

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.17
(341 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
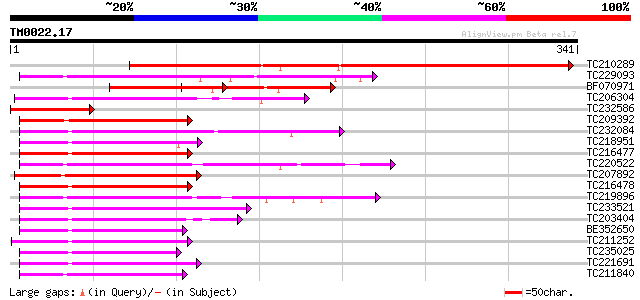
Score E
Sequences producing significant alignments: (bits) Value
TC210289 similar to UP|Q9FUK9 (Q9FUK9) MYB-related transcription... 345 2e-95
TC229093 homologue to PIR|T51621|T51621 myb-like protein [import... 112 2e-25
BF070971 similar to GP|14719883|gb myb-related transcription fac... 109 2e-24
TC206304 similar to GB|AAS10028.1|41619120|AY519558 MYB transcri... 101 6e-22
TC232586 homologue to UP|Q9FUK9 (Q9FUK9) MYB-related transcripti... 99 4e-21
TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb... 96 2e-20
TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragmen... 95 5e-20
TC218951 homologue to UP|P93474 (P93474) Myb26, partial (77%) 92 4e-19
TC216477 homologue to UP|P93474 (P93474) Myb26, partial (67%) 92 4e-19
TC220522 similar to UP|O49020 (O49020) MYB-like DNA-binding doma... 91 6e-19
TC207892 homologue to UP|Q9FGY3 (Q9FGY3) Myb-related transcripti... 91 1e-18
TC216478 similar to UP|P93474 (P93474) Myb26, partial (83%) 91 1e-18
TC219896 myb 91 1e-18
TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%) 90 2e-18
TC203404 GmMYB16 90 2e-18
BE352650 89 2e-18
TC211252 similar to UP|Q9ATD3 (Q9ATD3) GHMYB36, partial (37%) 89 3e-18
TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor,... 89 4e-18
TC221691 GmMYB6 86 2e-17
TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4,... 86 3e-17
>TC210289 similar to UP|Q9FUK9 (Q9FUK9) MYB-related transcription factor
PHAN1, partial (68%)
Length = 1009
Score = 345 bits (885), Expect = 2e-95
Identities = 188/280 (67%), Positives = 216/280 (77%), Gaps = 13/280 (4%)
Frame = +2
Query: 73 LQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKYEHMLEG 132
LQA YGNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQREK ++N I PI+++KYEH+LE
Sbjct: 5 LQAKYGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREKKDVNRISDPINNSKYEHILES 184
Query: 133 FAEKLVKEHTLPSFAMAASSNEAFLHTNS----SAMLPSWLSNYDSTSTP-PSSISVTLS 187
FAEKLVKE PSF MAAS N AFLHT++ S++LPSWLSN S + PSS+SVTLS
Sbjct: 185 FAEKLVKERPSPSFLMAAS-NGAFLHTDTPAPASSLLPSWLSNSSSPAADGPSSLSVTLS 361
Query: 188 LSPSTVATP--------RGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEG 239
LS S V P RG +NAPFVL N A G +P+ SD++LMS++V KELEEG
Sbjct: 362 LSSSMVIAPPFSWLPPKRG-PDNAPFVLGNAAALQGVIPTLSDNMLMSQMVEHRKELEEG 538
Query: 240 HRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACR 299
HRALA HKKEA WRL R+ELQLESEKA RRRE +EEFEA IKALQEE+ AAL RIE R
Sbjct: 539 HRALATHKKEAAWRLSRVELQLESEKANRRREKIEEFEAKIKALQEEEKAALGRIEAEYR 718
Query: 300 EQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQTG 339
EQL LRRDAE+KEQKLAE+W +KHLR TRLLEQ+ + G
Sbjct: 719 EQLAALRRDAENKEQKLAEQWDAKHLRFTRLLEQLGCRAG 838
>TC229093 homologue to PIR|T51621|T51621 myb-like protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(44%)
Length = 964
Score = 112 bits (281), Expect = 2e-25
Identities = 76/251 (30%), Positives = 125/251 (49%), Gaps = 36/251 (14%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+Q++GP W V NT L+R +KSC RW NYL+PGIK+G+ T +E
Sbjct: 198 WTPEEDIILVSYIQEHGPGNWKAVPA--NTGLSRCSKSCRLRWTNYLRPGIKRGNFTDQE 371
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIE------------- 113
++++I LQA GN+W IAA +P RT + +W Y +K+ +K+E
Sbjct: 372 EKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLNKKLEAGSDQGSLLGGGH 551
Query: 114 ----INGIVSPISDTKYEHMLE---GFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLP 166
+ + P+S ++E L+ A+K + E P+ A ++SS+ +NS+
Sbjct: 552 NNIGFSSVSQPMSRGQWERRLQTDIHMAKKALSEALSPNKA-SSSSSTLLAESNSNLSNE 728
Query: 167 SWLSNYDSTSTPPSSISVTLSLSPSTVA---------TPRGLENNAPFVLRN-------V 210
+ +T+ P ++ S+ + S +A TP+ A V +N +
Sbjct: 729 NSTFYNSNTNKPTTTQSMCYASSADNIARLLKGWMKNTPKSASCAASAVTQNSCSSSEGI 908
Query: 211 TAHNGSVPSFS 221
T GS S S
Sbjct: 909 TTTKGSTTSTS 941
>BF070971 similar to GP|14719883|gb myb-related transcription factor
{Medicago truncatula}, partial (39%)
Length = 424
Score = 109 bits (273), Expect = 2e-24
Identities = 55/75 (73%), Positives = 61/75 (81%), Gaps = 4/75 (5%)
Frame = +2
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA YGNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQREK EIN I P
Sbjct: 2 SLTEEEQRLVIHLQAKYGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREKKEINRIARP 181
Query: 121 ----ISDTKYEHMLE 131
+S + Y +L+
Sbjct: 182 NLTIVSTSTYSRVLQ 226
Score = 84.0 bits (206), Expect = 9e-17
Identities = 52/98 (53%), Positives = 66/98 (67%), Gaps = 5/98 (5%)
Frame = +1
Query: 104 KEKQQREKIEINGIVSPISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTN--- 160
+E +REK + + +++KYEH+LE FAEKLVKE PSF MAAS AFL T+
Sbjct: 133 REAAEREKGDQ*NCPTQFNNSKYEHILESFAEKLVKERPSPSFVMAASDG-AFLLTDTPA 309
Query: 161 -SSAMLPSWLSNYDSTST-PPSSISVTLSLSPSTVATP 196
+S++ PSWLSN S + PSS+SV LSLS STVATP
Sbjct: 310 PASSLRPSWLSNSSSAAAIGPSSLSVKLSLSSSTVATP 423
>TC206304 similar to GB|AAS10028.1|41619120|AY519558 MYB transcription factor
{Arabidopsis thaliana;} , partial (46%)
Length = 1158
Score = 101 bits (251), Expect = 6e-22
Identities = 62/181 (34%), Positives = 96/181 (52%), Gaps = 4/181 (2%)
Frame = +3
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+ EED LL Y+ +G WNL+++R L R KSC RW NYLKP +K+G+LT
Sbjct: 60 RGPWTLEEDNLLSQYIFNHGEGRWNLLAKRSG--LKRTGKSCRLRWLNYLKPDVKRGNLT 233
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE-KIEINGIVSPIS 122
+EQ +++ L + +GN+W KIA +PGRT + +W +KQ R KI+ +
Sbjct: 234 PQEQLIILELHSKWGNRWSKIAQHLPGRTDNEIKNYWRTRIQKQARHLKID--------T 389
Query: 123 DTKYEHMLEGFAEKLVKEHTLPSFAMAA--SSNEAFLHTNSSAMLP-SWLSNYDSTSTPP 179
D++ ++LV+ +P A SS+ A N + +P +S + + T P
Sbjct: 390 DSRE-------FQELVRRFWMPRLLQKAKESSSSAMSIQNQATPMPFDGVSQHSTVGTIP 548
Query: 180 S 180
S
Sbjct: 549 S 551
>TC232586 homologue to UP|Q9FUK9 (Q9FUK9) MYB-related transcription factor
PHAN1, partial (15%)
Length = 481
Score = 98.6 bits (244), Expect = 4e-21
Identities = 44/51 (86%), Positives = 47/51 (91%)
Frame = +1
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKN 51
MKERQRW +EEDALL +YV+QYGPREWNLVSQRMNT LNRD KSCLERWKN
Sbjct: 328 MKERQRWRAEEDALLRSYVKQYGPREWNLVSQRMNTYLNRDAKSCLERWKN 480
>TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb protein
(Fragment), partial (59%)
Length = 866
Score = 96.3 bits (238), Expect = 2e-20
Identities = 42/104 (40%), Positives = 69/104 (65%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+Q++GP W V + T L+R +KSC RW NYL+PGIK+G+ T++E
Sbjct: 477 WTPEEDIILVSYIQEHGPGNWRAVPAK--TGLSRCSKSCRLRWTNYLRPGIKRGNFTEQE 650
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
++++I LQ GN+W IA+ +P RT + +W + K+ ++
Sbjct: 651 EKMIIHLQDLLGNRWAAIASYLPQRTDNDIKNYWNTHLRKKLKK 782
>TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragment), complete
Length = 1383
Score = 94.7 bits (234), Expect = 5e-20
Identities = 58/200 (29%), Positives = 97/200 (48%), Gaps = 5/200 (2%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+Q++G W + + LNR KSC RW NYL+P IK+G ++EE
Sbjct: 101 WTPEEDRILVDYIQKHGHGSWRALPKHAR--LNRCGKSCRLRWTNYLRPDIKRGKFSEEE 274
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
++L+I L A GNKW IA +PGRT + +W + +K+ + P SD +
Sbjct: 275 EQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLLQMGLDPVTHRPRSD--H 448
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSW-----LSNYDSTSTPPSS 181
++L + L + + + +N A + + L + + +TP S+
Sbjct: 449 LNLLSNLQQLLAATNIVSNLTNTWDTNNALRLQSDATQLAKLQLLQNILHVLGDTTPTSN 628
Query: 182 ISVTLSLSPSTVATPRGLEN 201
+ + PS+ + P G N
Sbjct: 629 LDLLNPFGPSSSSLPDGFLN 688
>TC218951 homologue to UP|P93474 (P93474) Myb26, partial (77%)
Length = 866
Score = 91.7 bits (226), Expect = 4e-19
Identities = 44/112 (39%), Positives = 67/112 (59%), Gaps = 2/112 (1%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+ +G WN +++ L R KSC RW NYL+P +++G++T EE
Sbjct: 63 WTMEEDLILINYIANHGEGVWNSLAKASG--LKRTGKSCRLRWLNYLRPDVRRGNITPEE 236
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWW--EVYKEKQQREKIEING 116
Q L+I L A +GN+W KIA +PGRT + +W + K +Q E + +G
Sbjct: 237 QLLIIELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKHIKQAETSQQHG 392
>TC216477 homologue to UP|P93474 (P93474) Myb26, partial (67%)
Length = 1001
Score = 91.7 bits (226), Expect = 4e-19
Identities = 40/104 (38%), Positives = 63/104 (60%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+ +G WN +++ L R KSC RW NYL+P +++G++T EE
Sbjct: 83 WTMEEDLILITYIANHGEGVWNSLAKAAG--LKRTGKSCRLRWLNYLRPDVRRGNITPEE 256
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
Q L++ L A +GN+W KIA +PGRT + +W +K ++
Sbjct: 257 QLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNYWRTRIQKHLKQ 388
>TC220522 similar to UP|O49020 (O49020) MYB-like DNA-binding domain protein,
partial (51%)
Length = 947
Score = 91.3 bits (225), Expect = 6e-19
Identities = 60/229 (26%), Positives = 109/229 (47%), Gaps = 3/229 (1%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED++L YV + WN +++ ++ L R KSC RW NYL+P +++G++T +E
Sbjct: 82 WTEEEDSVLINYVNVHSEGHWNSLAR--SSGLKRTGKSCRLRWFNYLRPNVRRGNITLQE 255
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
Q L++ L +++GN+W KIA ++PGRT + +W KQ ++ + ++ ++
Sbjct: 256 QLLILELHSHWGNRWAKIAEQLPGRTDNEIKNYWRTRVVKQAKQ------LKCHVNSKQF 417
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNS---SAMLPSWLSNYDSTSTPPSSIS 183
L + E S + +T + ++M+ S+ S D PPS
Sbjct: 418 RDALRFVWMPRLMEQIQASSSSYGLDQTTLCNTQTHRDNSMVSSYSSEVD--LQPPSLSD 591
Query: 184 VTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGF 232
+++ S + GL +A GS+ S H S++ F
Sbjct: 592 TSITSSSYNLIGDGGLSTE--------SAEKGSIYSLWQHWDYSDIQAF 714
>TC207892 homologue to UP|Q9FGY3 (Q9FGY3) Myb-related transcription factor
(MYB transcription factor), partial (48%)
Length = 1082
Score = 90.5 bits (223), Expect = 1e-18
Identities = 42/113 (37%), Positives = 68/113 (60%), Gaps = 1/113 (0%)
Frame = +1
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+ +ED L YV +G WN ++ ++ L R KSC RW NYL+P +++G++T
Sbjct: 184 RGPWTVDEDLTLINYVATHGEGRWNTLA--LSAGLKRTGKSCRLRWLNYLRPDVRRGNIT 357
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE-KIEIN 115
EEQ L++ L + +GN+W KIA +PGRT + +W +K ++ K ++N
Sbjct: 358 LEEQLLILELHSRWGNRWSKIAQYLPGRTDNEIKNYWRTRVQKHAKQLKCDVN 516
>TC216478 similar to UP|P93474 (P93474) Myb26, partial (83%)
Length = 925
Score = 90.5 bits (223), Expect = 1e-18
Identities = 40/104 (38%), Positives = 63/104 (60%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+ +G WN +++ L R KSC RW NYL+P +++G++T EE
Sbjct: 115 WTMEEDLILINYIANHGEGVWNSLAKAAG--LKRTGKSCRLRWLNYLRPDVRRGNITPEE 288
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
Q L++ L A +GN+W KIA +PGRT + +W +K ++
Sbjct: 289 QLLIMELHAKWGNRWSKIAKHLPGRTDNEIKNYWRTRIQKHIKQ 420
>TC219896 myb
Length = 842
Score = 90.5 bits (223), Expect = 1e-18
Identities = 68/228 (29%), Positives = 102/228 (43%), Gaps = 11/228 (4%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WS EED L YV +G +W V+Q N L R KSC +RW NYLKPGIK+G ++ +E
Sbjct: 24 WSREEDETLSKYVSIHGEGKWQKVAQ--NAGLKRCGKSCRQRWLNYLKPGIKRGHISVDE 197
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
+ ++I L GN+W IA +PGRT + +W ++ ++ VS + ++
Sbjct: 198 EDMIIRLHRLLGNRWALIAKRLPGRTDNEIKNYWNTNLSRKLQK--HPTSSVSSLQHKRH 371
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAASSN----EAFLHTNSSAMLPSWL---SNYDSTSTPP 179
E EK + H P N L PS SN + S+
Sbjct: 372 E------KEKTKQMHVAPEAPRRRRVNAVEYSKNLENGGCGNRPSTTPSPSNKEEGSSEE 533
Query: 180 SSISVTL----SLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDH 223
+S S L + S P+ E++ + +T ++ S S SDH
Sbjct: 534 ASFSDFLMDIDQIDDSNSKVPQKEEDHHHNKVELITNNSPSSSSLSDH 677
>TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%)
Length = 886
Score = 89.7 bits (221), Expect = 2e-18
Identities = 44/139 (31%), Positives = 74/139 (52%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++EED +L +++Q+YG W + ++ L R KSC RW NYL+P IK+G +KEE
Sbjct: 68 WTAEEDQILVSHIQRYGHGNWRALPKQAG--LLRCGKSCRLRWINYLRPDIKRGKFSKEE 241
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
+ ++ L GN+W IAA +PGRT + +W + ++ Q+ + S I
Sbjct: 242 EDTILKLHGILGNRWSAIAASLPGRTDNEIKNFWHTHLKRIQKSGVHNGNPSSRILQEAQ 421
Query: 127 EHMLEGFAEKLVKEHTLPS 145
+ + ++ + LPS
Sbjct: 422 ANTSSNASSVMIANYGLPS 478
>TC203404 GmMYB16
Length = 1138
Score = 89.7 bits (221), Expect = 2e-18
Identities = 48/134 (35%), Positives = 72/134 (52%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ +EDALL Y+Q +G +W + ++ L R KSC RW NYL+P IK+G++ EE
Sbjct: 154 WTPKEDALLTKYIQAHGEGQWKSLPKKAG--LLRCGKSCRLRWMNYLRPDIKRGNIAPEE 327
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
L+I + + GN+W IA +PGRT + +W + K K++I G DT
Sbjct: 328 DDLIIRMHSLLGNRWSLIAGRLPGRTDNEIKNYWNTHLSK----KLKIQG----TEDTDT 483
Query: 127 EHMLEGFAEKLVKE 140
MLE E+ +
Sbjct: 484 HKMLENPQEEAASD 525
>BE352650
Length = 528
Score = 89.4 bits (220), Expect = 2e-18
Identities = 40/101 (39%), Positives = 61/101 (59%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L Y+ ++G W + +R LNR KSC RW NYL+P IK+G ++++
Sbjct: 154 WTPEEDEKLIDYISKHGHGSWRTLPKRAG--LNRCGKSCRLRWTNYLRPDIKRGKFSEDD 327
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+R++I + GNKW KIAA +PGRT + +W + K+
Sbjct: 328 ERIIINFHSVLGNKWSKIAAHLPGRTDNEIKNYWTTHIRKK 450
>TC211252 similar to UP|Q9ATD3 (Q9ATD3) GHMYB36, partial (37%)
Length = 819
Score = 89.0 bits (219), Expect = 3e-18
Identities = 42/109 (38%), Positives = 62/109 (56%)
Frame = +1
Query: 2 KERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGS 61
++ + WSS ED +L YVQ G W + +R L R +SC +RW NYLKP I +G+
Sbjct: 22 EKEEAWSSHEDEILLNYVQVRGEGNWRNLPKRAG--LKRCGESCKQRWLNYLKPTISRGN 195
Query: 62 LTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
++ +E L+I L GN+W IA +PGRT + + +W Y K+ E
Sbjct: 196 ISLDEHELIIRLHKLLGNRWSIIAGRLPGRTEEEIKNYWNTYLRKEAEE 342
>TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor, partial
(35%)
Length = 434
Score = 88.6 bits (218), Expect = 4e-18
Identities = 40/97 (41%), Positives = 58/97 (59%)
Frame = +1
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L Y+ ++G W + +R LNR KSC RW NYL P IK+G ++E+
Sbjct: 148 WTPEEDEKLIDYISKHGHGSWRTLPKRAG--LNRCGKSCRLRWTNYLTPDIKRGKFSEED 321
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVY 103
+R++I L + GNKW KIA +PGRT + +W +
Sbjct: 322 ERIIINLHSVLGNKWSKIATHLPGRTDNEIKNYWNTH 432
>TC221691 GmMYB6
Length = 590
Score = 85.9 bits (211), Expect = 2e-17
Identities = 42/109 (38%), Positives = 59/109 (53%)
Frame = +3
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED L AYV +YG W + + L R KSC RW NYL+P +K+G+ T++E
Sbjct: 126 WTPEEDRKLIAYVTRYGSWNWRQLPRFAG--LARCGKSCRLRWMNYLRPNVKRGNFTQQE 299
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEIN 115
+I + GNKW IAAE+PGRT + W +K ++ N
Sbjct: 300 DECIIRMHKKLGNKWSAIAAELPGRTDNEIKNHWHTTLKKWSQQNAITN 446
>TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4, partial
(38%)
Length = 516
Score = 85.5 bits (210), Expect = 3e-17
Identities = 40/101 (39%), Positives = 59/101 (57%)
Frame = +2
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++EED L Y+Q++G W + + N L R KSC RW NYL+P IK+G + EE
Sbjct: 11 WTTEEDQKLIDYIQKHGYGNWRTLPK--NAGLQRCGKSCRLRWTNYLRPDIKRGRFSFEE 184
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+ +I L + GNKW IA+ +PGRT + +W + K+
Sbjct: 185 EETIIQLHSILGNKWSAIASRLPGRTDNEIKNYWNTHIRKR 307
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.127 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,129,669
Number of Sequences: 63676
Number of extensions: 203511
Number of successful extensions: 1501
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 1409
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1438
length of query: 341
length of database: 12,639,632
effective HSP length: 98
effective length of query: 243
effective length of database: 6,399,384
effective search space: 1555050312
effective search space used: 1555050312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0022.17


