

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0023.5
(266 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
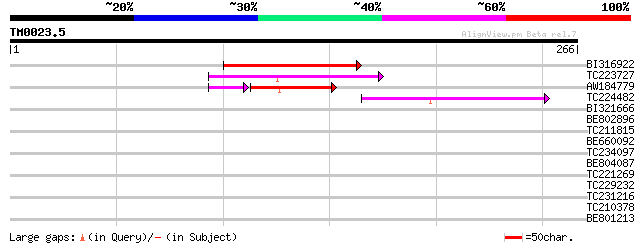
Score E
Sequences producing significant alignments: (bits) Value
BI316922 72 2e-13
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 67 8e-12
AW184779 42 8e-06
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 45 4e-05
BI321666 40 0.001
BE802896 39 0.003
TC211815 34 0.060
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 32 0.23
TC234097 similar to UP|Q9ZR25 (Q9ZR25) UDP-glucose:anthocyanin 5... 29 1.9
BE804087 28 4.3
TC221269 similar to UP|Q7RHR7 (Q7RHR7) Retinitis pigmentosa GTPa... 28 4.3
TC229232 similar to GB|AAP37829.1|30725614|BT008470 At5g12470 {A... 28 5.6
TC231216 28 5.6
TC210378 27 7.3
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 27 9.5
>BI316922
Length = 405
Score = 72.4 bits (176), Expect = 2e-13
Identities = 32/65 (49%), Positives = 46/65 (70%)
Frame = +3
Query: 101 EQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNF 160
E+L + APWPFAI G DIL F ++K Q+K+++V +D FTKWIE E + I++A +R F
Sbjct: 168 EELHNIVAPWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKF 347
Query: 161 LWRRI 165
+ R I
Sbjct: 348 V*RNI 362
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 67.0 bits (162), Expect = 8e-12
Identities = 31/83 (37%), Positives = 47/83 (56%), Gaps = 1/83 (1%)
Frame = +1
Query: 94 DLSGAPPEQLTTMTAPWPFAIWGADILGLFS-VAKAQMKFIVVAVDYFTKWIEAEAVETI 152
D APP L M++PWPF++WG D++G A +FI+VA+DYFTKW+EA + +
Sbjct: 472 DNVNAPPHPLNVMSSPWPFSMWGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDV 651
Query: 153 TVAKIRNFLWRRITSTGETPYKL 175
+ F+ + I P K+
Sbjct: 652 MRGVVVRFIKKEIICRYGLPRKI 720
>AW184779
Length = 432
Score = 41.6 bits (96), Expect(2) = 8e-06
Identities = 18/41 (43%), Positives = 27/41 (64%), Gaps = 1/41 (2%)
Frame = +3
Query: 114 IWGADILGLFSV-AKAQMKFIVVAVDYFTKWIEAEAVETIT 153
+WG D++G A FI+VA+DYFTKW+EA + ++T
Sbjct: 291 MWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVT 413
Score = 25.0 bits (53), Expect(2) = 8e-06
Identities = 9/19 (47%), Positives = 11/19 (57%)
Frame = +2
Query: 94 DLSGAPPEQLTTMTAPWPF 112
D APP L + APWP+
Sbjct: 236 DNVNAPPIPLNVLAAPWPY 292
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 44.7 bits (104), Expect = 4e-05
Identities = 30/91 (32%), Positives = 44/91 (47%), Gaps = 3/91 (3%)
Frame = +1
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQSWRVIT---MNEEENNDNLIANLIMLPELQREAHL 222
TSTG TP+ L YG +A+LP EVE S R++ + E E L ++ + A
Sbjct: 112 TSTGATPFSLVYGMEAVLPFEVEVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMS 291
Query: 223 RNEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ R+ + KV RK DLVL++
Sbjct: 292 HGRLYQQRMKSAFDKKVCLRKFHEGDLVLKK 384
>BI321666
Length = 430
Score = 40.0 bits (92), Expect = 0.001
Identities = 22/64 (34%), Positives = 35/64 (54%)
Frame = +2
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRITSTGET 171
F WG D +G F + + ++I+V VDY +KW+EA A + + FL ++I S
Sbjct: 155 FDYWGIDFVGPFPPSFSN-EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGV 331
Query: 172 PYKL 175
P+ L
Sbjct: 332 PWVL 343
>BE802896
Length = 416
Score = 38.5 bits (88), Expect = 0.003
Identities = 24/80 (30%), Positives = 36/80 (45%)
Frame = -2
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P P L ++ A+ +L Q+ K ++IY+ S TL A+ Y EK LAI+
Sbjct: 247 QAPDWTAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIV 68
Query: 64 KTARRLRPYFQSFQIKVKTD 83
+ Y +I V D
Sbjct: 67 FALEKFHSYLLGTRIIVYID 8
>TC211815
Length = 704
Score = 34.3 bits (77), Expect = 0.060
Identities = 23/76 (30%), Positives = 39/76 (51%)
Frame = +2
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPELQREAHLRNE 225
++T ETP+ L YG MLP+EV E N + L +L ++ +++ + +
Sbjct: 233 STTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIMT* 412
Query: 226 AGKGRVARKYASKVDS 241
A K R+ R + SK+ S
Sbjct: 413 AFKQRMTRCFNSKLPS 460
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 32.3 bits (72), Expect = 0.23
Identities = 21/73 (28%), Positives = 39/73 (52%), Gaps = 7/73 (9%)
Frame = -3
Query: 156 KIRNFLWRRITS----TGETPYKLTYGSDAMLPVEVENQS-WRVITMN--EEENNDNLIA 208
++ + LW T+ G +PY++ +G LPVE+E+++ W V T N ++ +
Sbjct: 226 RLDDALWAHRTAYKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKL 47
Query: 209 NLIMLPELQREAH 221
L L E++ EA+
Sbjct: 46 QLSELDEIRLEAY 8
>TC234097 similar to UP|Q9ZR25 (Q9ZR25) UDP-glucose:anthocyanin
5-O-glucosyltransferase, partial (30%)
Length = 435
Score = 29.3 bits (64), Expect = 1.9
Identities = 17/48 (35%), Positives = 28/48 (57%)
Frame = +2
Query: 7 TPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQK 54
T + PLIL+LAVTS+A +++ + + +F+SH + A R K
Sbjct: 50 TGKTPLILHLAVTSSAPRMIVVNGWTQSQRCRWFMSHLVVFACCRRSK 193
>BE804087
Length = 160
Score = 28.1 bits (61), Expect = 4.3
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +2
Query: 166 TSTGETPYKLTYGSDA 181
TSTG TPY L YG +A
Sbjct: 113 TSTGATPYSLVYGKEA 160
>TC221269 similar to UP|Q7RHR7 (Q7RHR7) Retinitis pigmentosa GTPase
regulator-like protein, partial (3%)
Length = 802
Score = 28.1 bits (61), Expect = 4.3
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 4/59 (6%)
Frame = -1
Query: 141 TKWIEAEAVETITVAKIRNFLWRRITST----GETPYKLTYGSDAMLPVEVENQSWRVI 195
T W E + + KI + LW T PY TY A L + + Q+W V+
Sbjct: 241 TNWGERPTITCVQKKKIISCLWLIFQLTEYRLSPKPYFCTYSRRAWLHISMFRQNWTVM 65
>TC229232 similar to GB|AAP37829.1|30725614|BT008470 At5g12470 {Arabidopsis
thaliana;} , partial (73%)
Length = 1309
Score = 27.7 bits (60), Expect = 5.6
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = +2
Query: 6 PTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFV 41
P P +P LYL++ +T S L YK IYF+
Sbjct: 983 PGPRLPHDLYLSLLTTFFSTCLFASVLDLYKTIYFL 1090
>TC231216
Length = 726
Score = 27.7 bits (60), Expect = 5.6
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -1
Query: 36 KMIYFVSHTLQGAEVRYQKIEK 57
K I++VSHT+ E+R KI+K
Sbjct: 393 KNIFYVSHTMHTTEIRKNKIKK 328
>TC210378
Length = 751
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/45 (26%), Positives = 28/45 (61%)
Frame = +3
Query: 120 LGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRR 164
L + + ++ ++KF++ DY+ + ++ E V+ + AK+R +W R
Sbjct: 240 LWIVATSRWELKFLMRIGDYWRRLLQGERVQDL--AKVRILVWPR 368
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 26.9 bits (58), Expect = 9.5
Identities = 14/49 (28%), Positives = 28/49 (56%), Gaps = 5/49 (10%)
Frame = +2
Query: 155 AKIRNFLWR----RITSTGETPYKLTYGSDAMLPVEVENQS-WRVITMN 198
+K+ + LW + T G TP+++ Y LPVE+++++ W + +N
Sbjct: 260 SKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELKHKAYWAMKCLN 406
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,524,551
Number of Sequences: 63676
Number of extensions: 118654
Number of successful extensions: 507
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 505
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 506
length of query: 266
length of database: 12,639,632
effective HSP length: 96
effective length of query: 170
effective length of database: 6,526,736
effective search space: 1109545120
effective search space used: 1109545120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0023.5


