

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
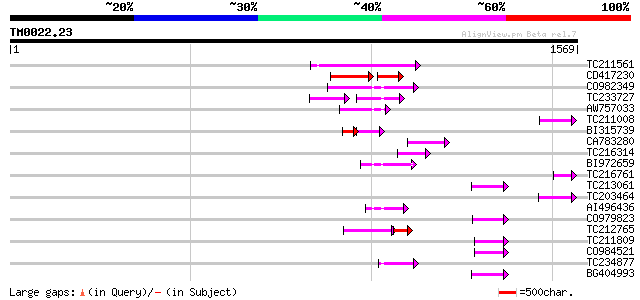
Query= TM0022.23
(1569 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 154 3e-37
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 97 3e-32
CO982349 128 2e-29
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 61 2e-17
AW757033 88 3e-17
TC211008 83 1e-15
BI315739 50 1e-13
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 68 4e-11
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 63 9e-10
BI972659 62 2e-09
TC216761 similar to UP|Q9FQE4 (Q9FQE4) Glutathione S-transferase... 60 6e-09
TC213061 57 5e-08
TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leish... 57 6e-08
AI496436 57 8e-08
CO979823 57 8e-08
TC212765 55 2e-07
TC211809 53 9e-07
CO984521 52 2e-06
TC234877 52 2e-06
BG404993 51 5e-06
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase (Fragment)
, partial (59%)
Length = 1077
Score = 154 bits (389), Expect = 3e-37
Identities = 99/306 (32%), Positives = 153/306 (49%), Gaps = 1/306 (0%)
Frame = +1
Query: 832 IDNAIILQKIVHHMNSRRGKCQNVVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIM 891
+ +A+I +++ C +VFK+D EKA SVSW FL + F P + I
Sbjct: 1 LHSALIANEVIDEAKRSNKSC--LVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIE 174
Query: 892 FCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELV 951
CV S+S+S+L NG P RGLR GDPL+P+LF + E L ++ AV++N +
Sbjct: 175 ECVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPY 354
Query: 952 RISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSV 1011
+ +G PIS L +ADD + +A V ++ IL F + S L +N AKS F V
Sbjct: 355 LVGANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKS-CFGVFGV 531
Query: 1012 PRAKKADIFVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKA 1071
K + A YLG PI + +D ++ + RK + WK R ++
Sbjct: 532 TDQWKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFG 711
Query: 1072 GKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSRG-ERRGMNMVAWRTVTLP 1130
G+ L K VL +IP+Y F +V D L + + F+W G +++ ++ + W + LP
Sbjct: 712 GRVTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLP 891
Query: 1131 KCNGGL 1136
K GG+
Sbjct: 892 KERGGI 909
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like protein
{Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 97.4 bits (241), Expect(2) = 3e-32
Identities = 48/120 (40%), Positives = 78/120 (65%)
Frame = -3
Query: 887 VSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQN 946
V L+ +C+S++++SLLWNG L +P+ G+R DP++PYLFVLC+ERL+ I +++
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKK 507
Query: 947 QGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDF 1006
++++R G ISHL F DD++L +A + QV + L F +SG VN+ K++ F
Sbjct: 506 L*RSIQVNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTKSF 327
Score = 61.6 bits (148), Expect(2) = 3e-32
Identities = 30/73 (41%), Positives = 44/73 (60%)
Frame = -2
Query: 1018 DIFVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILF 1077
DI + T +L +YLG PIF T FDF++D+VN++F+ WK LL+ G+ L
Sbjct: 291 DISSGLGLQSTDDLGKYLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLT 112
Query: 1078 KLVLNAIPVYPMQ 1090
K V+ A+P + MQ
Sbjct: 111 K*VV*ALPSHIMQ 73
>CO982349
Length = 795
Score = 128 bits (322), Expect = 2e-29
Identities = 87/262 (33%), Positives = 141/262 (53%), Gaps = 11/262 (4%)
Frame = +3
Query: 880 FGFPP-TSVS----LIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVER 934
F FPP T++ I C+ S+S+S+L N + +P RGLR GDPL+P LF + E
Sbjct: 27 FYFPP*TAIRH*QHWIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEG 206
Query: 935 LTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMAS 994
LT +++AVD+ + + ++ P+S L +ADD + +A + ++ ++ IL F +AS
Sbjct: 207 LTGLMREAVDRKRFNSFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELAS 386
Query: 995 GLNVNIAKSRDFSSVSVPR--AKKADIFV---VTSIPFTANLERYLGFPIFQGRQTQEHF 1049
GL +N A+SR F ++ P K+A F+ + S+PF+ YLG PI + +E
Sbjct: 387 GLKINFAESR-FGAIWKPDHWCKEAAEFLNCSMLSMPFS-----YLGIPIEANPRCREI* 548
Query: 1050 DFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNF 1109
D ++ + K A WK R ++ G+ L +L A+P+Y F V L ++ + F
Sbjct: 549 DPIIRKCETKLARWKQRHISLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKF 728
Query: 1110 IWSRGE-RRGMNMVAWRTVTLP 1130
+W G +R + V W TV LP
Sbjct: 729 LWGGGSXQRRIAWVKWETVCLP 794
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 61.2 bits (147), Expect(2) = 2e-17
Identities = 39/137 (28%), Positives = 67/137 (48%), Gaps = 5/137 (3%)
Frame = -2
Query: 961 SHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADI- 1019
SH+ ADD+++ K V + N + + G ++ KS+ F S+P + I
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIG-SIPGQRLRVIS 275
Query: 1020 ----FVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFI 1075
+ + IPF Y G PIF+G+ T+ H V D++ K ASWKG +L+ G+
Sbjct: 274 DLLQYFHSDIPFN-----YFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQ 110
Query: 1076 LFKLVLNAIPVYPMQLF 1092
L V++ + +Y ++
Sbjct: 109 LVNSVIHGMLLYSFSVY 59
Score = 47.8 bits (112), Expect(2) = 2e-17
Identities = 35/110 (31%), Positives = 57/110 (51%), Gaps = 1/110 (0%)
Frame = -3
Query: 831 TIDNAIILQKIVHHMNSRRGKCQNVVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLI 890
TI + + V +M ++ N+ K+D+ KA ++ FL L FG+ T + I
Sbjct: 837 TIKDCTCVTSEVINMLDKKVFGGNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWI 658
Query: 891 MFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLF-VLCVERLTISI 939
+ S+ LS+ NG + + RG+R GDPLSP L+ C+E+ TI +
Sbjct: 657 RVILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLYRQRCLEQGTIYV 508
>AW757033
Length = 441
Score = 87.8 bits (216), Expect = 3e-17
Identities = 56/147 (38%), Positives = 87/147 (59%), Gaps = 5/147 (3%)
Frame = -2
Query: 912 APSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLL 971
+P RGLR GDPL+P LF + E LT +++AV +N + + + P+S L +ADD +
Sbjct: 428 SPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTIF 249
Query: 972 LAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPR--AKKADIFV---VTSIP 1026
+A + VR ++ IL F +ASGL +N AKSR F ++SVP K+A F+ + S+P
Sbjct: 248 FGEATMENVRVIKTILRGFELASGLKINFAKSR-FGAISVPD*WRKEAAEFMNCSLLSLP 72
Query: 1027 FTANLERYLGFPIFQGRQTQEHFDFVL 1053
F+ YLG PI + +E +D ++
Sbjct: 71 FS-----YLGIPIGANPRRRETWDPII 6
>TC211008
Length = 516
Score = 82.8 bits (203), Expect = 1e-15
Identities = 42/101 (41%), Positives = 60/101 (58%)
Frame = +1
Query: 1467 GGVVRDAVGCWLFGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLL 1526
GG+VRD+ GC+L G++ +G + AEL V+ GL+ WD G +++ V DS L L+
Sbjct: 154 GGLVRDSSGCYLGGFTVNLGNTSVTLAELWGVVHGLKLAWDLGCKKVKVDIDSGNALGLV 333
Query: 1527 RHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
RHG A+V I EL+R++W VE H+ RE N AD
Sbjct: 334 RHGPVANDPAFALVSEINELVRKEWLVEFSHVFRESNRAAD 456
>BI315739
Length = 442
Score = 50.1 bits (118), Expect(2) = 1e-13
Identities = 31/76 (40%), Positives = 43/76 (55%)
Frame = +2
Query: 960 ISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADI 1019
ISHLFF D+ LL KA SQV+ V+++L FC+ GL + KSR + +V K A
Sbjct: 164 ISHLFFPDNCLLFVKATSSQVKLVRDVLQQFCLT*GLKAGLQKSRFLA*KNVSNRKIARF 343
Query: 1020 FVVTSIPFTANLERYL 1035
+ I + +N ER L
Sbjct: 344 SFIIVISW-SNQERGL 388
Score = 45.8 bits (107), Expect(2) = 1e-13
Identities = 21/43 (48%), Positives = 31/43 (71%)
Frame = +3
Query: 922 PLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLF 964
PLSPYLF +C+E+L I IQ V++ E ++ISR+G IS ++
Sbjct: 48 PLSPYLF*ICMEKLAILIQSKVNEETWEPIKISRNGPGISPIY 176
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase homolog
T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 67.8 bits (164), Expect = 4e-11
Identities = 41/123 (33%), Positives = 62/123 (50%), Gaps = 8/123 (6%)
Frame = +2
Query: 1102 LDSLAKNFIWS-RGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSH 1160
L+SL + F+W + R + V W+TV LPK GGLGI+ R NT++LGK WDL
Sbjct: 41 LESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTLLGKWRWDLFYIQ 220
Query: 1161 DKLWV*VLLDLYCTNSDFLH-APRRRGSCVWNAVMKTRDI-----IQDGFTWRIGRGD-L 1213
+ W VL Y + + S W ++KT+ + ++ W++G GD +
Sbjct: 221 QEPWAKVLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQQLQRNIPLKRETIWKVGGGDRI 400
Query: 1214 SFW 1216
FW
Sbjct: 401 KFW 409
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 63.2 bits (152), Expect = 9e-10
Identities = 29/94 (30%), Positives = 51/94 (53%), Gaps = 1/94 (1%)
Frame = +2
Query: 1072 GKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSRGERRG-MNMVAWRTVTLP 1130
G+ L K VL+A+P+ + F + D L +L + F+W + + V W + P
Sbjct: 8 GRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICNP 187
Query: 1131 KCNGGLGIRKARLINTSMLGKLVWDLVNSHDKLW 1164
K +GGLGI+ N ++ G+ +W L ++H++LW
Sbjct: 188 KIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLW 289
>BI972659
Length = 453
Score = 62.4 bits (150), Expect = 2e-09
Identities = 44/158 (27%), Positives = 76/158 (47%), Gaps = 5/158 (3%)
Frame = +1
Query: 972 LAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTS-----IP 1026
+ +A + ++++L F M SGL +N AKS+ F V D + + IP
Sbjct: 1 VGEASWDNIIVLKSMLRGFEMVSGLRINYAKSQ-FGIVGFQPNWAHDAAQLLNCRQLDIP 177
Query: 1027 FTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPV 1086
F YLG PI ++ ++ ++++ K + W + L+ G+ L K VLNA+P+
Sbjct: 178 F-----HYLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPI 342
Query: 1087 YPMQLFWFH*AVCDHLDSLAKNFIWSRGERRGMNMVAW 1124
Y + F + D L SL + F+W G + N ++W
Sbjct: 343 YLLSFFKIPQRIVDKLVSLQRTFMW--GGNQHHNRISW 450
>TC216761 similar to UP|Q9FQE4 (Q9FQE4) Glutathione S-transferase GST 14
(Fragment) , complete
Length = 1353
Score = 60.5 bits (145), Expect = 6e-09
Identities = 29/62 (46%), Positives = 37/62 (58%)
Frame = +1
Query: 1506 WDRGFRQLVVSSDSMVVLSLLRHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVV 1565
W RG R L+ SDS + L L+ GV H AA+V IR + +DW + +H LREGN V
Sbjct: 1024 WTRGIRNLICESDSSMALQLISTGVYITHPRAALVTTIRSFMDKDWCLSFQHTLREGNFV 1203
Query: 1566 AD 1567
AD
Sbjct: 1204 AD 1209
>TC213061
Length = 823
Score = 57.4 bits (137), Expect = 5e-08
Identities = 33/104 (31%), Positives = 47/104 (44%), Gaps = 1/104 (0%)
Frame = +2
Query: 1278 DDEDLVVWAPSLEGKYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHR 1337
D ED VWA G Y +S YR + + ++ LWK +VP K+ +FAW+
Sbjct: 119 DIEDQWVWAAESSGSYLARSAYRVIREGIPEEEQDREFKELWKLKVPMKVTMFAWRLLKD 298
Query: 1338 SLPTVDTFHDRGVAIPRW-CSLCHHHVET*MHCLRDCDASLQVW 1380
LPT D + V + + C LC E+ H C L +W
Sbjct: 299 ILPTRDNLRRKRVELHEYVCPLCRSMDESASHLFFHCSKILPIW 430
>TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leishmania major;}
, partial (8%)
Length = 1386
Score = 57.0 bits (136), Expect = 6e-08
Identities = 38/106 (35%), Positives = 57/106 (52%), Gaps = 2/106 (1%)
Frame = +2
Query: 1464 AEVGGVVRDAVGCWLFGYSGFIGMA-EILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVV 1522
A GGV+RDA WL G++ + + + EL A+L GL+ + ++L+V SDS V
Sbjct: 560 AGCGGVLRDASAKWLRGFAKKLNPTYAVHQTELEAILTGLKVASEMNVKKLIVESDSDSV 739
Query: 1523 LSLLRHGVTRFHVYAAIVGAIRELLRR-DWRVELKHMLREGNVVAD 1567
+S++ +GV H +V IR RR DW V + + N VAD
Sbjct: 740 VSMVENGVKPNHPDYGVVELIRTKRRRFDWEVRFVSVSNKANRVAD 877
>AI496436
Length = 414
Score = 56.6 bits (135), Expect = 8e-08
Identities = 41/123 (33%), Positives = 63/123 (50%), Gaps = 5/123 (4%)
Frame = +1
Query: 986 ILADFCMASGLNVNIAKSRDFSSV---SVPRAKKADIF--VVTSIPFTANLERYLGFPIF 1040
IL F ++SGL +N AKS+ F ++ + R + AD V S+PF YLG PI
Sbjct: 4 ILRCFELSSGLKINFAKSQ-FGTICKSDIWRKEAADYLNSSVLSMPFV-----YLGIPIG 165
Query: 1041 QGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCD 1100
+ + ++ V+ + RK ASWK + ++ G+ L VL A+P+Y + F V D
Sbjct: 166 ANSRHSDVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVVD 345
Query: 1101 HLD 1103
D
Sbjct: 346 KAD 354
>CO979823
Length = 853
Score = 56.6 bits (135), Expect = 8e-08
Identities = 32/102 (31%), Positives = 46/102 (44%), Gaps = 1/102 (0%)
Frame = -2
Query: 1280 EDLVVWAPSLEGKYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRSL 1339
ED +W P G YS +S Y L D+A +WK ++P K +FAW+ L
Sbjct: 705 EDKWLWKPEPGGHYSTKSGYHVLWGELTEEIQDADFAEIWKLKIPTKAAVFAWRLVRDRL 526
Query: 1340 PTVDTFHDRGVAIPRW-CSLCHHHVET*MHCLRDCDASLQVW 1380
PT R V + C LC++ E H +C +L +W
Sbjct: 525 PTKSNLRRRQVMVQDMVCPLCNNIEEGAAHLFFNCTKTLPLW 400
>TC212765
Length = 637
Score = 55.1 bits (131), Expect = 2e-07
Identities = 40/146 (27%), Positives = 63/146 (42%)
Frame = +3
Query: 925 PYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQ 984
PYLFVLC+ERL + I D Q + +S D + +L DD+ L A
Sbjct: 15 PYLFVLCMERLALCIHDLTSQ*IWRPILVSDDNRLVPYLMLVDDIFLFINAIGD*AYLAS 194
Query: 985 NILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQGRQ 1044
L +F GL +N+ KS + S + + I +I +L +YL GR
Sbjct: 195 LTL*NFFQTFGLKLNLDKSSMYCS*RMAPVLRDLISNTLNIVHAPSLGKYLCLKFIHGRS 374
Query: 1045 TQEHFDFVLDRVNRKFASWKGRLLNK 1070
F +++R++ + +G L K
Sbjct: 375 KIVDF-LLVERIHNPWPRGRGSFLTK 449
Score = 52.8 bits (125), Expect = 1e-06
Identities = 23/53 (43%), Positives = 35/53 (65%)
Frame = +2
Query: 1061 ASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSR 1113
ASWKG+LLN+A K L LVL+A P+YPM+ W ++ +D ++ IW++
Sbjct: 419 ASWKGKLLNQASKHCLT*LVLHAFPIYPMEASWLPQSIFQAIDKASRVTIWNK 577
>TC211809
Length = 895
Score = 53.1 bits (126), Expect = 9e-07
Identities = 30/95 (31%), Positives = 43/95 (44%), Gaps = 1/95 (1%)
Frame = +2
Query: 1287 PSLEGKYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFH 1346
P+L G YS +S YR L++ G V + LW ++P K +FAW+ LPT
Sbjct: 188 PTLFGNYSTKSAYRLLMEATGEFPVDRTYVDLWNLKIPSKAIVFAWRLIKDRLPTWTNLR 367
Query: 1347 DRGVAI-PRWCSLCHHHVET*MHCLRDCDASLQVW 1380
R V + C LC+ E H C + +W
Sbjct: 368 VRQVELNDSRCPLCNSSEEDAAHLFFHCTKTEXLW 472
>CO984521
Length = 716
Score = 52.4 bits (124), Expect = 2e-06
Identities = 30/97 (30%), Positives = 42/97 (42%), Gaps = 1/97 (1%)
Frame = -2
Query: 1285 WAPSLEGKYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRSLPTVDT 1344
WA G YS +S Y+ L + G + LWK +VP K+ +FAW+ LPT
Sbjct: 634 WAAEPSGCYSTKSAYKALHHVTVGEEQDGKFKELWKLRVPLKVAIFAWRLIQDKLPTKAN 455
Query: 1345 FHDRGVAIPRW-CSLCHHHVET*MHCLRDCDASLQVW 1380
+ V + + C LC ET H C +W
Sbjct: 454 LRKKRVELQEYLCPLCRSVEETASHLFFHCSKVSPLW 344
>TC234877
Length = 534
Score = 52.0 bits (123), Expect = 2e-06
Identities = 31/111 (27%), Positives = 52/111 (45%)
Frame = +1
Query: 1020 FVVTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKL 1079
F V S+PF YL P+FQ + H + DR+ + SWKG LL+ G L
Sbjct: 22 FSVGSLPFI-----YLSVPLFQEKPRPVHLRPIADRIKAELQSWKGNLLSFMGCVQLIIS 186
Query: 1080 VLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSRGERRGMNMVAWRTVTLP 1130
+++ + + L+ + A+ + + +NFIW+RG + W +P
Sbjct: 187 IIDGMLL*SFHLYSWPKALLIEMQTWTRNFIWTRGYFKEKTGYCWLKNCVP 339
>BG404993
Length = 412
Score = 50.8 bits (120), Expect = 5e-06
Identities = 29/103 (28%), Positives = 44/103 (42%), Gaps = 1/103 (0%)
Frame = -3
Query: 1279 DEDLVVWAPSLEGKYSVQSCYRWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRS 1338
+ D W G YS ++ Y L + G + LWK ++P K +F W+ +
Sbjct: 338 EADSWTWKADPSGNYSTKTAYELLQEAIRGDNEDGIFVQLWKLKIPPKAPVFTWRLINDR 159
Query: 1339 LPTVDTFHDRGVAI-PRWCSLCHHHVET*MHCLRDCDASLQVW 1380
LPT R V I C LC++ E H +C+ +L W
Sbjct: 158 LPTKVNLRRRQVEISDSLCPLCNNSEEDAAHLFFNCNKTLPFW 30
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.338 0.147 0.507
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 87,917,773
Number of Sequences: 63676
Number of extensions: 1552623
Number of successful extensions: 11990
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 11623
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11967
length of query: 1569
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1459
effective length of database: 5,635,272
effective search space: 8221861848
effective search space used: 8221861848
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0022.23


