

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0020.6
(795 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
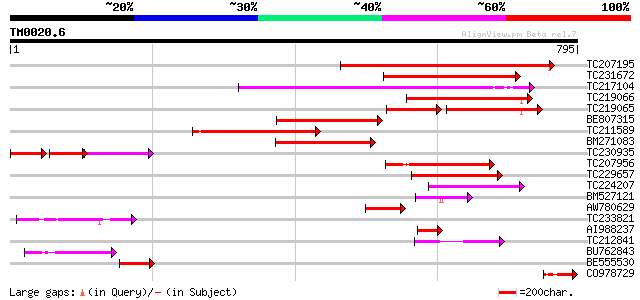
Score E
Sequences producing significant alignments: (bits) Value
TC207195 374 e-104
TC231672 similar to UP|Q8S026 (Q8S026) OJ1029_F04.25 protein, pa... 284 1e-76
TC217104 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), ... 269 4e-72
TC219066 233 2e-61
TC219065 weakly similar to UP|Q8S026 (Q8S026) OJ1029_F04.25 prot... 134 3e-59
BE807315 219 3e-57
TC211589 219 5e-57
BM271083 178 9e-45
TC230935 similar to UP|Q9MAV5 (Q9MAV5) F24O1.4, partial (20%) 84 2e-41
TC207956 weakly similar to UP|Q94A87 (Q94A87) At1g10080/T27I1_10... 122 6e-28
TC229657 similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21 pr... 114 2e-25
TC224207 similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21 pr... 104 1e-22
BM527121 similar to GP|20146379|dbj OJ1029_F04.25 {Oryza sativa ... 104 2e-22
AW780629 93 4e-19
TC233821 weakly similar to UP|Q75GI5 (Q75GI5) Expressed protein,... 82 7e-16
AI988237 71 2e-12
TC212841 similar to UP|Q75LY1 (Q75LY1) Expressed protein (With a... 70 3e-12
BU762843 65 9e-11
BE555530 64 3e-10
CO978729 62 1e-09
>TC207195
Length = 1194
Score = 374 bits (960), Expect = e-104
Identities = 179/304 (58%), Positives = 235/304 (76%), Gaps = 4/304 (1%)
Frame = +3
Query: 464 AKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGW 523
++YY F VN+FLG+I+TGTAFEQL SF+HQ + P TIG +IP+KA+FF+TYIMVDGW
Sbjct: 3 SRYYLFNFVNIFLGNILTGTAFEQLDSFIHQPANEYPITIGTAIPLKASFFITYIMVDGW 182
Query: 524 AGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAV 583
AGIA E+L LKPL+IYHLKN F+VKTE+DR +AMDPGS+ F P +QLYFLLG+VYA
Sbjct: 183 AGIAAEVLMLKPLIIYHLKNFFLVKTEKDREEAMDPGSIGFNTGEPRIQLYFLLGLVYAS 362
Query: 584 VTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGL 643
VTP +LPFI+VFF LAY+V+RHQIINVYNQ+YES AAFWP VH R+I +L++SQ++++GL
Sbjct: 363 VTPTVLPFIIVFFGLAYVVFRHQIINVYNQEYESGAAFWPDVHFRVIIALIVSQIVLMGL 542
Query: 644 LSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIK 703
L+TKKAA STP L +LP+LT FH YC+ RFEPAF +YPL+EAM KD LE+ T+P+ N+K
Sbjct: 543 LTTKKAASSTPFLIVLPVLTIWFHIYCKGRFEPAFVRYPLQEAMMKDTLERATDPNFNLK 722
Query: 704 AYLADAYLHPIFRSFEVEEELVEVRVDKQPETQ-VASPTLSEPSSPSPLHDHV---HQPS 759
AYL +AY+HP+F++ +E+ E + + ET+ + PT + +PL + PS
Sbjct: 723 AYLQNAYVHPVFKASLFDEDEDEEVMSLKLETESLTVPTKRQSRRNTPLPSRISGASSPS 902
Query: 760 PPQH 763
P H
Sbjct: 903 LPDH 914
>TC231672 similar to UP|Q8S026 (Q8S026) OJ1029_F04.25 protein, partial (32%)
Length = 706
Score = 284 bits (726), Expect = 1e-76
Identities = 134/193 (69%), Positives = 163/193 (84%)
Frame = +1
Query: 524 AGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAV 583
AG AGEILRLKPL+ YHLKN F+VKTE+DR +AMDPG+ F P +QLYFLLG+VYAV
Sbjct: 28 AGCAGEILRLKPLIFYHLKNFFLVKTEKDREEAMDPGTFGFNTGEPQIQLYFLLGLVYAV 207
Query: 584 VTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGL 643
VTP LLP+I+VFF LAY+VYRHQIINVYNQ+YESAAAFWP VH RII +L+ISQLL++GL
Sbjct: 208 VTPFLLPYIIVFFGLAYVVYRHQIINVYNQEYESAAAFWPDVHGRIIFALVISQLLLMGL 387
Query: 644 LSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIK 703
LSTK+AA STPLL LPILT +FH YC+ R+EPAF K+PL+EAM KD LE+ EP+ N+K
Sbjct: 388 LSTKEAANSTPLLITLPILTISFHLYCKGRYEPAFVKHPLQEAMMKDTLERAREPNFNLK 567
Query: 704 AYLADAYLHPIFR 716
+L +AY+HP+F+
Sbjct: 568 EFLQNAYIHPVFK 606
>TC217104 similar to UP|Q94II2 (Q94II2) ERD4 protein (Fragment), partial
(62%)
Length = 1535
Score = 269 bits (687), Expect = 4e-72
Identities = 145/416 (34%), Positives = 237/416 (56%), Gaps = 1/416 (0%)
Frame = +3
Query: 321 AFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSV 380
A + FSSR A+ +Q+ ++ W APEP + W NL I + +R+ ++ V
Sbjct: 15 AVVFFSSRVVAASASQSLHAQMVDTWSVFDAPEPNQLIWPNLKIKYFQRELRQYLVYFIV 194
Query: 381 FALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILP 440
+FFYMIPI F+ + L+ L + PF++P++ +K +++ L+ +LP LAL IFL +LP
Sbjct: 195 ALTIFFYMIPITFISAFTTLDNLVKYLPFIKPIVNIKALRTVLEAYLPQLALIIFLALLP 374
Query: 441 AVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFL-HQSPTQI 499
+L+ +SK EG +S R + KY+YF ++NVF+G + GT F+ H + +I
Sbjct: 375 KLLLFLSKFEGIPTESHAVRAASGKYFYFTVLNVFIGVTIGGTLFKAFKRIREHPTLDEI 554
Query: 500 PRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDP 559
+ S+P ATFF+TY+ + + G E+ R+ PL+IYHLK ++ KTE + +A P
Sbjct: 555 SSLLAESLPGNATFFLTYVALKFFIGYGLELSRIVPLIIYHLKRKYLCKTEAELKEAWRP 734
Query: 560 GSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAA 619
G + + +P L + Y+V+ P+++PF ++F L +LV R+Q + VY +ES
Sbjct: 735 GDLGYGTRVPGDMLIVTIVFCYSVIAPVIIPFGALYFGLGWLVLRNQALKVYVPTFESYG 914
Query: 620 AFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFR 679
WPH+H+RI+ASL++ Q+ M G T+K TPL+ LPIL+ F C +F PAF
Sbjct: 915 RMWPHIHNRILASLILYQITMFGYFGTQK-FYYTPLVLPLPILSLVFGFVCAKKFYPAF- 1088
Query: 680 KYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFEVEEELVEVRVDKQPET 735
++P E A L E P++ + + AY+ P RS +++ + VE + + T
Sbjct: 1089QHPALEVAANTLKE---VPNMEL---IFGAYIPPSLRSEKIDGDRVEDALSQASRT 1238
>TC219066
Length = 586
Score = 233 bits (595), Expect = 2e-61
Identities = 113/189 (59%), Positives = 143/189 (74%), Gaps = 13/189 (6%)
Frame = +2
Query: 557 MDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYE 616
+DPGS+ F P +QLYFLLG+VYA VTP +LPFI VFF LAYLV+RHQIINVYNQ+YE
Sbjct: 2 LDPGSIGFNTGEPRIQLYFLLGLVYAAVTPAVLPFITVFFGLAYLVFRHQIINVYNQEYE 181
Query: 617 SAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEP 676
S AAFWP VH RI+ +L++SQ++++GLL+TKKAA STP L +LPILT FH+YC+ RFE
Sbjct: 182 SGAAFWPDVHFRIVMALIVSQIVLMGLLTTKKAASSTPFLIVLPILTIWFHRYCKGRFES 361
Query: 677 AFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFR-------------SFEVEEE 723
AF K PL+EAM KD LE+TTEP+LN+K YL +AY+HP+F+ S ++E E
Sbjct: 362 AFVKCPLQEAMMKDTLERTTEPNLNLKGYLQNAYVHPVFKDSMDDDDDEEDILSMDLETE 541
Query: 724 LVEVRVDKQ 732
V VR +Q
Sbjct: 542 SVTVRTKRQ 568
>TC219065 weakly similar to UP|Q8S026 (Q8S026) OJ1029_F04.25 protein, partial
(35%)
Length = 1081
Score = 134 bits (337), Expect(2) = 3e-59
Identities = 69/148 (46%), Positives = 94/148 (62%), Gaps = 13/148 (8%)
Frame = +1
Query: 613 QQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQH 672
Q+YES AAFWP VH + +L Q++++GLL+TKKAA STP L +LPILT FH+YC+
Sbjct: 253 QEYESGAAFWPDVH*KKKMALRGGQIVLMGLLTTKKAASSTPFLIVLPILTIWFHRYCKG 432
Query: 673 RFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFR-------------SFE 719
R E AF K+PL+EAM KD LE+ TEP+LN K YL +AY+HP+ + S +
Sbjct: 433 RLESAFVKFPLQEAMMKDTLERATEPNLNSKGYLQNAYVHPVSKDSMDDDDDEEDRLSID 612
Query: 720 VEEELVEVRVDKQPETQVASPTLSEPSS 747
+E E V VR +Q P+ + +S
Sbjct: 613 LETESVTVRTKRQSRRNTPLPSKNNDAS 696
Score = 114 bits (284), Expect(2) = 3e-59
Identities = 52/77 (67%), Positives = 63/77 (81%)
Frame = +2
Query: 529 EILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPIL 588
E+L LKPL++YHLKN F+VKTE+DR +AMDPGS+ F P +QLYFLLG+VYA VTP +
Sbjct: 2 EVLMLKPLIVYHLKNFFLVKTEKDREEAMDPGSIGFNTGEPRIQLYFLLGLVYAAVTPAV 181
Query: 589 LPFILVFFALAYLVYRH 605
LPFI FF LAYLV+RH
Sbjct: 182 LPFISFFFGLAYLVFRH 232
>BE807315
Length = 450
Score = 219 bits (559), Expect = 3e-57
Identities = 104/149 (69%), Positives = 128/149 (85%)
Frame = +3
Query: 374 LVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALK 433
L+I+++ F L FF+MIPIAFVQSLAN+EG+E+ APFL+ IE++FIKSF+QGFLPG+ALK
Sbjct: 3 LIIAVAFFFLTFFFMIPIAFVQSLANIEGIEKAAPFLKSFIEMQFIKSFIQGFLPGIALK 182
Query: 434 IFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLH 493
IFL LPA+LMIMSK EG I+ S LER+ A +YY F +NVFLGSI+TGTAF+QL F+H
Sbjct: 183 IFLIFLPAILMIMSKFEGFISTSALERRAATRYYIFQFINVFLGSIITGTAFQQLDKFIH 362
Query: 494 QSPTQIPRTIGVSIPMKATFFMTYIMVDG 522
QS +IP+TIGVSIPMKATFF+TYIMVDG
Sbjct: 363 QSANEIPKTIGVSIPMKATFFITYIMVDG 449
>TC211589
Length = 536
Score = 219 bits (557), Expect = 5e-57
Identities = 102/179 (56%), Positives = 139/179 (76%)
Frame = +3
Query: 257 LDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKC 316
L YYQ K +R +RP I TG LGL G KVDAI++++ I +L K + LER + DPK
Sbjct: 3 LVYYQNKVER-TSERPQIKTGFLGLCGNKVDAIDHHNTEIDKLSKEIALERDNVSNDPKS 179
Query: 317 ILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVI 376
I+P AF+SF +RWGA+VCAQTQQ+ NPT+WLT+WAPEPRDIYW NLAIP+VSL++R+L++
Sbjct: 180 IMPAAFVSFKTRWGAAVCAQTQQTTNPTMWLTEWAPEPRDIYWSNLAIPYVSLTVRRLIM 359
Query: 377 SLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIF 435
+++ F L FF+MIPIA VQ LA++EG+ + AP+L +I++ FIK +Q LPG+ALK+F
Sbjct: 360 AVAFFFLTFFFMIPIAIVQGLASIEGIRKRAPWLNALIDIPFIKPLIQRCLPGIALKLF 536
>BM271083
Length = 422
Score = 178 bits (451), Expect = 9e-45
Identities = 81/140 (57%), Positives = 117/140 (82%)
Frame = +1
Query: 373 KLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLAL 432
+L+I+++ F L FF+MIPIAFVQ+LA+L+G+++ AP+L+P++++ FIKSF+QGFLPG+ L
Sbjct: 1 RLIIAVTFFFLTFFFMIPIAFVQTLASLDGIQKAAPWLKPLVDIPFIKSFIQGFLPGIVL 180
Query: 433 KIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFL 492
K+FL LP +LMIMSK EG + S+LER++A++YY F VN+FLG+I+TGTAF+QL SF+
Sbjct: 181 KLFLIFLPTILMIMSKFEGFGSISSLERRSASRYYLFNFVNIFLGNILTGTAFQQLSSFI 360
Query: 493 HQSPTQIPRTIGVSIPMKAT 512
HQ Q P TIG +IP+KA+
Sbjct: 361 HQPADQYPVTIGTAIPLKAS 420
>TC230935 similar to UP|Q9MAV5 (Q9MAV5) F24O1.4, partial (20%)
Length = 990
Score = 83.6 bits (205), Expect(3) = 2e-41
Identities = 37/51 (72%), Positives = 44/51 (85%)
Frame = +3
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSP 51
MATL+DIGV+A +NILSAF F +AFA+LR+QP NDRVYFPKWY+ G RT P
Sbjct: 81 MATLSDIGVAAGLNILSAFIFFVAFAILRLQPFNDRVYFPKWYLKGLRTDP 233
Score = 72.8 bits (177), Expect(3) = 2e-41
Identities = 33/52 (63%), Positives = 43/52 (82%)
Frame = +1
Query: 57 FVGKFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKI 108
F KFVNL++++YL FLNWMP ALRM E E+I+HAGLDS V+LRIY +G ++
Sbjct: 250 FGRKFVNLDWRSYLRFLNWMPAALRMPELELIDHAGLDSVVYLRIYLVGYEL 405
Score = 53.1 bits (126), Expect(3) = 2e-41
Identities = 37/99 (37%), Positives = 54/99 (54%), Gaps = 3/99 (3%)
Frame = +3
Query: 106 LKIFVPVTVVALLILIPVNVSSGALFFLKRDLVVS-DIDKLSISNVPSKSSRFFVH--IG 162
LKIFVP+ +A +L+PVN +S L D + S DIDKLSISNV S S RF H +
Sbjct: 480 LKIFVPIAFLAWAVLVPVNATSTGLESAGLDNITSSDIDKLSISNVHSTSERFLNHHAVL 659
Query: 163 LEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFT 201
L++ +++ F + + H L+ RR +F+
Sbjct: 660 LQFCC*IFVLFSALVSNPIL*LSWQHVLSLWRRTSVRFS 776
>TC207956 weakly similar to UP|Q94A87 (Q94A87) At1g10080/T27I1_10, partial
(20%)
Length = 1050
Score = 122 bits (306), Expect = 6e-28
Identities = 66/155 (42%), Positives = 101/155 (64%), Gaps = 2/155 (1%)
Frame = +3
Query: 527 AGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFP--ETIPSLQLYFLLGIVYAVV 584
A E++++ PL + +L FI++ + D A+D GS+ FP +P + L+ LG A++
Sbjct: 15 AVEVMQIFPL-LRNLFQRFILRLKED---ALD-GSLSFPYHTEVPRILLFGFLGFTCAIL 179
Query: 585 TPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLL 644
P++LPF+LV+F +AYLVYR+QIINVY +Y+S FWP VH+ + SLL SQL+ LG+
Sbjct: 180 APLMLPFLLVYFFIAYLVYRNQIINVYITKYDSGGQFWPIVHNTTVFSLLFSQLIALGVF 359
Query: 645 STKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFR 679
K+++ ++ L I T FH+YC+ RF P FR
Sbjct: 360 GLKRSSVASGFTIPLLIGTLLFHQYCRQRFLPVFR 464
>TC229657 similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21 protein),
partial (24%)
Length = 895
Score = 114 bits (284), Expect = 2e-25
Identities = 49/127 (38%), Positives = 83/127 (64%)
Frame = +3
Query: 564 FPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWP 623
+ + IP + + LLGI Y + P++LPF+L +F LAY+++R+Q INVY +Y++A FWP
Sbjct: 9 YHKDIPRVLFFGLLGITYFFLAPLILPFLLAYFCLAYIIFRNQFINVYAPKYDTAGKFWP 188
Query: 624 HVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPL 683
+H+ +I SL++ ++ +G+ + KK + ++ L LP+LT F++YC+ RF P F Y
Sbjct: 189 IIHNSMIFSLVLMHIIAVGIFALKKLSLASTLTMPLPVLTLLFNEYCRKRFLPIFVAYSA 368
Query: 684 EEAMAKD 690
E +D
Sbjct: 369 ESLKKRD 389
>TC224207 similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21 protein),
partial (27%)
Length = 619
Score = 104 bits (260), Expect = 1e-22
Identities = 50/135 (37%), Positives = 78/135 (57%)
Frame = +1
Query: 588 LLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTK 647
+LPF+L++F L Y+++R+Q++ VY +YE+ FWP VHS I SL++ ++ +GL K
Sbjct: 10 ILPFLLIYFCLGYIIFRNQLLKVYVPKYETGGEFWPTVHSSTIFSLILMHIIAIGLFGLK 189
Query: 648 KAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLA 707
K ++ L+ LPILT F++YCQ RF P F+ Y E + KD ++ LA
Sbjct: 190 KLPLASILILPLPILTLLFNEYCQKRFFPIFKNYSAECLIKKDRADQNEHNMSEFYDKLA 369
Query: 708 DAYLHPIFRSFEVEE 722
+AY P + E
Sbjct: 370 NAYNDPALMRVKYSE 414
>BM527121 similar to GP|20146379|dbj OJ1029_F04.25 {Oryza sativa (japonica
cultivar-group)}, partial (12%)
Length = 421
Score = 104 bits (259), Expect = 2e-22
Identities = 63/138 (45%), Positives = 75/138 (53%), Gaps = 57/138 (41%)
Frame = +3
Query: 569 PSLQLYFLLGIVYAVVTPILLPFILVFFALAYLV-------------------------- 602
P +QLYFLLG+VYAVVTP LLP+I+VFF LAY+V
Sbjct: 6 PQIQLYFLLGLVYAVVTPFLLPYIIVFFGLAYVVYRHQVKVSS*KVFCFNAI*SLFLQTD 185
Query: 603 YRH-------------------------------QIINVYNQQYESAAAFWPHVHSRIIA 631
+RH QIINVYNQ+YESAAAFWP VH RII
Sbjct: 186 HRHEKF*G*IFTMYYSIFNN*IFCYVA*CLKSKWQIINVYNQEYESAAAFWPDVHGRIIF 365
Query: 632 SLLISQLLMLGLLSTKKA 649
+L+ISQLL++GLLSTK+A
Sbjct: 366 ALVISQLLLMGLLSTKEA 419
>AW780629
Length = 341
Score = 93.2 bits (230), Expect = 4e-19
Identities = 45/56 (80%), Positives = 47/56 (83%)
Frame = +2
Query: 499 IPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRG 554
IPR IGV IPMKAT FMTYIMVDGWAGIAGEILRLKPLVIYHL+NMF+ D G
Sbjct: 173 IPRKIGVFIPMKATLFMTYIMVDGWAGIAGEILRLKPLVIYHLRNMFLSNGYLDEG 340
>TC233821 weakly similar to UP|Q75GI5 (Q75GI5) Expressed protein, partial
(4%)
Length = 671
Score = 82.4 bits (202), Expect = 7e-16
Identities = 57/173 (32%), Positives = 90/173 (51%), Gaps = 5/173 (2%)
Frame = +2
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTY 69
SAAINI A L F++L+ QP N +Y+ + S R F +LN +
Sbjct: 38 SAAINIGLALVTLPLFSVLKKQPSNAPIYYAR------PLSRRHHLPFDDSSSSLN--RF 193
Query: 70 LTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVN----- 124
L L W+ +A R++E+EI+ GLD+ V +R++ G+K F ++V L++L+P N
Sbjct: 194 LPSLAWLSRAFRVTEDEIVQDHGLDALVIIRLFKFGIKFFTVCSLVGLVVLLPTNYGAQE 373
Query: 125 VSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYK 177
V +G+ F +D +ISNV S+R +VH +L+ +LLYK
Sbjct: 374 VQNGSYF---------TMDSFTISNVKRGSNRLWVHFAFLCFISLYGMYLLYK 505
>AI988237
Length = 411
Score = 70.9 bits (172), Expect = 2e-12
Identities = 33/36 (91%), Positives = 35/36 (96%)
Frame = +2
Query: 572 QLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQI 607
QLYFLLGIVYAVVTPILLPFI+VFFA AYLVYRHQ+
Sbjct: 2 QLYFLLGIVYAVVTPILLPFIVVFFAFAYLVYRHQV 109
>TC212841 similar to UP|Q75LY1 (Q75LY1) Expressed protein (With alternative
splicing), partial (5%)
Length = 472
Score = 70.5 bits (171), Expect = 3e-12
Identities = 42/128 (32%), Positives = 67/128 (51%), Gaps = 1/128 (0%)
Frame = +2
Query: 568 IPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHS 627
IP + L L+GIVYAVV P+LLPF++++F L Y+VY
Sbjct: 29 IPLVSLSVLIGIVYAVVAPLLLPFLILYFCLGYVVY------------------------ 136
Query: 628 RIIASLLISQLLMLGLLSTK-KAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEA 686
++Q+ M+GL K K A S + ++ + T+ F++YC+ RF P+F Y L++A
Sbjct: 137 -------VNQITMVGLFGLKLKPAASISTIPLI-LFTWMFNEYCKMRFLPSFHHYTLQDA 292
Query: 687 MAKDLLEK 694
D L++
Sbjct: 293 AENDELDE 316
>BU762843
Length = 423
Score = 65.5 bits (158), Expect = 9e-11
Identities = 41/131 (31%), Positives = 65/131 (49%), Gaps = 1/131 (0%)
Frame = +2
Query: 21 FLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTYLTFLNWMPQAL 80
F +++LR QP N VY P+ G TS R + + + W+ +A
Sbjct: 5 FFTLYSILRKQPSNYEVYVPRLLTEG--TSKRRS--------RFKLERLIPSAGWVAKAW 154
Query: 81 RMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLK-RDLVV 139
R+SE E+ + +GLD VF+R+ T LK F ++ + +L+PVN L + D V
Sbjct: 155 RLSEEELFSLSGLDGVVFMRMITFSLKTFTFAGIIGIFVLLPVNCWGNQLKDIDIADFVN 334
Query: 140 SDIDKLSISNV 150
+ +D +ISNV
Sbjct: 335 NSLDVFTISNV 367
>BE555530
Length = 347
Score = 63.5 bits (153), Expect = 3e-10
Identities = 26/48 (54%), Positives = 37/48 (76%)
Frame = +1
Query: 155 SRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTV 202
+RF+ HI + Y FT W C++L KEY+ +A MRL FLA+++RR +QFTV
Sbjct: 1 TRFWAHILVAYAFTFWTCYILLKEYEKVASMRLQFLAAEKRRPDQFTV 144
>CO978729
Length = 377
Score = 61.6 bits (148), Expect = 1e-09
Identities = 31/49 (63%), Positives = 33/49 (67%), Gaps = 2/49 (4%)
Frame = -2
Query: 749 SPLHDHVHQPSPPQHVNQYSHYPTSPPGYY--YHPPPPPHYAYQYQDEP 795
SP H H+H+PS PQ YS TSPPGYY YHPP PPHY YQY EP
Sbjct: 376 SPPH-HIHEPSVPQ----YSVXQTSPPGYYYQYHPPSPPHYVYQY*VEP 245
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,102,969
Number of Sequences: 63676
Number of extensions: 672586
Number of successful extensions: 12280
Number of sequences better than 10.0: 341
Number of HSP's better than 10.0 without gapping: 7146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8987
length of query: 795
length of database: 12,639,632
effective HSP length: 105
effective length of query: 690
effective length of database: 5,953,652
effective search space: 4108019880
effective search space used: 4108019880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0020.6


