

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.15
(313 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
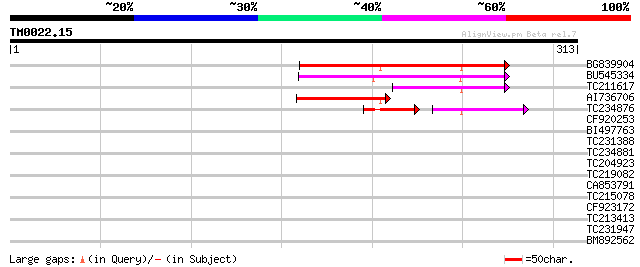
Score E
Sequences producing significant alignments: (bits) Value
BG839904 116 1e-26
BU545334 85 4e-17
TC211617 61 6e-10
AI736706 47 9e-06
TC234876 similar to UP|Q7RNR5 (Q7RNR5) 60S ribosomal protein L27... 35 4e-04
CF920253 37 0.009
BI497763 37 0.015
TC231388 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At... 30 1.1
TC234881 29 3.1
TC204923 similar to GB|AAO39903.1|28372842|BT003675 At5g67590 {A... 29 3.1
TC219082 similar to UP|Q9LYS5 (Q9LYS5) Receptor-like protein kin... 28 4.1
CA853791 28 5.4
TC215078 similar to UP|O80439 (O80439) 30S ribosomal protein S31... 28 5.4
CF923172 28 7.0
TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 28 7.0
TC231947 similar to UP|Q84NI9 (Q84NI9) Myb51 protein, partial (13%) 27 9.2
BM892562 27 9.2
>BG839904
Length = 712
Score = 116 bits (291), Expect = 1e-26
Identities = 57/127 (44%), Positives = 82/127 (63%), Gaps = 11/127 (8%)
Frame = -3
Query: 161 DYTQSFTTDKVFASRDEILEWARNLGKQHGFIIVITRSDNGGL--------KRKTFMILG 212
D + +F T +VF SR+ +L WAR++ K++GF+++I RS+ +RKTF+I+G
Sbjct: 518 DCSDAFNTTEVFPSREAMLNWARDVAKENGFVLIILRSETSTTCTRSTRCNQRKTFVIMG 339
Query: 213 CERCDKYI-PYKEVLKHQSTGTKKCYCPFRLRARGTK--VGWKVSVMNGYHIHERSETLL 269
C+R KY PYK L + +GT+KC CPF+L+ + K GW V VM G H H+ ETL+
Sbjct: 338 CDRSGKYRGPYKNELSRKVSGTRKCECPFKLKGKALKKAEGWIVKVMCGCHNHDLXETLV 159
Query: 270 GHPYVGR 276
GHPY GR
Sbjct: 158 GHPYAGR 138
>BU545334
Length = 556
Score = 85.1 bits (209), Expect = 4e-17
Identities = 44/121 (36%), Positives = 73/121 (59%), Gaps = 4/121 (3%)
Frame = -1
Query: 160 VDYTQSFTTDKVFASRDEILEWARNLGKQHGFIIVITRSD--NGGLKRKTFMILGCERCD 217
VD + +F T +VF +RD +L+ R++ ++GF VI RSD G R +F+++GCER
Sbjct: 457 VDCSYAFNTXQVFGTRDYVLQ*XRSVAHENGFAAVIMRSDTHTGSRGRSSFVLIGCERNG 278
Query: 218 KYIPYKEVLKHQSTGTKKCYCPFRLRARGT--KVGWKVSVMNGYHIHERSETLLGHPYVG 275
+Y + ++ G++K CPF+LR + GW + ++ G H HE +++L+ HPY G
Sbjct: 277 QYKCRNKXXVIRAIGSRKYGCPFKLRGKPVIGGQGWMMKLICGIHNHELAKSLVEHPYAG 98
Query: 276 R 276
R
Sbjct: 97 R 95
>TC211617
Length = 534
Score = 61.2 bits (147), Expect = 6e-10
Identities = 29/67 (43%), Positives = 40/67 (59%), Gaps = 2/67 (2%)
Frame = -1
Query: 212 GCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRARGTK--VGWKVSVMNGYHIHERSETLL 269
GCER KY K+ + T T+KC CPF++R + GW V ++ G H HE ++TL+
Sbjct: 534 GCERSGKYKCRKKEFVRRDTCTRKCGCPFKIRGKPMHGGEGWTVKLICGIHNHELAKTLV 355
Query: 270 GHPYVGR 276
HPY GR
Sbjct: 354 EHPYAGR 334
>AI736706
Length = 426
Score = 47.4 bits (111), Expect = 9e-06
Identities = 23/63 (36%), Positives = 39/63 (61%), Gaps = 11/63 (17%)
Frame = +3
Query: 159 AVDYTQSFTTDKVFASRDEILEWARNLGKQHGFIIVITRSDNGGL-----------KRKT 207
+VD + +F T +VF SR+ +L WAR++ K++GF+++I RS+ +RKT
Sbjct: 237 SVDCSDAFNTTEVFPSREAMLNWARDVAKENGFVLIILRSETSTRSTKYTRSTRCNQRKT 416
Query: 208 FMI 210
F+I
Sbjct: 417 FVI 425
>TC234876 similar to UP|Q7RNR5 (Q7RNR5) 60S ribosomal protein L27 homolog,
partial (15%)
Length = 920
Score = 34.7 bits (78), Expect(2) = 4e-04
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 2/55 (3%)
Frame = -3
Query: 234 KKCYCPFRLRARGTK--VGWKVSVMNGYHIHERSETLLGHPYVGRPFTNKRLADG 286
+KC C F+L+ + GW ++ H H ++ L+ HPYV R N+++ G
Sbjct: 600 RKCGCLFKLQVKPVLGGEGWMKKLIRESHNHALAKLLVRHPYVDRLTINEKIIIG 436
Score = 26.2 bits (56), Expect(2) = 4e-04
Identities = 12/31 (38%), Positives = 20/31 (63%)
Frame = -2
Query: 196 TRSDNGGLKRKTFMILGCERCDKYIPYKEVL 226
T ++N G R +++++GCE KY YK+ L
Sbjct: 703 THTENRG--RISYVLIGCESSRKYTTYKKDL 617
>CF920253
Length = 856
Score = 37.4 bits (85), Expect = 0.009
Identities = 19/51 (37%), Positives = 32/51 (62%), Gaps = 2/51 (3%)
Frame = -2
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSD-NGGLKRKT-FMILGCERCDKY 219
VF RD++L W + + + F+ +I D N G++ KT F+++GCER +Y
Sbjct: 282 VFVIRDDVL*WTKFVAYEIEFVTLIMSLDTNTGMREKTSFVLIGCERNGQY 130
>BI497763
Length = 421
Score = 36.6 bits (83), Expect = 0.015
Identities = 14/27 (51%), Positives = 20/27 (73%)
Frame = +2
Query: 250 GWKVSVMNGYHIHERSETLLGHPYVGR 276
GW V ++ G H HE +++L+ HPYVGR
Sbjct: 44 GWMVKLLCGSHNHELTKSLVEHPYVGR 124
>TC231388 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At3g15920),
partial (10%)
Length = 684
Score = 30.4 bits (67), Expect = 1.1
Identities = 18/64 (28%), Positives = 32/64 (49%)
Frame = +1
Query: 150 AEDVQDEPLAVDYTQSFTTDKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFM 209
A+DV+D +AV+ T+ TTD + DE+ + H F+ DN L+++
Sbjct: 235 AQDVEDTVVAVEETRDITTDNTGKTHDELRKML-----AHMFV------DNASLRKQINS 381
Query: 210 ILGC 213
++ C
Sbjct: 382 VIRC 393
>TC234881
Length = 690
Score = 28.9 bits (63), Expect = 3.1
Identities = 14/42 (33%), Positives = 19/42 (44%)
Frame = +3
Query: 239 PFRLRARGTKVGWKVSVMNGYHIHERSETLLGHPYVGRPFTN 280
P+ AR WK++ N +H H + G P RP TN
Sbjct: 279 PYDSHARRNTKPWKLAFDNHHHHHRSKDKDPGKPSKTRPLTN 404
>TC204923 similar to GB|AAO39903.1|28372842|BT003675 At5g67590 {Arabidopsis
thaliana;} , partial (75%)
Length = 812
Score = 28.9 bits (63), Expect = 3.1
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Frame = -3
Query: 235 KCYCPF-RLRARGTKVGWKVSVMNGYHIHERSETLLGHPYVGRPFTN 280
KC+C F ++++G +V V +G H+ LLG V PF+N
Sbjct: 414 KCFCCFFTIKSQGRITNMRVGVPSGCPAHQWIFPLLGR*EVDLPFSN 274
>TC219082 similar to UP|Q9LYS5 (Q9LYS5) Receptor-like protein kinase, partial
(14%)
Length = 664
Score = 28.5 bits (62), Expect = 4.1
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = +3
Query: 89 IPKLKRGF--LELSGCRKRKLPSEERLETSREMGVRVWNLHL 128
IPK KRG ++L GC K ER+E ++VW L+L
Sbjct: 117 IPKQKRGPK*VKL*GCLKLTNIHSERIEEIERAALQVWKLNL 242
>CA853791
Length = 443
Score = 28.1 bits (61), Expect = 5.4
Identities = 16/50 (32%), Positives = 29/50 (58%)
Frame = -3
Query: 7 TTEFHHNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKL 56
+T H+Q +SI T+ T+SSI E +S + + RR++ + +L + L
Sbjct: 321 STHGMHSQFTITNSISTEKTQSSILVEGVSAIKKQRNRRQNPNIILRVDL 172
>TC215078 similar to UP|O80439 (O80439) 30S ribosomal protein S31
(At2g38140/F16M14.7) (Plastid-specific ribosomal protein
4 precursor), partial (48%)
Length = 1030
Score = 28.1 bits (61), Expect = 5.4
Identities = 36/122 (29%), Positives = 49/122 (39%), Gaps = 3/122 (2%)
Frame = -2
Query: 4 RAKTTEFHHNQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMR 63
R K T H NQ E + S ++ T S S LS + S PL G
Sbjct: 945 RHKITIHHQNQPEKRDSSISIFTTS--LSSSLSFLEGAGGA*GLSGPLPFFLSFGLAFPN 772
Query: 64 E-NENFAILIFPAPSSVNSKIGFGDRIPKLKRGFLELSGCR--KRKLPSEERLETSREMG 120
E + F +FP PS+ + G+G E GCR +R+ S ER + +G
Sbjct: 771 E*AKRFPFAVFPVPSATVHRRGWG-----------ESGGCRG*ERRKGSGER--*GKSLG 631
Query: 121 VR 122
VR
Sbjct: 630 VR 625
>CF923172
Length = 371
Score = 27.7 bits (60), Expect = 7.0
Identities = 16/61 (26%), Positives = 29/61 (47%)
Frame = -2
Query: 197 RSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRARGTKVGWKVSVM 256
R +NG L+ + ++ C + K LK +S +C+ + LR + K WK+ +
Sbjct: 301 RLNNGKLRLQHSLL----NCRXLLSMKHYLKERSVN*MRCFVFYILRKQNCKEWWKI*RL 134
Query: 257 N 257
N
Sbjct: 133 N 131
>TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (5%)
Length = 761
Score = 27.7 bits (60), Expect = 7.0
Identities = 19/58 (32%), Positives = 27/58 (45%)
Frame = +1
Query: 233 TKKCYCPFRLRARGTKVGWKVSVMNGYHIHERSETLLGHPYVGRPFTNKRLADGPRPV 290
TK+ Y PF ++ R KV +++ + IH L +VG P T GP PV
Sbjct: 79 TKRYYGPFEVQERLGKVVYRLKLTAHSRIHPVFHVSLLKAFVGDPETTHA---GPLPV 243
>TC231947 similar to UP|Q84NI9 (Q84NI9) Myb51 protein, partial (13%)
Length = 756
Score = 27.3 bits (59), Expect = 9.2
Identities = 16/62 (25%), Positives = 30/62 (47%), Gaps = 15/62 (24%)
Frame = +3
Query: 124 WNLHLKLRIEKVFIN--QEKPLTGKLVRA-------------EDVQDEPLAVDYTQSFTT 168
WN H+K +++K+ I+ KPL+ K + ++ Q+EP+ V+ F
Sbjct: 6 WNTHIKKKLKKMGIDPVTHKPLSNKTEQTQAQPDEQQTHQPLQEQQEEPIPVEKDTKFEP 185
Query: 169 DK 170
+K
Sbjct: 186 EK 191
>BM892562
Length = 422
Score = 27.3 bits (59), Expect = 9.2
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +3
Query: 13 NQSETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSS 49
+Q E +HS+L K+ RS S SE+ + SSS
Sbjct: 204 DQGENKHSLLEKLHRSDSSSSSSSEEEGENRG*SSSS 314
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,414,209
Number of Sequences: 63676
Number of extensions: 198236
Number of successful extensions: 1002
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 993
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 997
length of query: 313
length of database: 12,639,632
effective HSP length: 97
effective length of query: 216
effective length of database: 6,463,060
effective search space: 1396020960
effective search space used: 1396020960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0022.15


