

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.7
(1703 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
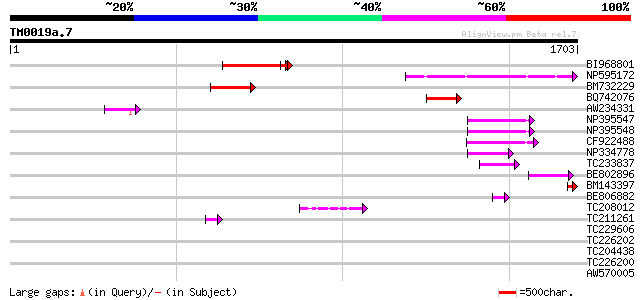
Sequences producing significant alignments: (bits) Value
BI968801 similar to GP|24461860|gb CTV.20 {Poncirus trifoliata},... 257 4e-68
NP595172 polyprotein [Glycine max] 206 7e-53
BM732229 183 5e-46
BQ742076 126 9e-29
AW234331 95 3e-19
NP395547 reverse transcriptase [Glycine max] 88 4e-17
NP395548 reverse transcriptase [Glycine max] 79 1e-14
CF922488 79 2e-14
NP334778 reverse transcriptase [Glycine max] 67 5e-11
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 61 5e-09
BE802896 58 4e-08
BM143397 51 4e-06
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 45 4e-04
TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting prote... 44 8e-04
TC211261 44 8e-04
TC229606 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein H... 39 0.015
TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 39 0.020
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 39 0.020
TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 39 0.020
AW570005 39 0.020
>BI968801 similar to GP|24461860|gb CTV.20 {Poncirus trifoliata}, partial
(0%)
Length = 660
Score = 257 bits (656), Expect = 4e-68
Identities = 127/206 (61%), Positives = 159/206 (76%), Gaps = 1/206 (0%)
Frame = -1
Query: 638 NMTMVATAYVTANNCPESLIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNP 697
+MTMVATAY T++ C I+++LVAGF GQLK WWDNYLT +EK +I +AVKTD G
Sbjct: 657 HMTMVATAYQTSHECSXXXIIDILVAGFSGQLKRWWDNYLTNEEK*KIYSAVKTDLNGKV 478
Query: 698 IM-EDGKFISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTR 756
I +D K I DAVN+LIFTIAQHF+GDPSL KD+S +LLSNLKC++L DFRWY+DTFLTR
Sbjct: 477 ITNDDDKEILDAVNTLIFTIAQHFIGDPSLWKDRSAELLSNLKCRTLADFRWYRDTFLTR 298
Query: 757 VYTREDSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVAL 816
VYTREDSQQ F KEKFLAGLP+S GDKVR+K+RSQ+ G+IPY +LSYGQLI+ +Q+VA
Sbjct: 297 VYTREDSQQPF*KEKFLAGLPRSLGDKVRDKIRSQSANGDIPYESLSYGQLISYVQKVAX 118
Query: 817 KICQDDKIQQQLTKEKSQNRRDLGTF 842
K + + +++ K N+ F
Sbjct: 117 KNLSG*QNSKAISQRKGSNKEGFRFF 40
Score = 58.2 bits (139), Expect = 3e-08
Identities = 25/36 (69%), Positives = 31/36 (85%)
Frame = -2
Query: 812 QRVALKICQDDKIQQQLTKEKSQNRRDLGTFCEQFG 847
+R KICQDDKIQ+QL KEK+Q ++DLG+FCEQFG
Sbjct: 131 KR*XXKICQDDKIQRQLAKEKAQTKKDLGSFCEQFG 24
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 206 bits (524), Expect = 7e-53
Identities = 154/528 (29%), Positives = 261/528 (49%), Gaps = 12/528 (2%)
Frame = +1
Query: 1188 EIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGITVQHL-GQPILFKFSEPPIDKTLN 1246
E++ P L++ +I+G+ ++ L P+ D +T++ + E + T
Sbjct: 1378 EVKVPVYLLQISGADVILGSTWLATLGPHVADYAALTLKFFQNDKFITLQGEGNSEAT-- 1551
Query: 1247 VISYKEKQINFLKEEISYRTIEEQLQQPSVKSRI-EDILENIQSSICSDLP---NAFWER 1302
+ Q++ + + ++IEE ++ + ED L+++ ++I +L + + +
Sbjct: 1552 -----QAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPTNIDPELAILLHTYAQV 1716
Query: 1303 KSHMVELPYEKDFSD----KQIS--TKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSP 1356
+ LP +++ KQ S K RP + +K I ++L + +I+ S SP
Sbjct: 1717 FAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSP 1896
Query: 1357 WSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDM 1416
+S V K+ G+ R +Y+ LN +P+P +LL H A+ FSK D+
Sbjct: 1897 FSLPILLVKKKD----GSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDL 2064
Query: 1417 KSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFN-PYSKLTIV 1475
+SG+ QI +Q +D+ KTAF G YEW VMPFGL NAP+ FQ +MN+IF K +V
Sbjct: 2065 RSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLV 2244
Query: 1476 YIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPI 1535
+ DD+LI+S + H KHL + + +K++ L +K S T++ +LGH + +
Sbjct: 2245 FFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSME 2424
Query: 1536 NRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHT 1595
N ++ +P K QL+ FLG Y F + I PL D L+KD ++
Sbjct: 2425 NTKVQAVLDWPTPNNVK-QLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDSFLWNNEAE 2601
Query: 1596 NVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNP 1655
++K + P L LP+ I+ETDAS IG G +L Q IA+ SK P
Sbjct: 2602 AAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQ----NGHPIAYFSKKLAP 2769
Query: 1656 AQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKSAKEILQKDV 1703
Q S +E+LAI ++SKF+ L+ KF++ D +S K ++ + +
Sbjct: 2770 RMQKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSL 2913
>BM732229
Length = 421
Score = 183 bits (465), Expect = 5e-46
Identities = 92/139 (66%), Positives = 109/139 (78%), Gaps = 3/139 (2%)
Frame = -3
Query: 602 DLLLEE--SSNFKSFSANNVYEWNIDGENEYGITKILQNMTMVATAYVTANNCPESLIVE 659
DLLLEE +NFKSFSANN+YEWNID + EY I LQ+MTMVATAY T++ C E I++
Sbjct: 419 DLLLEERGENNFKSFSANNIYEWNIDAQTEYNIMNTLQHMTMVATAYQTSHECSEETIID 240
Query: 660 VLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIM-EDGKFISDAVNSLIFTIAQ 718
+LVAGF GQLKGWWDNYLT +EK +I +AVKTD G I +D K I DAVN+LIFTIAQ
Sbjct: 239 ILVAGFSGQLKGWWDNYLTNEEKSKIYSAVKTDLNGKVITNDDDKEIPDAVNTLIFTIAQ 60
Query: 719 HFVGDPSLIKDKSGDLLSN 737
HF+GDPSL D+S +LLSN
Sbjct: 59 HFIGDPSLW*DRSAELLSN 3
>BQ742076
Length = 429
Score = 126 bits (316), Expect = 9e-29
Identities = 60/106 (56%), Positives = 80/106 (74%)
Frame = -2
Query: 1251 KEKQINFLKEEISYRTIEEQLQQPSVKSRIEDILENIQSSICSDLPNAFWERKSHMVELP 1310
K QINFLK+E+S+ I+ QL++P +K +I +L++IQS+IC DL AF RK H++ LP
Sbjct: 323 KINQINFLKDEVSFSNIQIQLEKPQMKEKISSLLKHIQSTICFDLARAF*NRKKHILNLP 144
Query: 1311 YEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSP 1356
+KDF +K I TK+RPI MNEELL +CQKEI DL K LIR+SK+P
Sbjct: 143 CDKDFKEKHIPTKSRPI*MNEELLQYCQKEIKDLFNKGLIRKSKNP 6
>AW234331
Length = 370
Score = 94.7 bits (234), Expect = 3e-19
Identities = 48/119 (40%), Positives = 71/119 (59%), Gaps = 12/119 (10%)
Frame = +2
Query: 286 FVIDKGSIKADFYSDRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQDY 345
F IDK +++ DFYS QR+WFF Y+G RK+I++KFY F+E+ + NI FF+WF Y
Sbjct: 11 FSIDKETLRKDFYSSENEPQRRWFFQHYKGTNRKQIKDKFYKFVERVKINILFFDWFHAY 190
Query: 346 ANKNWKSYPFQIDMV---------KWQHKDGKETLSNLPPKTPFLL---KGAQSTPVLA 392
A + YP++ D++ W+ KDG+ S LPP T + L K + + PV+A
Sbjct: 191 AIRKDIDYPWKQDIIGDPTTNVITNWKIKDGELIQSELPPATQYQLPNVKDSNNKPVMA 367
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 87.8 bits (216), Expect = 4e-17
Identities = 58/201 (28%), Positives = 94/201 (45%), Gaps = 1/201 (0%)
Frame = +1
Query: 1376 RLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAF 1435
R+ I+Y+ LN+A YP+P +L R + D SG+ QI + +D+ KTAF
Sbjct: 163 RMCIDYRKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAF 342
Query: 1436 TVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNP-YSKLTIVYIDDVLIFSQTLDQHFKHL 1494
T PF + + MPFGL NA + FQR M IF+ K V++DD F + +L
Sbjct: 343 TCPFSVFAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANL 522
Query: 1495 NTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQ 1554
+ +++ L ++ K + LGH I + I + ++ DK P + K
Sbjct: 523 EKVLQRCEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVK-G 699
Query: 1555 LQRFLGCLNYVADFCPQLSTI 1575
+ FLG + + F + +
Sbjct: 700 IHSFLGHVGFYRRFIKDFTKV 762
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 79.3 bits (194), Expect = 1e-14
Identities = 58/201 (28%), Positives = 91/201 (44%), Gaps = 1/201 (0%)
Frame = +1
Query: 1376 RLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAF 1435
+L I+Y+ LN+A +P+P +L R + D G+ QI + KD+ K AF
Sbjct: 163 KLCIDYRKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAF 342
Query: 1436 TVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTI-VYIDDVLIFSQTLDQHFKHL 1494
T PFG + + +PFGL NAP+ FQ M IF + +I V++DD +F +L+ K L
Sbjct: 343 TCPFGVFAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKL 522
Query: 1495 NTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQ 1554
+ L ++ K + LGH I I I+ +K P K
Sbjct: 523 EMVLQRCVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVK-G 699
Query: 1555 LQRFLGCLNYVADFCPQLSTI 1575
++ FLG + F + +
Sbjct: 700 IRSFLGQARFYRRFIKDFTKV 762
>CF922488
Length = 741
Score = 78.6 bits (192), Expect = 2e-14
Identities = 63/219 (28%), Positives = 107/219 (48%), Gaps = 3/219 (1%)
Frame = +3
Query: 1371 ERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDK 1430
E G + ++Y+ LN A ++P+P+ L+ FS D SG+ QI++ +D
Sbjct: 12 EDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAPEDM 191
Query: 1431 YKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIF-NPYSKLTIVYIDDVLIFSQTLDQ 1489
KT F +G + + M FGLKN + +QR M +F + K VY+DD+++ S+T ++
Sbjct: 192 EKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRTEEE 371
Query: 1490 HFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIH--QGTIIPINRAIEFTDKFPD 1547
H +L +++ L ++ K +F+ K R L I +G + N+ +
Sbjct: 372 HLVNLRKLFRRLRKYRLRLNPAK-CMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMAKP 548
Query: 1548 QIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKD 1586
+ Q+Q FLG LNY+ F L +PL L K+
Sbjct: 549 H--TEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKN 659
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 67.4 bits (163), Expect = 5e-11
Identities = 39/138 (28%), Positives = 72/138 (51%), Gaps = 1/138 (0%)
Frame = +3
Query: 1376 RLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAF 1435
R+ ++Y+ LN+A +P+P+ L+A +FS D SG+ QI++ +D KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 1436 TVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTI-VYIDDVLIFSQTLDQHFKHL 1494
+G + + VM FGLKN + + R M +F I Y+D+++ S+ ++H +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 1495 NTFISVIKRNGLAVSKTK 1512
+++ L ++ K
Sbjct: 363 QNLFGQLRKYRLRLNPRK 416
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 60.8 bits (146), Expect = 5e-09
Identities = 38/120 (31%), Positives = 62/120 (51%), Gaps = 1/120 (0%)
Frame = +2
Query: 1411 FSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYS 1470
FS D SG+ QI + +D KT F +G + + VM FGLKN + +QR M +F+
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 1471 KLTI-VYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQ 1529
I VY+DD++ S+T +H +L +++ L ++ TK + + LG + Q
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
>BE802896
Length = 416
Score = 57.8 bits (138), Expect = 4e-08
Identities = 39/135 (28%), Positives = 68/135 (49%), Gaps = 1/135 (0%)
Frame = -2
Query: 1558 FLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNP 1616
FLG + F + PL + L+K+ +D +K + P + P+
Sbjct: 412 FLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPDW 233
Query: 1617 QAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISK 1676
A + DAS+ G +L QK+ ++I ++S+ + AQ NY+T +KE+LAIV ++ K
Sbjct: 232 TAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALEK 53
Query: 1677 FQSDLINQKFLVCVD 1691
F S L+ + +V +D
Sbjct: 52 FHSYLLGTRIIVYID 8
>BM143397
Length = 408
Score = 51.2 bits (121), Expect = 4e-06
Identities = 24/30 (80%), Positives = 27/30 (90%)
Frame = -3
Query: 1674 ISKFQSDLINQKFLVCVDCKSAKEILQKDV 1703
ISKFQSDL+NQK L+ VDCK AK+ILQKDV
Sbjct: 406 ISKFQSDLLNQKILIRVDCKLAKDILQKDV 317
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 44.7 bits (104), Expect = 4e-04
Identities = 21/51 (41%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Frame = -1
Query: 1451 LKNAPSEFQRIMNEIF-NPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISV 1500
L NAP+ FQ +MN IF + K +V+ DD+L++S T +H HL V
Sbjct: 153 LTNAPTSFQCLMNHIFQHALRKYVLVFFDDILVYSSTWHEHLCHLEVVFKV 1
>TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting protein 117
(Fragment), partial (38%)
Length = 993
Score = 43.5 bits (101), Expect = 8e-04
Identities = 53/214 (24%), Positives = 93/214 (42%), Gaps = 10/214 (4%)
Frame = +1
Query: 872 SSRND--HRKPKPRSKPQSS-QIPKNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRRIS 928
SS ND + KP S P + ++P + P+ CY C K GH+SR C+ ++
Sbjct: 46 SSENDTLDQGKKPGSGPSDAPKVPSDAPKI---------CYKCKKAGHLSRDCK-EQPDG 195
Query: 929 ELHLEP--EIEDKINNLLIQTSDEEESNPSDSEVSEDLNQIQNDDSQSSSSVNTLSINTL 986
LH E E+ + I TS + + +D+N+I ++ + + V+ L+ N L
Sbjct: 196 LLHRNAIGEAEENPKSTAIDTSQADRVAMEE----DDINEIGEEEKEKLNDVDYLTGNPL 363
Query: 987 TNEQDLLFRAINSIPDPEEKKIYLERLRSTLEDRPPK----SPITINKFNLRDTFKRLEK 1042
N D+L A+ + Y R++ + P K + +N F+ EK
Sbjct: 364 PN--DILLYAVPVCGPYSAVQSYKYRVK--IIPGPAKKGKAAKTAMNLFSHMSEATTREK 531
Query: 1043 STVKPVTIQDLHFEI-NSLKTEVKSLKQIQKSQQ 1075
+K T +L I ++K L Q+++ Q+
Sbjct: 532 ELMKACTDPELVAAIVGNVKISAAGLTQLKQKQK 633
>TC211261
Length = 520
Score = 43.5 bits (101), Expect = 8e-04
Identities = 24/55 (43%), Positives = 31/55 (55%), Gaps = 3/55 (5%)
Frame = -2
Query: 588 RKETKLYYPRATAPDLLLEE---SSNFKSFSANNVYEWNIDGENEYGITKILQNM 639
R TKLYY + T PDL L+E +S + F + NIDG+ EY T LQ+M
Sbjct: 504 RNSTKLYYQKVTIPDLWLQERDLNSKLQRFRTMGIC**NIDGQIEYNTTHTLQHM 340
>TC229606 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein HUA1, partial
(50%)
Length = 1569
Score = 39.3 bits (90), Expect = 0.015
Identities = 25/88 (28%), Positives = 42/88 (47%), Gaps = 7/88 (7%)
Frame = +1
Query: 506 IPETPEPPETPEPTPEAVSNP--TITAPPPPKTRTSTRNTSTSLNNIED-----GKVESD 558
+P PPE+P P P +S+P ++ PPPP T N +L+ I+ GKV+
Sbjct: 7 LPLPASPPESPPPPPPRISSPPGPLSPPPPPCLPPLTSNAPPTLSTIQQFWVQLGKVKL- 183
Query: 559 SDIQSMKSDVQSIKSIPVNTVIHNSKNQ 586
+Q+ +K +P+ IH + +
Sbjct: 184 GILQTHWPSALDMKVLPI*LYIHRGQER 267
>TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (72%)
Length = 1003
Score = 38.9 bits (89), Expect = 0.020
Identities = 13/24 (54%), Positives = 18/24 (74%)
Frame = +3
Query: 901 SQGKDVTCYNCGKPGHISRYCRLK 924
S G D+ CY CG+PGH +R CR++
Sbjct: 813 SGGSDLKCYECGEPGHFARECRMR 884
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 38.9 bits (89), Expect = 0.020
Identities = 33/115 (28%), Positives = 54/115 (46%), Gaps = 17/115 (14%)
Frame = +1
Query: 866 QQWRKRSSRNDHR-KPKPRSKP----QSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRY 920
+Q+ K +R D R KP R+ P + S+ K E +PS K + C+ C GHI
Sbjct: 766 KQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKKSDE-KPSHSKGIQCHGCEGYGHIKAE 942
Query: 921 C--RLKRR----------ISELHLEPEIEDKINNLLIQTSDEEESNPSDSEVSED 963
C LK++ +E E + + +N L + E+S+ +DSE++ D
Sbjct: 943 CPTHLKKQRKGLSVCRSDDTESEQESDSDRDVNALTGRFESAEDSSDTDSEITFD 1107
>TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (89%)
Length = 936
Score = 38.9 bits (89), Expect = 0.020
Identities = 13/24 (54%), Positives = 18/24 (74%)
Frame = +2
Query: 901 SQGKDVTCYNCGKPGHISRYCRLK 924
S G D+ CY CG+PGH +R CR++
Sbjct: 317 SGGSDLKCYECGEPGHFARECRMR 388
>AW570005
Length = 413
Score = 38.9 bits (89), Expect = 0.020
Identities = 38/131 (29%), Positives = 56/131 (42%), Gaps = 3/131 (2%)
Frame = -2
Query: 1555 LQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDP---PP*SDIHTNVVKQIKLRVKNLPCL 1611
L+ FL + F + + PL L KD P +D+ +K + V N L
Sbjct: 388 LRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDSFVWSPEADVAFQALKNV---VTNTLVL 218
Query: 1612 YLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIV 1671
LP+ +ETDAS G +L Q + IAF SK + P ST E+ AI
Sbjct: 217 ALPDFTKPFTVETDASGSDMGAVLSQ----EGHPIAFFSKEFCPKLVRSSTYVHELAAIT 50
Query: 1672 LSISKFQSDLI 1682
+ K++ L+
Sbjct: 49 NVVKKWRQYLL 17
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 73,761,575
Number of Sequences: 63676
Number of extensions: 1116455
Number of successful extensions: 15053
Number of sequences better than 10.0: 280
Number of HSP's better than 10.0 without gapping: 9489
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 13704
length of query: 1703
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1593
effective length of database: 5,635,272
effective search space: 8976988296
effective search space used: 8976988296
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0019a.7


