

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
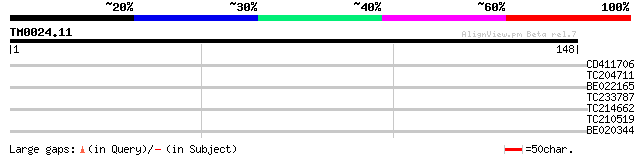
Query= TM0024.11
(148 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CD411706 28 2.1
TC204711 similar to UP|ATP7_ARATH (Q9SJ12) Probable ATP synthase... 27 3.6
BE022165 similar to PIR|T31068|T31 N-methyl-D-aspartate receptor... 27 4.7
TC233787 similar to UP|Q7XAV5 (Q7XAV5) Dehydration responsive el... 26 6.2
TC214662 26 8.1
TC210519 weakly similar to UP|Q6NMK2 (Q6NMK2) At5g49400, partial... 26 8.1
BE020344 26 8.1
>CD411706
Length = 630
Score = 27.7 bits (60), Expect = 2.1
Identities = 16/58 (27%), Positives = 28/58 (47%), Gaps = 2/58 (3%)
Frame = +3
Query: 92 LVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHARRKAANSC--HAYTDSTRESEINL 147
++E RIF ++ S L +S+ + Y GH+ +A + H ++RE E L
Sbjct: 237 ILERSNMRIFSDTRGSPESLNKSTLHFVGYKGHSAERACSLIVPHPTVQTSREREREL 410
>TC204711 similar to UP|ATP7_ARATH (Q9SJ12) Probable ATP synthase 24 kDa
subunit, mitochondrial precursor , partial (86%)
Length = 1143
Score = 26.9 bits (58), Expect = 3.6
Identities = 8/34 (23%), Positives = 22/34 (64%)
Frame = -2
Query: 18 YDFSTIKFRNHVHQGILREILASEFQNSKQISKI 51
Y + ++KF H+H+ +L+ +++ + + + SK+
Sbjct: 356 YSYGSLKFVLHLHKNVLQHLISLKGRRGRLFSKV 255
>BE022165 similar to PIR|T31068|T31 N-methyl-D-aspartate receptor homolog
NMDAR-L - rat, partial (1%)
Length = 425
Score = 26.6 bits (57), Expect = 4.7
Identities = 23/79 (29%), Positives = 33/79 (41%), Gaps = 7/79 (8%)
Frame = +3
Query: 50 KIWEKRYLKSQKRWF-----RSKEISVARLPESEYYRAYSPEF--RNETLVEGKIKRIFK 102
K WE + +K WF R++ I + R +YR+YSP +T V + R K
Sbjct: 123 KTWE---MDVKKYWFGSRSQRTQNIRLGRNYTKPHYRSYSPSSCDGTKTTVWQMLWRKLK 293
Query: 103 FSKDSKSHLKRSSKSEIAY 121
S D K + E Y
Sbjct: 294 RSSDKKKVFSSHTTMEGVY 350
>TC233787 similar to UP|Q7XAV5 (Q7XAV5) Dehydration responsive element
binding protein, partial (25%)
Length = 505
Score = 26.2 bits (56), Expect = 6.2
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = -3
Query: 84 SPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSG 123
SPE+ N +F FS +S+S RSS+ +YSG
Sbjct: 344 SPEYEN--------MHLFSFSFNSRSTHSRSSRVRASYSG 249
>TC214662
Length = 1129
Score = 25.8 bits (55), Expect = 8.1
Identities = 21/71 (29%), Positives = 33/71 (45%)
Frame = -3
Query: 6 SVTTIPKGTKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFR 65
S TTI G + Y+ S + I + A S I I +K+YLKS W +
Sbjct: 455 SATTIAGGRHAAYE-SIMNNPIQASVAIEQAEAAGSAFFSSIIYSICKKKYLKSHATW-K 282
Query: 66 SKEISVARLPE 76
+K +S ++P+
Sbjct: 281 TKRLSTKQIPK 249
>TC210519 weakly similar to UP|Q6NMK2 (Q6NMK2) At5g49400, partial (8%)
Length = 517
Score = 25.8 bits (55), Expect = 8.1
Identities = 22/80 (27%), Positives = 38/80 (47%)
Frame = -2
Query: 14 TKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVAR 73
T + YD + I +NH HQ + N K I K +K +++ R + KEISV
Sbjct: 348 TNNHYDKTLILIKNHQHQNTWK-YST*ILSNHKSIHK-RKKNRNRNRNREKKRKEISV-- 181
Query: 74 LPESEYYRAYSPEFRNETLV 93
+P+ +++P + L+
Sbjct: 180 VPQKTLKTSHNPTLWHYKLI 121
>BE020344
Length = 354
Score = 25.8 bits (55), Expect = 8.1
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +3
Query: 41 EFQNSKQISKIWEKRYLKSQKRWFR 65
+++N ++ I KR LKSQK+W R
Sbjct: 267 QYKN*PVLNSIC*KRNLKSQKKWMR 341
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,726,080
Number of Sequences: 63676
Number of extensions: 63763
Number of successful extensions: 363
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 363
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 363
length of query: 148
length of database: 12,639,632
effective HSP length: 88
effective length of query: 60
effective length of database: 7,036,144
effective search space: 422168640
effective search space used: 422168640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0024.11


