

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.11
(334 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
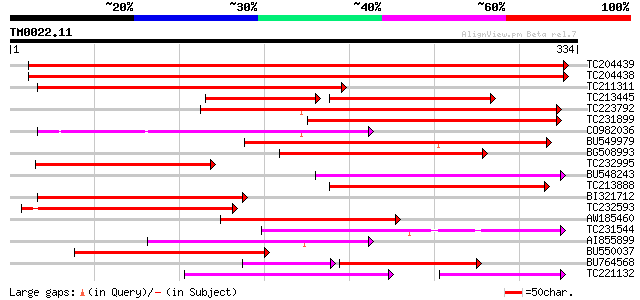
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 327 4e-90
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 325 1e-89
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 198 3e-51
TC213445 118 7e-46
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 164 7e-41
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 134 8e-32
CO982036 130 8e-31
BU549979 124 8e-29
BG508993 118 3e-27
TC232995 117 7e-27
BU548243 114 6e-26
TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%) 112 3e-25
BI321712 107 6e-24
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 105 3e-23
AW185460 102 3e-22
TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, ... 100 7e-22
AI855899 similar to GP|2244960|emb| retrotransposon like protein... 99 4e-21
BU550037 93 2e-19
BU764568 77 2e-19
TC221132 weakly similar to UP|O23529 (O23529) RETROTRANSPOSON li... 68 8e-19
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 327 bits (839), Expect = 4e-90
Identities = 157/318 (49%), Positives = 227/318 (71%)
Frame = +1
Query: 12 LVSEQIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQS 71
L+ QIYVDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q ++ ++ QS
Sbjct: 3763 LMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQS 3942
Query: 72 KYTKEFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDIL 131
+Y K +KKF M + +TP L K++ V Q LYR MIGSLLYLTASRPDI
Sbjct: 3943 RYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDIT 4122
Query: 132 FSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTE 191
++V +CA++Q++P+ +HLT VKRILKY+ GT++ G+MY S L GYCDAD+AG +
Sbjct: 4123 YAVGVCARYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADD 4302
Query: 192 RKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILES 251
RKST G C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + +
Sbjct: 4303 RKSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQD 4482
Query: 252 NISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSK 311
+++YCDN +AI++SKN + HSR KHI++++H+IRD V V+ LK VDT+ Q ADIF+K
Sbjct: 4483 VMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTK 4662
Query: 312 PLAEDRFKFILKNLNMDL 329
L ++F+ + L + L
Sbjct: 4663 ALDANQFEKLRGKLGICL 4716
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 325 bits (834), Expect = 1e-89
Identities = 156/318 (49%), Positives = 226/318 (71%)
Frame = +1
Query: 12 LVSEQIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQS 71
L+ QIYVDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q ++ ++ QS
Sbjct: 3766 LMIAQIYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQS 3945
Query: 72 KYTKEFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDIL 131
KY K +KKF M + +TP L K++ V Q LYR MIGSLLYLTASRPDI
Sbjct: 3946 KYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDIT 4125
Query: 132 FSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTE 191
++V +CA++Q++P+ +HL VKRILKY+ GT++ G+MY S L GYCDAD+AG +
Sbjct: 4126 YAVGVCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSADD 4305
Query: 192 RKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILES 251
RKST G C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + +
Sbjct: 4306 RKSTSGGCFYLGTNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQD 4485
Query: 252 NISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSK 311
+++YCDN +AI++SKN + HSR KHI++++H+IRD V V+ L+ VDT+ Q ADIF+K
Sbjct: 4486 VMTLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLEHVDTEEQIADIFTK 4665
Query: 312 PLAEDRFKFILKNLNMDL 329
L ++F+ + L + L
Sbjct: 4666 ALDANQFEKLRGKLGICL 4719
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 198 bits (503), Expect = 3e-51
Identities = 100/182 (54%), Positives = 128/182 (69%)
Frame = +3
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKE 76
IYVDDIIFG+ + +CKEF ++M+ FE SM GELK+ LG+Q+ Q+ +IHQ KYTK
Sbjct: 501 IYVDDIIFGATSKRMCKEFFELMKDGFETSMKGELKFLLGLQIIQKVYGIFIHQEKYTKS 680
Query: 77 FLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVHL 136
LK+F M E P TPMH + I++K++ K Y GMI SL YLT+SRPDI+F V L
Sbjct: 681 HLKRFRMDEAKPMATPMHRSTIIDKDEKGNHTS*KEYSGMIDSLSYLTSSRPDIVFVVCL 860
Query: 137 CAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTY 196
CA+FQS P+ +H+T VKRIL+YL GTTN L +KK S + L GYCD +AGD+ ERKST
Sbjct: 861 CARFQSYPKISHVTAVKRILRYLVGTTNHCLWFKKRSEFDLLGYCDVYFAGDKVERKSTS 1040
Query: 197 GN 198
N
Sbjct: 1041RN 1046
>TC213445
Length = 705
Score = 118 bits (296), Expect(2) = 7e-46
Identities = 56/98 (57%), Positives = 75/98 (76%)
Frame = +1
Query: 189 RTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI 248
+T+R+ST C F+GS LVSW SK+Q+++ LSTAEAEYISA Q+ WM+ QL DY +
Sbjct: 400 KTDRESTSDTCHFIGSALVSWHSKKQNSVVLSTAEAEYISARSYYAQIFWMRQQLFDYGL 579
Query: 249 LESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIR 286
+I I CDNT+AI+LSKN IL+SR KHIE+++HF+R
Sbjct: 580 KLDHIPIRCDNTSAINLSKNHILYSRTKHIEIRHHFLR 693
Score = 83.6 bits (205), Expect(2) = 7e-46
Identities = 38/68 (55%), Positives = 52/68 (75%)
Frame = +2
Query: 116 MIGSLLYLTASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*Y 175
MI S LYL+ SRP I+FSV +C ++Q++P+E+HL+ +KRI++YL G NLGL Y K+S Y
Sbjct: 197 MIESFLYLSTSRPHIMFSVCMCVRYQANPKESHLSVIKRIMRYLLGIINLGLWYPKNSSY 376
Query: 176 KLSGYCDA 183
L GY DA
Sbjct: 377 NLVGYSDA 400
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 164 bits (414), Expect = 7e-41
Identities = 87/217 (40%), Positives = 137/217 (63%), Gaps = 4/217 (1%)
Frame = +1
Query: 113 YRGMIGSLLYLTASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYK-- 170
+R +IGSL YL SRP+I F+V L ++F PR +H+ KR+L+ +KGT G+++
Sbjct: 22 FRRLIGSLRYLCNSRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVLFPFK 201
Query: 171 -KSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISA 229
KS L GY D+D+ D + KST G V+ +SK+Q IALST EAEY++A
Sbjct: 202 AKSGKPDLLGYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAEYVAA 381
Query: 230 AICSTQMLWMKHQLEDYQILESN-ISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDY 288
++ + Q +WM + LE+ ++ E +++ DN +AI+L+K+ LH R+KHIE+++H+IRD
Sbjct: 382 SLGACQAVWMMNLLEELKLRERKPVNLLIDNKSAINLAKHPTLHGRSKHIELRFHYIRDQ 561
Query: 289 VQKGVLLLKFVDTDHQWADIFSKPLAEDRFKFILKNL 325
V KG + +++ + Q AD+ +KP+ RFK I L
Sbjct: 562 VSKGNVTVEYCKAEEQLADLMTKPIQVSRFKQICSEL 672
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment),
partial (30%)
Length = 687
Score = 134 bits (336), Expect = 8e-32
Identities = 62/151 (41%), Positives = 99/151 (65%), Gaps = 1/151 (0%)
Frame = +2
Query: 176 KLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQ 235
+LSGYCDAD+AG +R+ST G C F+G NLVSW SK+Q+ +A S+AEAEY S A+ + +
Sbjct: 14 QLSGYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCE 193
Query: 236 MLWMKHQLEDYQILES-NISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVL 294
++W+K L++ + E + +YCDN AA+ ++ N + H R KHIE+ HFIR+ + +
Sbjct: 194 LMWIKQFLQELRFCEELQMKLYCDNQAALHIASNPVFHERTKHIEIDCHFIREKLLSKEI 373
Query: 295 LLKFVDTDHQWADIFSKPLAEDRFKFILKNL 325
+ +F+ ++ Q DI +K L + + + L
Sbjct: 374 VTEFIGSNDQPVDILTKSLRGPKIQIVCSKL 466
>CO982036
Length = 674
Score = 130 bits (327), Expect = 8e-31
Identities = 74/201 (36%), Positives = 114/201 (55%), Gaps = 3/201 (1%)
Frame = -2
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKE 76
+YVD II GS+ T L + + + + F + ++G+L YF+ I+V P+ + ++ +
Sbjct: 640 VYVDIIITGSSCT-LIQNLTSKLNSSFPLKLLGKLDYFVEIEVKSMPDLLFSLRTSIFEI 464
Query: 77 FLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVHL 136
F +K + P +PM TC L K D YR ++G+L Y T RP+I F+V+
Sbjct: 463 FCRK-PR*QAQPISSPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTVIRPEISFAVNK 287
Query: 137 CAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYK---KSS*YKLSGYCDADYAGDRTERK 193
QF S+P ++H T VKRIL+YLKG+ + GL K S + G+CDAD+A +++
Sbjct: 286 VCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAISSQPLPIRGFCDADWASAVDDKR 107
Query: 194 STYGNCQFLGSNLVSWASKRQ 214
ST G FLG NL+SW +Q
Sbjct: 106 STSGAAVFLGPNLISWWXXKQ 44
>BU549979
Length = 615
Score = 124 bits (310), Expect = 8e-29
Identities = 62/184 (33%), Positives = 113/184 (60%), Gaps = 3/184 (1%)
Frame = -1
Query: 139 QFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGN 198
++QS+P H T K++++YL+GT + LMYK+++ ++ GY D+D+AG R+ST G
Sbjct: 606 RYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYKQTNCLEVIGYSDSDFAGCVDSRRSTSGY 427
Query: 199 CQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILES---NISI 255
L +VSW S +Q+ IA ST E E++ ++ +W+K + ++++S + +
Sbjct: 426 IFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSLRVVDSISRPLKL 247
Query: 256 YCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPLAE 315
YCDN AA+ ++KN +R+KHI++KY IR+ V++ ++++ V+T+ D +K +
Sbjct: 246 YCDNFAAVFMAKNNKSGNRSKHIDIKYLVIRERVKEKKVVIEHVNTELMIVDPLTKGMTP 67
Query: 316 DRFK 319
FK
Sbjct: 66 KNFK 55
>BG508993
Length = 374
Score = 118 bits (296), Expect = 3e-27
Identities = 54/123 (43%), Positives = 82/123 (65%), Gaps = 1/123 (0%)
Frame = +1
Query: 160 KGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIAL 219
KGT + GL Y S+ YKL G+CD+D+AGD +RKST G F+G + +W+SK+Q + L
Sbjct: 4 KGTIDFGLFYSPSNNYKLVGFCDSDFAGDVDDRKSTTGFVFFMGDCVFTWSSKKQGIVTL 183
Query: 220 STAEAEYISAAICSTQMLWMKHQLEDYQILE-SNISIYCDNTAAISLSKNLILHSRAKHI 278
T EAEY++A C+ +W++ LE+ Q+L+ + IY DN +A L+KN + H R+KHI
Sbjct: 184 FTCEAEYVAATSCTCHAIWLRRLLEELQLLQKESTKIYVDNRSAQELAKNSVFHERSKHI 363
Query: 279 EVK 281
+ +
Sbjct: 364 DTR 372
>TC232995
Length = 1009
Score = 117 bits (293), Expect = 7e-27
Identities = 58/106 (54%), Positives = 71/106 (66%)
Frame = +2
Query: 16 QIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTK 75
QIYVDDIIFGS N LCKEFS MQ+EFEMSMMGELKYFLG+Q+ Q +I+QSKY K
Sbjct: 203 QIYVDDIIFGSTNDSLCKEFSLDMQSEFEMSMMGELKYFLGLQIKQTQ*GIFINQSKYCK 382
Query: 76 EFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLL 121
E +K+F M TPM C L+K++ + K YR IG ++
Sbjct: 383 ELIKRFGMDSAKHMSTPMSTNCYLDKDESGQSIDIKQYRDAIGEVV 520
>BU548243
Length = 599
Score = 114 bits (285), Expect = 6e-26
Identities = 57/147 (38%), Positives = 87/147 (58%)
Frame = -1
Query: 181 CDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMK 240
CDA +A D + +ST G+ FLG NL+SW S++Q A S+ EAEY S A S ++ W++
Sbjct: 590 CDAGWASDVDDHRSTLGSAIFLGPNLISWWSRKQQVTAQSSTEAEYRSIAQTSAELTWIQ 411
Query: 241 HQLEDYQILESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVD 300
L + QI + I CDN +A++++ NL+ HSR KH+E+ F+ + V L + +
Sbjct: 410 ALLMELQIPFTPPVILCDNKSAVAIAHNLVFHSRTKHMEIDVFFVHEKVLSKQLQIFHIP 231
Query: 301 TDHQWADIFSKPLAEDRFKFILKNLNM 327
QWA I +KPL+ RF F+ L +
Sbjct: 230 ALDQWAGILTKPLSSARFTFLKSKLTV 150
>TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%)
Length = 493
Score = 112 bits (279), Expect = 3e-25
Identities = 53/131 (40%), Positives = 86/131 (65%), Gaps = 1/131 (0%)
Frame = +3
Query: 189 RTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI 248
R +RKST G F+G +W SK+Q + LST EAEY++A C +W+++ L++ ++
Sbjct: 6 RDDRKSTTGFVFFMGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELKM 185
Query: 249 -LESNISIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWAD 307
E + I DN +A++L+KN + H ++KHI+ +YHFIR+ ++K + LK+V + Q AD
Sbjct: 186 PQEEPMEICVDNKSALALAKNPVFHEKSKHIDTRYHFIRECIEKKEVKLKYVMSQDQAAD 365
Query: 308 IFSKPLAEDRF 318
IF+KPL + F
Sbjct: 366 IFTKPLKLETF 398
>BI321712
Length = 399
Score = 107 bits (268), Expect = 6e-24
Identities = 55/124 (44%), Positives = 76/124 (60%)
Frame = -3
Query: 17 IYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKE 76
+YVDD+IF N + +EF K M EFEM+ MG + Y+LGI+V Q+ + +I Q Y KE
Sbjct: 379 LYVDDLIFTGNNPSMFEEFKKDMSNEFEMTDMGLMAYYLGIEVKQEDKGIFITQEGYAKE 200
Query: 77 FLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVHL 136
LKKF M + P TPM L K + V LY+ +IGSL YLT +RPDIL+ V +
Sbjct: 199 VLKKFKMDDANPVGTPMECGSKLSKHEKGENVDPTLYKSLIGSLRYLTCTRPDILYVVGV 20
Query: 137 CAQF 140
+++
Sbjct: 19 VSRY 8
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 105 bits (262), Expect = 3e-23
Identities = 55/127 (43%), Positives = 77/127 (60%)
Frame = +1
Query: 8 SYLQLVSEQIYVDDIIFGSANTFLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQQPEATY 67
+Y +VS +YVDD++ + L +EF + M FEM+ +G + YFLGI++ Q
Sbjct: 184 NYFLIVS--LYVDDLLVTRDDARLVEEFKQEMMQAFEMTNLGLMTYFLGIEIKQSQNKVL 357
Query: 68 IHQSKYTKEFLKKFNMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASR 127
I Q KY KE LKKF M EC TPM+ K D + K+ + YR +IG L+YLTA+R
Sbjct: 358 ICQRKYAKEILKKFQMEECKSVSTPMNQKEKFNKVDGADKIDEGYYRSLIGCLMYLTATR 537
Query: 128 PDILFSV 134
PDILF++
Sbjct: 538 PDILFAI 558
>AW185460
Length = 411
Score = 102 bits (253), Expect = 3e-22
Identities = 49/106 (46%), Positives = 70/106 (65%)
Frame = +2
Query: 125 ASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDAD 184
A+RPDI+++ L ++F P + H KRIL+YL+GT G+ Y + +L GY D+D
Sbjct: 89 ATRPDIMYATSLLSRFMQSPSQIHFGAGKRILRYLQGTKAFGIWYTTETNSELLGYTDSD 268
Query: 185 YAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAA 230
+AG + KST G LGS + SWASK+Q+T+A STAEAEY++ A
Sbjct: 269 WAGSTDDMKSTSGYAFSLGSGMFSWASKKQATVAQSTAEAEYVAVA 406
>TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, partial
(16%)
Length = 662
Score = 100 bits (250), Expect = 7e-22
Identities = 63/183 (34%), Positives = 103/183 (55%), Gaps = 4/183 (2%)
Frame = +3
Query: 149 LTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVS 208
L R+LKYLKG GL + + S ++ G+ DAD+A KS C FLGS+L+S
Sbjct: 12 LCAATRVLKYLKGCPRKGLSFSRESPIQILGFSDADWATCIDSSKSITWYCFFLGSSLIS 191
Query: 209 WASKRQSTIALSTAEAEYISAAICST--QMLWMKHQLEDYQILESNISIYCDNTAAIS-L 265
W +K+Q+T++ S++ +E A+ ST ++ W+ + L+D + IYCDN +A+ L
Sbjct: 192 WKAKKQNTVSRSSSSSEAKYRALTSTTCELQWLTYLLKDLHV----TLIYCDNQSALQ*L 359
Query: 266 SKNLILHSRAKHIEVKYHFIRDYVQKGVL-LLKFVDTDHQWADIFSKPLAEDRFKFILKN 324
+I H + +E+ H +R+ Q+G++ L V + +Q ADIF+K L+ F L
Sbjct: 360 PIKVIYHGQ---LEIDCHIVREKTQQGLMHCLLPVSSSNQLADIFTKALSPKLFSSNLSK 530
Query: 325 LNM 327
L +
Sbjct: 531 LGL 539
>AI855899 similar to GP|2244960|emb| retrotransposon like protein
{Arabidopsis thaliana}, partial (18%)
Length = 418
Score = 98.6 bits (244), Expect = 4e-21
Identities = 52/136 (38%), Positives = 80/136 (58%), Gaps = 3/136 (2%)
Frame = +1
Query: 82 NMTECTPAKTPMHPTCILEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVHLCAQFQ 141
NM +C TPM + L K YR ++G+L Y+T +RP+I ++V+ ++F
Sbjct: 10 NMLDCNGISTPMVSSYKLSKFGSELLPNAHQYRDIVGALQYVTLTRPNIAYNVNKVSEFM 189
Query: 142 SDPRETHLTTVKRILKYLKGTTNLGLMYKKS---S*YKLSGYCDADYAGDRTERKSTYGN 198
S P +++ TVKRIL+YL GT GL+ + + + L Y D D+ D E +ST G+
Sbjct: 190 SSPLQSY*LTVKRILRYLSGTVTQGLLLQPAHMDAKISLRAYNDLDWGSDPAEMRSTSGS 369
Query: 199 CQFLGSNLVSWASKRQ 214
C F GSNL++W+SK+Q
Sbjct: 370 CIFSGSNLIAWSSKKQ 417
>BU550037
Length = 728
Score = 92.8 bits (229), Expect = 2e-19
Identities = 44/115 (38%), Positives = 72/115 (62%)
Frame = -3
Query: 39 MQAEFEMSMMGELKYFLGIQVDQQPEATYIHQSKYTKEFLKKFNMTECTPAKTPMHPTCI 98
M++EFEM+ +G++KYFLG+ + Q + +I Q KY E L+KF+M C P T +
Sbjct: 363 MESEFEMTDLGQMKYFLGM*IFQSEDGIFISQKKYAWEILRKFHMERCKPIATVLVVNEK 184
Query: 99 LEKEDVSAKVCQKLYRGMIGSLLYLTASRPDILFSVHLCAQFQSDPRETHLTTVK 153
K++ + +YR +IGSLLYL+A+RP+++F+ L ++F P + HL + K
Sbjct: 183 FSKDEEDNQGDASVYRSLIGSLLYLSATRPNLMFAATLLSRFTKSPSQVHLGSSK 19
>BU764568
Length = 420
Score = 77.4 bits (189), Expect(2) = 2e-19
Identities = 36/85 (42%), Positives = 57/85 (66%), Gaps = 1/85 (1%)
Frame = +3
Query: 195 TYGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILE-SNI 253
T G C +G NL+SW SK+QS +A S+AEAEY + A+ + +++W+K L + + E + +
Sbjct: 165 TSGYCVLIGGNLISWKSKKQSVVAKSSAEAEYRAMALVTCELIWLKQLL*ELKFEEDTQM 344
Query: 254 SIYCDNTAAISLSKNLILHSRAKHI 278
++ CDN AA+ ++ N I H R KHI
Sbjct: 345 TLICDNQAALHIASNPIFH*RTKHI 419
Score = 35.8 bits (81), Expect(2) = 2e-19
Identities = 19/55 (34%), Positives = 28/55 (50%)
Frame = +1
Query: 138 AQFQSDPRETHLTTVKRILKYLKGTTNLGLMYKKSS*YKLSGYCDADYAGDRTER 192
+QF + P + H V ILK K GL+Y+ ++ GY DAD G ++R
Sbjct: 1 SQFLNSPCQDHWNAVS*ILK*TKSAPGKGLIYEDKGHSQIIGYSDAD*VGSPSDR 165
>TC221132 weakly similar to UP|O23529 (O23529) RETROTRANSPOSON like protein,
partial (5%)
Length = 799
Score = 68.2 bits (165), Expect(2) = 8e-19
Identities = 35/123 (28%), Positives = 65/123 (52%)
Frame = +1
Query: 104 VSAKVCQKLYRGMIGSLLYLTASRPDILFSVHLCAQFQSDPRETHLTTVKRILKYLKGTT 163
V + + L+ + + ++ S I S+ +QF DP + + KR+L+YLKGT
Sbjct: 31 VMSPLVMALFTASLSTRCNISLSHALI*HSLSTSSQFMKDPTKIRMQATKRVLRYLKGTI 210
Query: 164 NLGLMYKKSS*YKLSGYCDADYAGDRTERKSTYGNCQFLGSNLVSWASKRQSTIALSTAE 223
+ GL + S L + DA++ + ++ +ST + G +++SW+ K+QS I S+ +
Sbjct: 211 DFGLQLRSSPDQHLRAFYDANWVDNTSDIRSTGAYVVYFGLSVISWSCKKQSIIDKSSTK 390
Query: 224 AEY 226
EY
Sbjct: 391 VEY 399
Score = 43.1 bits (100), Expect(2) = 8e-19
Identities = 21/74 (28%), Positives = 38/74 (50%)
Frame = +2
Query: 254 SIYCDNTAAISLSKNLILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFSKPL 313
++Y N A+ L N + H KH+ + + F++D V L + V + H D+F+K L
Sbjct: 452 TMYSYNIGAMYLCANPVFHLCMKHLTIDHLFVQDLVANKQLRVSHVPSCH*HVDLFTKAL 631
Query: 314 AEDRFKFILKNLNM 327
R KF++ + +
Sbjct: 632 VSSRHKFMMDKIGV 673
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,066,006
Number of Sequences: 63676
Number of extensions: 203603
Number of successful extensions: 1317
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 1292
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1295
length of query: 334
length of database: 12,639,632
effective HSP length: 98
effective length of query: 236
effective length of database: 6,399,384
effective search space: 1510254624
effective search space used: 1510254624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0022.11


