query: 204 VVSWNAMITGYAQNRRLDEALELFERMPER----DMASWNAMLTGFFQNGELNRAEKLFA 259
V +W A+I G A + + EAL+ F +M + + ++ A+LT G + LF
Sbjct: 10 VCAWTAIIGGLAIHGKGREALDWFTQMQKAGINPNSITFTAILTACSHAGLTEEGKSLFE 189
Query: 260 ELP-----QKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSG 314
+ + + + M+ + GL +EA + M +KPN + +L AC
Sbjct: 190 SMSSVYNIKPSMEHYGCMVDLMGRAGLLKEAREFIESMP----VKPNAAIWGALLNACQL 357
Query: 315 LASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGE----LHIARKIFDDGLLRQR-- 368
G++I +++ + + R + L ++Y+ GE + + +I GLL
Sbjct: 358 HKHFELGKEIGKILIELDPDHSGRYIH-LASIYAAAGEWNQVVRVRSQIKHRGLLNHPGC 534
Query: 369 DLISWNGMI 377
I+ NG++
Sbjct: 535 SSITLNGVV 561
Score = 28.5 bits (62), Expect = 8.8
Identities = 15/63 (23%), Positives = 28/63 (43%), Gaps = 5/63 (7%)
Frame = +1
Query: 57 GRTDHARKLFDQMP-----QRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTT 111
G T+ + LF+ M + + M+D + G +KEAR+ + K + W
Sbjct: 157 GLTEEGKSLFESMSSVYNIKPSMEHYGCMVDLMGRAGLLKEAREFIESMPVKPNAAIWGA 336
Query: 112 MVN 114
++N
Sbjct: 337 LLN 345
>TC224522 weakly similar to UP|Q9LUL5 (Q9LUL5) Emb|CAB66100.1, partial (18%)
Length = 534
Score = 157 bits (396), Expect = 2e-38
Identities = 70/158 (44%), Positives = 107/158 (67%)
Frame = +2
Query: 406 VTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIE 465
+T++ LL+AC AG V+E + F ++ N I + +HYACLVD+ RAG+L+ A II
Sbjct: 8 ITFLSLLSACCRAGKVNESMNLFSLMVDNYGIPPRSEHYACLVDVMSRAGQLQRACKIIN 187
Query: 466 GLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKE 525
+ K S+WG +LA C+VH N ++G+L A++IL ++ N+G Y +LSN+YA+ GKWK+
Sbjct: 188 EMPFKADSSIWGAVLAACSVHLNVELGELAARRILNLDPFNSGAYVMLSNIYAAAGKWKD 367
Query: 526 AANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSH 563
+R+ MK++G+KKQ SW+++GN FV GD SH
Sbjct: 368 VHRIRVLMKEQGVKKQTAYSWLQIGNKTHYFVGGDPSH 481
>TC232145
Length = 817
Score = 155 bits (393), Expect = 4e-38
Identities = 75/183 (40%), Positives = 115/183 (61%)
Frame = +3
Query: 406 VTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIE 465
+T+V +L+ACSH G +D+G +YF+ + +I+ +HYAC+VDL GR GRL+EA IE
Sbjct: 24 ITFVSVLSACSHTGKIDDGFKYFNSMANVHNIKPGLEHYACMVDLLGRVGRLEEACRFIE 203
Query: 466 GLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKE 525
+ + VWG LL C H N ++G+ VA+++ K+E +N G Y LLSN+Y G +E
Sbjct: 204 SMPFEPDSLVWGALLGACGKHANVEMGREVAERLFKLEPDNPGNYMLLSNIYIRHGMLEE 383
Query: 526 AANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGD 585
A VR M G++K+ GCSWI+V N VF D+SHS+++ + +L L +K+ G
Sbjct: 384 ADEVRRLMGINGVRKESGCSWIDVKNRTFVFNANDRSHSRTQEIYGMLQKLKELIKRRGY 563
Query: 586 ILD 588
+ +
Sbjct: 564 VAE 572
Score = 38.5 bits (88), Expect = 0.009
Identities = 39/144 (27%), Positives = 66/144 (45%), Gaps = 9/144 (6%)
Frame = +3
Query: 174 IIKALARCGRIEDAQWHLIRCRKGMMPLRNVVS----WNAMITGYAQNRRLDEALELFER 229
++ A + G+I+D + M + N+ + M+ + RL+EA E
Sbjct: 39 VLSACSHTGKIDDG----FKYFNSMANVHNIKPGLEHYACMVDLLGRVGRLEEACRFIES 206
Query: 230 MP-ERDMASWNAML--TGFFQNGELNR--AEKLFAELPQKDVITWTSMMTGYAQHGLSEE 284
MP E D W A+L G N E+ R AE+LF +L + + + Y +HG+ EE
Sbjct: 207 MPFEPDSLVWGALLGACGKHANVEMGREVAERLF-KLEPDNPGNYMLLSNIYIRHGMLEE 383
Query: 285 ALKMFTKMQANGGLKPNNGTFVTV 308
A ++ M NG K + +++ V
Sbjct: 384 ADEVRRLMGINGVRKESGCSWIDV 455
Score = 32.0 bits (71), Expect = 0.80
Identities = 19/88 (21%), Positives = 41/88 (46%), Gaps = 1/88 (1%)
Frame = +3
Query: 297 GLKPNNGTFVTVLGACSGLASLTEG-QQIHQLISKTGFQENTRVVSALINMYSKCGELHI 355
G+ P TFV+VL ACS + +G + + + + + + ++++ + G L
Sbjct: 6 GVVPEYITFVSVLSACSHTGKIDDGFKYFNSMANVHNIKPGLEHYACMVDLLGRVGRLEE 185
Query: 356 ARKIFDDGLLRQRDLISWNGMIAAYAHH 383
A + F + + + D + W ++ A H
Sbjct: 186 ACR-FIESMPFEPDSLVWGALLGACGKH 266
>CK769031
Length = 827
Score = 85.9 bits (211), Expect(2) = 2e-32
Identities = 46/109 (42%), Positives = 63/109 (57%)
Frame = +1
Query: 468 GVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAA 527
GV + G LL+ CN GN + + K + VE +++G Y LLSN+YA+ KW E
Sbjct: 265 GVNKDFQIPGALLSACNT*GNVGFTQEMLKSLQNVEFQDSGIYVLLSNLYATNKKWAEVR 444
Query: 528 NVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGL 576
++R MK KG+ K PG S I V F+VGD SH QSE + Y+LL +
Sbjct: 445 SIRRLMKQKGISKAPGSSIISVDGMSHEFLVGDISHPQSEDI-YILLNI 588
Score = 72.0 bits (175), Expect(2) = 2e-32
Identities = 36/86 (41%), Positives = 56/86 (64%), Gaps = 1/86 (1%)
Frame = +2
Query: 381 AHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLK-NRSIQV 439
A +GYG EA+ F + E G + N+VT++ + TAC H GLVDEG +YF+++ + ++
Sbjct: 20 AINGYGKEALKRFEDLVESGARPNEVTFLAVYTACCHNGLVDEGRKYFNEMTSPHYNLSP 199
Query: 440 KEDHYACLVDLCGRAGRLKEAFYIIE 465
+HY C+VDL RAG + EA +I+
Sbjct: 200 CLEHYGCMVDLLCRAGLVGEAVELIK 277
>AW396441
Length = 459
Score = 133 bits (335), Expect = 2e-31
Identities = 67/155 (43%), Positives = 96/155 (61%)
Frame = +3
Query: 305 FVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGL 364
F V+ A S +ASL GQQ H + K G ++ V ++L++MY+KCG + + K F
Sbjct: 3 FAAVIAAASNIASLRHGQQFHNQVIKMGLDDDPFVTNSLVDMYAKCGSIEESHKAFSS-- 176
Query: 365 LRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEG 424
QRD+ WN MI+ YA HG +A+ +F +M G + N VT+V LL+ACSHAGL+D G
Sbjct: 177 TNQRDIACWNSMISTYAQHGDAAKALEVFERMIMEGVKPNYVTFVGLLSACSHAGLLDLG 356
Query: 425 IQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKE 459
+F+ + K I+ DHYAC+V L GRAG++ E
Sbjct: 357 FHHFESMSK-FGIEPXIDHYACMVSLLGRAGKIYE 458
Score = 68.6 bits (166), Expect = 8e-12
Identities = 35/116 (30%), Positives = 66/116 (56%)
Frame = +3
Query: 239 NAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGL 298
N+++ + + G + + K F+ Q+D+ W SM++ YAQHG + +AL++F +M G+
Sbjct: 111 NSLVDMYAKCGSIEESHKAFSSTNQRDIACWNSMISTYAQHGDAAKALEVFERMIME-GV 287
Query: 299 KPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELH 354
KPN TFV +L ACS L G + +SK G + + ++++ + G+++
Sbjct: 288 KPNYVTFVGLLSACSHAGLLDLGFHHFESMSKFGIEPXIDHYACMVSLLGRAGKIY 455
Score = 47.4 bits (111), Expect = 2e-05
Identities = 26/79 (32%), Positives = 42/79 (52%), Gaps = 4/79 (5%)
Frame = +3
Query: 111 TMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM----PKR 166
++V+ Y K IEE+ F +R+ WN MI Y Q+G KAL++F RM K
Sbjct: 114 SLVDMYAKCGSIEESHKAFSSTNQRDIACWNSMISTYAQHGDAAKALEVFERMIMEGVKP 293
Query: 167 NLVSWNIIIKALARCGRIE 185
N V++ ++ A + G ++
Sbjct: 294 NYVTFVGLLSACSHAGLLD 350
Score = 40.8 bits (94), Expect = 0.002
Identities = 21/59 (35%), Positives = 34/59 (57%)
Frame = +3
Query: 172 NIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERM 230
N ++ A+CG IE++ + R++ WN+MI+ YAQ+ +ALE+FERM
Sbjct: 111 NSLVDMYAKCGSIEESHKAFSSTNQ-----RDIACWNSMISTYAQHGDAAKALEVFERM 272
>TC234029 similar to UP|Q9LN01 (Q9LN01) T6D22.15, partial (32%)
Length = 732
Score = 133 bits (335), Expect = 2e-31
Identities = 60/152 (39%), Positives = 96/152 (62%)
Frame = +2
Query: 437 IQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVA 496
I K HY C++DL R+G+ EA ++ + ++ ++WG LL C +HG + G+ VA
Sbjct: 5 ISPKLQHYGCMIDLLARSGKFDEAKVLMGNMEMEPDGAIWGSLLNACRIHGQVEFGEYVA 184
Query: 497 KKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVF 556
+++ ++E EN+G Y LLSN+YA G+W + A +R K+ DKG+KK PGC+ IE+ V F
Sbjct: 185 ERLFELEPENSGAYVLLSNIYAGAGRWDDVAKIRTKLNDKGMKKVPGCTSIEIDGVVHEF 364
Query: 557 VVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
+VGDK H QSE + +L + +++ G + D
Sbjct: 365 LVGDKFHPQSENIFRMLDEVDRLLEETGFVPD 460
>TC222432
Length = 951
Score = 91.3 bits (225), Expect(2) = 1e-30
Identities = 47/140 (33%), Positives = 76/140 (53%)
Frame = +2
Query: 442 DHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILK 501
+HY C+VDL GRAG L EA +I+ + + + L C + + V K+++K
Sbjct: 221 EHYGCMVDLLGRAGCLDEAENLIQTMPYDANGIILSSFLFACGYFNDVLRAERVLKEVVK 400
Query: 502 VEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDK 561
++ + AG Y +L N+YA+ +W + +V+ MK +G K+ CS IE+G + F GD
Sbjct: 401 MDEDVAGNYVMLRNLYATRQRWTDVEDVKQMMKKRGTSKEVACSVIEIGGSFIEFAAGDY 580
Query: 562 SHSQSEMLEYLLLGLHTKMK 581
HS E+++ L L MK
Sbjct: 581 LHSHLEVIQLTLGQLSKHMK 640
Score = 60.8 bits (146), Expect(2) = 1e-30
Identities = 26/69 (37%), Positives = 47/69 (67%)
Frame = +3
Query: 372 SWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL 431
SWN +I +A +G EA+ +F +M E GF N+VT + +L+AC+H GLV+EG ++F+ +
Sbjct: 12 SWNALINGFAVNGCAKEALEVFARMIEEGFGPNEVTMIGVLSACNHCGLVEEGRRWFNAM 191
Query: 432 LKNRSIQVK 440
+ ++++
Sbjct: 192 ERYLDLRLR 218
Score = 42.7 bits (99), Expect = 5e-04
Identities = 19/59 (32%), Positives = 36/59 (60%)
Frame = +3
Query: 265 DVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQ 323
+ +W +++ G+A +G ++EAL++F +M G PN T + VL AC+ + EG++
Sbjct: 3 ETASWNALINGFAVNGCAKEALEVFARM-IEEGFGPNEVTMIGVLSACNHCGLVEEGRR 176
Score = 34.3 bits (77), Expect = 0.16
Identities = 16/27 (59%), Positives = 19/27 (70%)
Frame = +3
Query: 206 SWNAMITGYAQNRRLDEALELFERMPE 232
SWNA+I G+A N EALE+F RM E
Sbjct: 12 SWNALINGFAVNGCAKEALEVFARMIE 92
>CD397389 similar to GP|6691205|gb| F1K23.11 {Arabidopsis thaliana}, partial
(16%)
Length = 626
Score = 128 bits (322), Expect = 6e-30
Identities = 66/177 (37%), Positives = 98/177 (55%), Gaps = 2/177 (1%)
Frame = -2
Query: 373 WNGMIAAYAHHGYGNEAINLFNKMQ-ELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL 431
W M+ Y +G+ +EA+ LF KM E G N VT + L+AC+HAGLVD+G + +
Sbjct: 625 WTSMMDGYGKNGFPDEALELFVKMPTEYGIVPNYVTLLSALSACAHAGLVDKGWEIIQSM 446
Query: 432 LKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADI 491
++ +HYAC+VDL GRAG L +A+ I + K VW LL+ C +HGN ++
Sbjct: 445 ENEYLVKPGMEHYACMVDLLGRAGMLNQAWEFIMRIPEKPISDVWAALLSSCRLHGNIEL 266
Query: 492 GKLVAKKILKVEHE-NAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWI 547
KL A ++ K+ G Y LSN + G W+ +R MK++G+ K SW+
Sbjct: 265 AKLAANELFKLNATGRPGAYVALSNTLVAAG*WESVTELREIMKERGISKDTXRSWV 95
Score = 55.1 bits (131), Expect = 9e-08
Identities = 44/174 (25%), Positives = 81/174 (46%), Gaps = 5/174 (2%)
Frame = -2
Query: 269 WTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQI---- 324
WTSMM GY ++G +EAL++F KM G+ PN T ++ L AC+ + +G +I
Sbjct: 625 WTSMMDGYGKNGFPDEALELFVKMPTEYGIVPNYVTLLSALSACAHAGLVDKGWEIIQSM 446
Query: 325 -HQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHH 383
++ + K G + + ++++ + G L+ A + F + + W ++++ H
Sbjct: 445 ENEYLVKPGMEH----YACMVDLLGRAGMLNQAWE-FIMRIPEKPISDVWAALLSSCRLH 281
Query: 384 GYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSI 437
G A N++ +L YV L AG E + +++K R I
Sbjct: 280 GNIELAKLAANELFKLNATGRPGAYVALSNTLVAAG*W-ESVTELREIMKERGI 122
Score = 48.5 bits (114), Expect = 8e-06
Identities = 34/128 (26%), Positives = 65/128 (50%), Gaps = 6/128 (4%)
Frame = -2
Query: 140 WNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNII--IKALARCGR--IEDAQWHLIRCR 195
W M+ GYG+NG ++AL+LF +MP + N + + AL+ C + D W +I+
Sbjct: 625 WTSMMDGYGKNGFPDEALELFVKMPTEYGIVPNYVTLLSALSACAHAGLVDKGWEIIQSM 446
Query: 196 KGMMPLR-NVVSWNAMITGYAQNRRLDEALELFERMPERDMAS-WNAMLTGFFQNGELNR 253
+ ++ + + M+ + L++A E R+PE+ ++ W A+L+ +G +
Sbjct: 445 ENEYLVKPGMEHYACMVDLLGRAGMLNQAWEFIMRIPEKPISDVWAALLSSCRLHGNIEL 266
Query: 254 AEKLFAEL 261
A+ EL
Sbjct: 265 AKLAANEL 242
Score = 32.3 bits (72), Expect = 0.61
Identities = 26/122 (21%), Positives = 54/122 (43%), Gaps = 10/122 (8%)
Frame = -2
Query: 77 WTTMIDGYIKCGAIKEARKLFDQPDAKKSV----VTWTTMVNGYVKLNQIEEAESLFYEM 132
WT+M+DGY K G EA +LF + + + VT + ++ +++ + M
Sbjct: 625 WTSMMDGYGKNGFPDEALELFVKMPTEYGIVPNYVTLLSALSACAHAGLVDKGWEIIQSM 446
Query: 133 PERNELS-----WNIMIGGYGQNGQIEKALDLFRRMPKRNLVS-WNIIIKALARCGRIED 186
+ + M+ G+ G + +A + R+P++ + W ++ + G IE
Sbjct: 445 ENEYLVKPGMEHYACMVDLLGRAGMLNQAWEFIMRIPEKPISDVWAALLSSCRLHGNIEL 266
Query: 187 AQ 188
A+
Sbjct: 265 AK 260
>BI893521
Length = 423
Score = 127 bits (319), Expect = 1e-29
Identities = 59/143 (41%), Positives = 91/143 (63%)
Frame = +1
Query: 358 KIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSH 417
++F+D + R++I+W +I+ +A HG+ +A+ LF +M E+G + N+VTY+ +L+ACSH
Sbjct: 1 QVFND--MGYRNVITWTSIISGFAKHGFATKALELFYEMLEIGVKPNEVTYIAVLSACSH 174
Query: 418 AGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWG 477
GL+DE ++F+ + N SI + +HYAC+VDL GR+G L EA I + VW
Sbjct: 175 VGLIDEAWKHFNSMHYNHSISPRMEHYACMVDLLGRSGLLLEAIEFINSMPFDADALVWR 354
Query: 478 PLLAGCNVHGNADIGKLVAKKIL 500
L C VH N +G+ AKKIL
Sbjct: 355 TFLGSCRVHRNTKLGEHAAKKIL 423
Score = 58.5 bits (140), Expect = 8e-09
Identities = 26/65 (40%), Positives = 47/65 (72%)
Frame = +1
Query: 256 KLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGL 315
++F ++ ++VITWTS+++G+A+HG + +AL++F +M G+KPN T++ VL ACS +
Sbjct: 1 QVFNDMGYRNVITWTSIISGFAKHGFATKALELFYEM-LEIGVKPNEVTYIAVLSACSHV 177
Query: 316 ASLTE 320
+ E
Sbjct: 178 GLIDE 192
Score = 45.8 bits (107), Expect = 5e-05
Identities = 32/120 (26%), Positives = 55/120 (45%), Gaps = 5/120 (4%)
Frame = +1
Query: 128 LFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM----PKRNLVSWNIIIKALARCGR 183
+F +M RN ++W +I G+ ++G KAL+LF M K N V++ ++ A + G
Sbjct: 4 VFNDMGYRNVITWTSIISGFAKHGFATKALELFYEMLEIGVKPNEVTYIAVLSACSHVGL 183
Query: 184 IEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP-ERDMASWNAML 242
I++A H + + M+ ++ L EA+E MP + D W L
Sbjct: 184 IDEAWKHFNSMHYNHSISPRMEHYACMVDLLGRSGLLLEAIEFINSMPFDADALVWRTFL 363
Score = 38.9 bits (89), Expect = 0.007
Identities = 38/166 (22%), Positives = 70/166 (41%), Gaps = 5/166 (3%)
Frame = +1
Query: 199 MPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLF 258
M RNV++W ++I+G+A++ +ALELF M E +
Sbjct: 16 MGYRNVITWTSIISGFAKHGFATKALELFYEMLEIGV----------------------- 126
Query: 259 AELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASL 318
+ + +T+ ++++ + GL +EA K F M N + P + ++ L
Sbjct: 127 ----KPNEVTYIAVLSACSHVGLIDEAWKHFNSMHYNHSISPRMEHYACMVDLLGRSGLL 294
Query: 319 TEGQQIHQLISKTGFQEN-----TRVVSALINMYSKCGELHIARKI 359
E + I+ F + T + S ++ +K GE H A+KI
Sbjct: 295 LEA---IEFINSMPFDADALVWRTFLGSCRVHRNTKLGE-HAAKKI 420
Score = 38.9 bits (89), Expect = 0.007
Identities = 25/93 (26%), Positives = 48/93 (50%), Gaps = 9/93 (9%)
Frame = +1
Query: 104 KSVVTWTTMVNGYVKLNQIEEAESLFYEMPE----RNELSWNIMIGGYGQNGQIEKALDL 159
++V+TWT++++G+ K +A LFYEM E NE+++ ++ G I++A
Sbjct: 25 RNVITWTSIISGFAKHGFATKALELFYEMLEIGVKPNEVTYIAVLSACSHVGLIDEAWKH 204
Query: 160 FRRMPKRNLVS-----WNIIIKALARCGRIEDA 187
F M + +S + ++ L R G + +A
Sbjct: 205 FNSMHYNHSISPRMEHYACMVDLLGRSGLLLEA 303
>AW317686
Length = 447
Score = 124 bits (312), Expect = 9e-29
Identities = 56/140 (40%), Positives = 93/140 (66%), Gaps = 2/140 (1%)
Frame = +1
Query: 368 RDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQY 427
RD+++WN +I +A +G+G E++ +F +M E + N VT++ +L+ C+HAGL +EG+Q
Sbjct: 28 RDVVTWNTLITGFAQNGHGEESLAVFRRMIEAKVEPNHVTFLGVLSGCNHAGLDNEGLQL 207
Query: 428 FDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGL--GVKLSLSVWGPLLAGCNV 485
D + + ++ K +HYA L+DL GR RL EA +IE + G+K ++VWG +L C V
Sbjct: 208 VDLMERQYGVKPKAEHYALLIDLLGRRNRLMEAMSLIEKVPDGIKNHIAVWGAVLGACXV 387
Query: 486 HGNADIGKLVAKKILKVEHE 505
HGN D+ + A+K+ ++E E
Sbjct: 388 HGNLDLARKAAEKLFELEPE 447
Score = 67.4 bits (163), Expect = 2e-11
Identities = 44/149 (29%), Positives = 76/149 (50%), Gaps = 4/149 (2%)
Frame = +1
Query: 255 EKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSG 314
E LF P +DV+TW +++TG+AQ+G EE+L +F +M ++PN+ TF+ VL C+
Sbjct: 1 ENLFEMAPMRDVVTWNTLITGFAQNGHGEESLAVFRRM-IEAKVEPNHVTFLGVLSGCNH 177
Query: 315 LASLTEGQQIHQLISKT-GFQENTRVVSALINMYSKCGELHIARKIFD---DGLLRQRDL 370
EG Q+ L+ + G + + LI++ + L A + + DG+ + +
Sbjct: 178 AGLDNEGLQLVDLMERQYGVKPKAEHYALLIDLLGRRNRLMEAMSLIEKVPDGI--KNHI 351
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQEL 399
W ++ A HG + A K+ EL
Sbjct: 352 AVWGAVLGACXVHGNLDLARKAAEKLFEL 438
Score = 58.2 bits (139), Expect = 1e-08
Identities = 36/137 (26%), Positives = 64/137 (46%), Gaps = 4/137 (2%)
Frame = +1
Query: 226 LFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDV----ITWTSMMTGYAQHGL 281
LFE P RD+ +WN ++TGF QNG + +F + + V +T+ +++G GL
Sbjct: 7 LFEMAPMRDVVTWNTLITGFAQNGHGEESLAVFRRMIEAKVEPNHVTFLGVLSGCNHAGL 186
Query: 282 SEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVS 341
E L++ M+ G+KP + ++ L E + + + G + + V
Sbjct: 187 DNEGLQLVDLMERQYGVKPKAEHYALLIDLLGRRNRLMEAMSLIEKV-PDGIKNHIAVWG 363
Query: 342 ALINMYSKCGELHIARK 358
A++ G L +ARK
Sbjct: 364 AVLGACXVHGNLDLARK 414
Score = 53.5 bits (127), Expect = 3e-07
Identities = 42/149 (28%), Positives = 75/149 (50%), Gaps = 11/149 (7%)
Frame = +1
Query: 126 ESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNI-IIKALARCGR- 183
E+LF P R+ ++WN +I G+ QNG E++L +FRRM + + ++ + L+ C
Sbjct: 1 ENLFEMAPMRDVVTWNTLITGFAQNGHGEESLAVFRRMIEAKVEPNHVTFLGVLSGCNHA 180
Query: 184 -IEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNR-RLDEALELFERMPE---RDMASW 238
+++ L+ + ++ A++ R RL EA+ L E++P+ +A W
Sbjct: 181 GLDNEGLQLVDLMERQYGVKPKAEHYALLIDLLGRRNRLMEAMSLIEKVPDGIKNHIAVW 360
Query: 239 NAMLTGFFQNGELN----RAEKLFAELPQ 263
A+L +G L+ AEKLF P+
Sbjct: 361 GAVLGACXVHGNLDLARKAAEKLFELEPE 447
Score = 37.0 bits (84), Expect = 0.025
Identities = 34/142 (23%), Positives = 64/142 (44%), Gaps = 13/142 (9%)
Frame = +1
Query: 104 KSVVTWTTMVNGYVKLNQIEEAESLFYEM----PERNELSWNIMIGGYGQNGQIEKALDL 159
+ VVTW T++ G+ + EE+ ++F M E N +++ ++ G G + L L
Sbjct: 28 RDVVTWNTLITGFAQNGHGEESLAVFRRMIEAKVEPNHVTFLGVLSGCNHAGLDNEGLQL 207
Query: 160 FRRMPKRNLVS-----WNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGY 214
M ++ V + ++I L R R+ +A + + G+ ++ W A++
Sbjct: 208 VDLMERQYGVKPKAEHYALLIDLLGRRNRLMEAMSLIEKVPDGIK--NHIAVWGAVLGAC 381
Query: 215 AQNRRLD----EALELFERMPE 232
+ LD A +LFE PE
Sbjct: 382 XVHGNLDLARKAAEKLFELEPE 447
>BM177962
Length = 421
Score = 119 bits (297), Expect = 5e-27
Identities = 56/133 (42%), Positives = 89/133 (66%)
Frame = +2
Query: 300 PNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKI 359
PN+ T V VL AC+GLA+L +G+ IH I + G V++ALI MY +CGE+ + +++
Sbjct: 23 PNSVTMVNVLQACAGLAALEQGKLIHGYILRRGLDSILPVLNALITMYGRCGEILMGQRV 202
Query: 360 FDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAG 419
FD+ ++ RD++SWN +I+ Y HG+G +AI +F M G + ++++ +L ACSHAG
Sbjct: 203 FDN--MKNRDVVSWNSLISIYGMHGFGKKAIQIF*NMIHQGGSPSYISFITVLGACSHAG 376
Query: 420 LVDEGIQYFDKLL 432
LV+EG F+ +L
Sbjct: 377 LVEEGKILFESML 415
Score = 66.6 bits (161), Expect = 3e-11
Identities = 38/122 (31%), Positives = 69/122 (56%), Gaps = 1/122 (0%)
Frame = +2
Query: 210 MITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
+I GY R LD L + NA++T + + GE+ +++F + +DV++W
Sbjct: 92 LIHGYILRRGLDSILPVL-----------NALITMYGRCGEILMGQRVFDNMKNRDVVSW 238
Query: 270 TSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQ-LI 328
S+++ Y HG ++A+++F M GG P+ +F+TVLGACS + EG+ + + ++
Sbjct: 239 NSLISIYGMHGFGKKAIQIF*NMIHQGG-SPSYISFITVLGACSHAGLVEEGKILFESML 415
Query: 329 SK 330
SK
Sbjct: 416 SK 421
Score = 37.4 bits (85), Expect = 0.019
Identities = 20/81 (24%), Positives = 43/81 (52%), Gaps = 4/81 (4%)
Frame = +2
Query: 112 MVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKR----N 167
++ Y + +I + +F M R+ +SWN +I YG +G +KA+ +F M + +
Sbjct: 152 LITMYGRCGEILMGQRVFDNMKNRDVVSWNSLISIYGMHGFGKKAIQIF*NMIHQGGSPS 331
Query: 168 LVSWNIIIKALARCGRIEDAQ 188
+S+ ++ A + G +E+ +
Sbjct: 332 YISFITVLGACSHAGLVEEGK 394
Score = 30.0 bits (66), Expect = 3.0
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +2
Query: 141 NIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIII 175
N +I YG+ G+I +F M R++VSWN +I
Sbjct: 146 NALITMYGRCGEILMGQRVFDNMKNRDVVSWNSLI 250
Score = 28.5 bits (62), Expect = 8.8
Identities = 18/59 (30%), Positives = 29/59 (48%)
Frame = +2
Query: 172 NIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERM 230
N +I RCG I Q + R+VVSWN++I+ Y + +A+++F M
Sbjct: 146 NALITMYGRCGEILMGQRVFDNMKN-----RDVVSWNSLISIYGMHGFGKKAIQIF*NM 307
>TC230979 weakly similar to UP|Q6Z8F8 (Q6Z8F8) Selenium-binding protein-like,
partial (61%)
Length = 837
Score = 116 bits (290), Expect = 3e-26
Identities = 58/154 (37%), Positives = 88/154 (56%)
Frame = +2
Query: 428 FDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHG 487
F+++ I+ HY C+VDL GRAG LKEA+ +I+ + +K + VW LL+ C VH
Sbjct: 5 FNRMQFEHMIKPTIQHYGCMVDLMGRAGMLKEAYDLIKSMPIKPNDVVWRSLLSACKVHH 184
Query: 488 NADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWI 547
N +IG++ A I K+ N G Y +L+NMYA KW A +R +M +K L + PG S +
Sbjct: 185 NLEIGEIAADNIFKLNKHNPGDYLVLANMYARAQKWANVARIRTEMVEKNLVQTPGFSLV 364
Query: 548 EVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMK 581
E V FV DKS + E + ++ + ++K
Sbjct: 365 EANRNVYKFVSQDKSQPKCETIYDMIQQMEWQLK 466
>BG790334 weakly similar to GP|2160154|gb| F19K23.18 gene product
{Arabidopsis thaliana}, partial (19%)
Length = 390
Score = 111 bits (277), Expect = 1e-24
Identities = 49/128 (38%), Positives = 82/128 (63%)
Frame = +2
Query: 396 MQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAG 455
M+ L +T++ +L AC+HAGLV+EG + F ++ + I+ + +H+A LVD+ GR G
Sbjct: 5 MKRLKIHPTYITFISVLNACAHAGLVEEGWRQFKSMINDYGIEPRVEHFASLVDILGRQG 184
Query: 456 RLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSN 515
+L+EA +I + K +VWG LL C VH N ++ + A ++++E E++ Y LL N
Sbjct: 185 QLQEAMDLINTMPFKPDKAVWGALLGACRVHNNVELALVAADALIRLEPESSAPYVLLYN 364
Query: 516 MYASVGKW 523
MYA++G+W
Sbjct: 365 MYANLGQW 388
Score = 32.3 bits (72), Expect = 0.61
Identities = 20/87 (22%), Positives = 42/87 (47%), Gaps = 1/87 (1%)
Frame = +2
Query: 298 LKPNNGTFVTVLGACSGLASLTEG-QQIHQLISKTGFQENTRVVSALINMYSKCGELHIA 356
+ P TF++VL AC+ + EG +Q +I+ G + ++L+++ + G+L A
Sbjct: 20 IHPTYITFISVLNACAHAGLVEEGWRQFKSMINDYGIEPRVEHFASLVDILGRQGQLQEA 199
Query: 357 RKIFDDGLLRQRDLISWNGMIAAYAHH 383
+ + + + D W ++ A H
Sbjct: 200 MDLINT-MPFKPDKAVWGALLGACRVH 277
>BE802290
Length = 481
Score = 111 bits (277), Expect = 1e-24
Identities = 52/155 (33%), Positives = 91/155 (58%)
Frame = -1
Query: 393 FNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCG 452
F + ++ + N + ++ + ACSHA L +EG F+ + +I+ +HY C+VDL G
Sbjct: 481 FTQCRKQKSKPNSIPFMAISPACSHAALTEEGTSSFESMSSVYNIKPSMEHYGCMVDLMG 302
Query: 453 RAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSL 512
AG LKEA +E + VK + ++WG LL C +H ++GK K ++++ +++G Y
Sbjct: 301 GAGLLKEAREFVESMPVKPNAAIWGALLNACQLHKYFELGKETGKIQIELDPDHSGRYIH 122
Query: 513 LSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWI 547
L+++YA+ +W + VR ++K +GL GCS I
Sbjct: 121 LASIYAAAREWNQVTRVRSQIKHRGLLNYSGCSSI 17
>BE211268
Length = 422
Score = 109 bits (273), Expect = 3e-24
Identities = 53/139 (38%), Positives = 80/139 (57%)
Frame = +1
Query: 413 TACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLS 472
T CSHAGL++EGI+ F ++ + +HY +VDL GR G L +A +I + ++
Sbjct: 1 TRCSHAGLIEEGIKMFHVMVNEYQLMPNTEHYGIMVDLLGRMGELDKALDMINEMPMQAG 180
Query: 473 LSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMK 532
VWG LL C +H N IG+L A + ++ +AG Y+LLSN+Y W +AA +R
Sbjct: 181 PHVWGALLGACRIHQNIKIGELAALNLFLLDPNHAGYYTLLSNIYCVDKNWHDAAKLRTL 360
Query: 533 MKDKGLKKQPGCSWIEVGN 551
+K+ KK G S +E+ N
Sbjct: 361 IKENRFKKIVGQSMVEIKN 417
>TC223934
Length = 889
Score = 109 bits (273), Expect = 3e-24
Identities = 52/150 (34%), Positives = 95/150 (62%)
Frame = +2
Query: 199 MPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLF 258
MP R+V +W MI+ + ++ + A LF+ MPE+++A+WNAM+ G+ + G AE LF
Sbjct: 443 MPERDVFAWTTMISAHVRDGDMASAGRLFDEMPEKNVATWNAMIDGYGKLGNAESAEFLF 622
Query: 259 AELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASL 318
++P +D+I+WT+MM Y+++ +E + +F + + G+ P+ T TV+ AC+ L +L
Sbjct: 623 NQMPARDIISWTTMMNCYSRNKRYKEVIALFHDV-IDKGMIPDEVTMTTVISACAHLGAL 799
Query: 319 TEGQQIHQLISKTGFQENTRVVSALINMYS 348
G+++H + GF + + S+LI+MY+
Sbjct: 800 ALGKEVHLYLVLQGFDLDVYIGSSLIDMYA 889
Score = 92.4 bits (228), Expect = 5e-19
Identities = 47/132 (35%), Positives = 76/132 (56%), Gaps = 4/132 (3%)
Frame = +2
Query: 57 GRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGY 116
G +R++FD MP+RD WTTMI +++ G + A +LFD+ +K+V TW M++GY
Sbjct: 407 GDVGGSRRVFDDMPERDVFAWTTMISAHVRDGDMASAGRLFDEM-PEKNVATWNAMIDGY 583
Query: 117 VKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWN 172
KL E AE LF +MP R+ +SW M+ Y +N + ++ + LF + + + V+
Sbjct: 584 GKLGNAESAEFLFNQMPARDIISWTTMMNCYSRNKRYKEVIALFHDVIDKGMIPDEVTMT 763
Query: 173 IIIKALARCGRI 184
+I A A G +
Sbjct: 764 TVISACAHLGAL 799
Score = 87.8 bits (216), Expect = 1e-17
Identities = 49/132 (37%), Positives = 79/132 (59%), Gaps = 4/132 (3%)
Frame = +2
Query: 73 DKHLW--TTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFY 130
D H++ TT+I+ Y G + +R++FD ++ V WTTM++ +V+ + A LF
Sbjct: 356 DSHVFVQTTLIEFYSTFGDVGGSRRVFDDMP-ERDVFAWTTMISAHVRDGDMASAGRLFD 532
Query: 131 EMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIED--AQ 188
EMPE+N +WN MI GYG+ G E A LF +MP R+++SW ++ +R R ++ A
Sbjct: 533 EMPEKNVATWNAMIDGYGKLGNAESAEFLFNQMPARDIISWTTMMNCYSRNKRYKEVIAL 712
Query: 189 WHLIRCRKGMMP 200
+H + KGM+P
Sbjct: 713 FHDV-IDKGMIP 745
Score = 65.1 bits (157), Expect = 9e-11
Identities = 55/275 (20%), Positives = 112/275 (40%), Gaps = 3/275 (1%)
Frame = +2
Query: 153 IEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRC-RKGMMPLRNVVSWNAMI 211
I A F + N++ +N +I+ C E A H + R +MP S
Sbjct: 110 INLAASAFANVQNPNVLVFNALIRGCVHCCYSEQALVHYMHMLRNNVMPTSYSFSSLIKA 289
Query: 212 TGYAQNRRLDEALE--LFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
+ EA+ +++ + + ++ + G++ + ++F ++P++DV W
Sbjct: 290 CTLLVDSAFGEAVHGHVWKHGFDSHVFVQTTLIEFYSTFGDVGGSRRVFDDMPERDVFAW 469
Query: 270 TSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLIS 329
T+M++ + + G A ++F +M
Sbjct: 470 TTMISAHVRDGDMASAGRLFDEMP------------------------------------ 541
Query: 330 KTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEA 389
++N +A+I+ Y K G A +F+ + RD+ISW M+ Y+ + E
Sbjct: 542 ----EKNVATWNAMIDGYGKLGNAESAEFLFNQ--MPARDIISWTTMMNCYSRNKRYKEV 703
Query: 390 INLFNKMQELGFQANDVTYVELLTACSHAGLVDEG 424
I LF+ + + G ++VT +++AC+H G + G
Sbjct: 704 IALFHDVIDKGMIPDEVTMTTVISACAHLGALALG 808
Score = 65.1 bits (157), Expect = 9e-11
Identities = 33/101 (32%), Positives = 59/101 (57%)
Frame = +2
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F P ++ + IS ++G A +LFD+MP+++ W MIDGY K G + A
Sbjct: 434 FDDMPERDVFAWTTMISAHVRDGDMASAGRLFDEMPEKNVATWNAMIDGYGKLGNAESAE 613
Query: 95 KLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPER 135
LF+Q A + +++WTTM+N Y + + +E +LF+++ ++
Sbjct: 614 FLFNQMPA-RDIISWTTMMNCYSRNKRYKEVIALFHDVIDK 733
Score = 42.4 bits (98), Expect = 6e-04
Identities = 29/91 (31%), Positives = 41/91 (44%), Gaps = 3/91 (3%)
Frame = +2
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F P + N+ I K G + A LF+QMP RD WTTM++ Y + KE
Sbjct: 527 FDEMPEKNVATWNAMIDGYGKLGNAESAEFLFNQMPARDIISWTTMMNCYSRNKRYKEVI 706
Query: 95 KLFDQPDAKKSV---VTWTTMVNGYVKLNQI 122
LF K + VT TT+++ L +
Sbjct: 707 ALFHDVIDKGMIPDEVTMTTVISACAHLGAL 799
>TC222005 weakly similar to UP|Q9FIB2 (Q9FIB2) Selenium-binding protein-like,
partial (13%)
Length = 757
Score = 108 bits (271), Expect = 5e-24
Identities = 55/129 (42%), Positives = 78/129 (59%), Gaps = 1/129 (0%)
Frame = +2
Query: 469 VKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAAN 528
VK +W LL GC +HGN ++ K AK + ++E EN TY L+N+YA+ G W E AN
Sbjct: 8 VKPDKFLWASLLGGCRIHGNLELAKRAAKALYEIEPENPATYITLANIYANAGLWSEVAN 187
Query: 529 VRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
VR M + G+ K+PG SWIE+ V VF+VGD SH ++ + L L K+K+ G + D
Sbjct: 188 VRKDMDNMGIVKKPGKSWIEIKRQVHVFLVGDTSHPKTSDIHEFLGELSKKIKEEGYVPD 367
Query: 589 DD-LSRDVE 596
+ + DVE
Sbjct: 368 TNFVLHDVE 394
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,873,460
Number of Sequences: 63676
Number of extensions: 374149
Number of successful extensions: 2127
Number of sequences better than 10.0: 163
Number of HSP's better than 10.0 without gapping: 1705
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1996
length of query: 597
length of database: 12,639,632
effective HSP length: 103
effective length of query: 494
effective length of database: 6,081,004
effective search space: 3004015976
effective search space used: 3004015976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0026.14


