

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
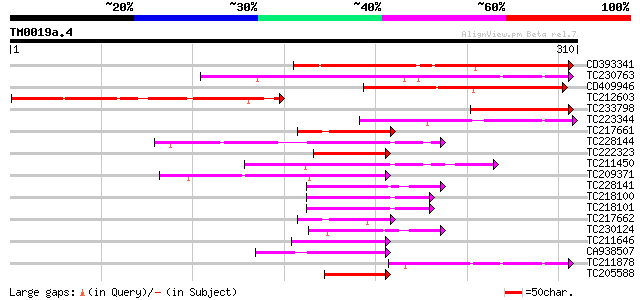
Score E
Sequences producing significant alignments: (bits) Value
CD393341 229 9e-61
TC230763 166 1e-41
CD409946 130 8e-31
TC212603 homologue to UP|Q9AGG0 (Q9AGG0) NLPE2, partial (6%) 124 7e-29
TC233798 83 2e-16
TC223344 82 4e-16
TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%) 74 8e-14
TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protei... 65 3e-11
TC222323 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (39%) 65 4e-11
TC211450 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-... 61 6e-10
TC209371 similar to PIR|T52104|T52104 GATA-binding transcription... 61 7e-10
TC228141 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F2... 59 2e-09
TC218100 similar to UP|Q9AVU3 (Q9AVU3) GATA-1 zinc finger protei... 59 2e-09
TC218101 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F2... 59 3e-09
TC217662 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%) 59 3e-09
TC230124 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-... 59 3e-09
TC211646 homologue to GB|AAP37701.1|30725358|BT008342 At4g32890 ... 57 1e-08
CA938507 homologue to PIR|T52104|T521 GATA-binding transcription... 57 1e-08
TC211878 weakly similar to UP|LIB5_HUMAN (O75023) Leukocyte immu... 55 3e-08
TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcriptio... 52 3e-07
>CD393341
Length = 612
Score = 229 bits (585), Expect = 9e-61
Identities = 122/158 (77%), Positives = 133/158 (83%), Gaps = 5/158 (3%)
Frame = -3
Query: 156 RYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAA 215
RYSQRSPRNNN SS T RVCSDCNTS+TPLWR+GP GPKSLCNACGIRQRKARRAMAEAA
Sbjct: 598 RYSQRSPRNNNGSS-TPRVCSDCNTSTTPLWRTGPKGPKSLCNACGIRQRKARRAMAEAA 422
Query: 216 NGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKS----TSSNAGSSQETKKLECFK 271
NGL TPI A KTR+ HNKEKK R NHFAQFKNK KS T++ GSS+ +KLE F
Sbjct: 421 NGLVTPI--ACEKTRL-HNKEKKSRMNHFAQFKNKYKSTTTTTTTTVGSSEGVRKLEYFN 251
Query: 272 DFAISLR-NNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+FAISLR NNS F+Q+FPRDEVAEAALLLMDLSCG+VH
Sbjct: 250 NFAISLRSNNSDFEQMFPRDEVAEAALLLMDLSCGFVH 137
>TC230763
Length = 1058
Score = 166 bits (421), Expect = 1e-41
Identities = 101/215 (46%), Positives = 127/215 (58%), Gaps = 11/215 (5%)
Frame = +1
Query: 105 EDIKNVHGSAKWMSSKMRLMKKMMSTPATD---KANNSTTIPISPRIQNQGHENRYSQRS 161
E + GS KWM +KMR+M+KM+ + TD ++N+TT + Q S
Sbjct: 31 ESVAAEDGSLKWMPAKMRIMRKMLVSDQTDTYTNSDNNTTHKFDDQKQQLSSPLGTDNSS 210
Query: 162 PRN-NNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEA---ANG 217
N +N+S+NT RVCSDC+T+ TPLWRSGP GPKSLCNACGIRQRKARRAMA A A+G
Sbjct: 211 SNNYSNHSNNTVRVCSDCHTTKTPLWRSGPRGPKSLCNACGIRQRKARRAMAAAAASASG 390
Query: 218 LATPI----NTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDF 273
T I + + ++ KEKK R AQ K K K +A +SQ K F+D
Sbjct: 391 NGTVIVEAKKSVKGRNKLQKKKEKKTRTEGAAQMKKKRKLGVGSAKASQSRNKFG-FEDL 567
Query: 274 AISLRNNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+ LR N QVFP+DE EAA+LLM LS G VH
Sbjct: 568 TLRLRKNLAMHQVFPQDE-KEAAILLMALSYGLVH 669
>CD409946
Length = 631
Score = 130 bits (327), Expect = 8e-31
Identities = 73/116 (62%), Positives = 88/116 (74%), Gaps = 4/116 (3%)
Frame = -1
Query: 194 KSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSK- 252
+SLCNACGIRQRKARRAMAEA NG A +N++STK RVHH KEKK R NHFA+F+ K K
Sbjct: 457 QSLCNACGIRQRKARRAMAEAVNGFAPSVNSSSTKIRVHH-KEKKSRTNHFARFRLKCKL 281
Query: 253 -STSSNAGSSQETKKLECFKDFAISLRNNSTF-QQVFP-RDEVAEAALLLMDLSCG 305
+TS+ G+SQ+ DF +SLR++S QQVFP DEVA+AA+LLMDLSCG
Sbjct: 280 ATTSTAEGTSQQENVKIDLNDFGLSLRDSSALKQQVFPIMDEVAQAAMLLMDLSCG 113
>TC212603 homologue to UP|Q9AGG0 (Q9AGG0) NLPE2, partial (6%)
Length = 433
Score = 124 bits (310), Expect = 7e-29
Identities = 93/158 (58%), Positives = 102/158 (63%), Gaps = 9/158 (5%)
Frame = +3
Query: 2 TPVSLNPPGPSIQ-GQNHLFN-SLNNQDYHASLFNILDRRQ-GIGIGELRENDHQDDKLV 58
TP SLNPPGPSIQ GQN LFN S NNQD + FNI D RQ I IG LREN QDDK++
Sbjct: 3 TPYSLNPPGPSIQAGQNQLFNISPNNQDCR-TFFNIFDPRQTSIEIGGLRENYRQDDKMI 179
Query: 59 VWHDGSSSSSSSNHLYNSTFISPQP-VMDNPSSSTCD-PNLSFSKMEEEDIKNVHGSA-K 115
+ HDGSSS+ +S S ISP+ VM +P SS CD NL EEE N HGS K
Sbjct: 180 L-HDGSSSNCNS-----SFNISPETVVMVDPLSSACDRRNL---PSEEESKNNDHGSGNK 332
Query: 116 WMSSKMRLMKKMMS---TPATDKANNSTTIPISPRIQN 150
WMSSKMRLMKKMM +P TDKA NS SPR QN
Sbjct: 333 WMSSKMRLMKKMMRPSISPTTDKAINS-----SPRFQN 431
>TC233798
Length = 423
Score = 82.8 bits (203), Expect = 2e-16
Identities = 42/57 (73%), Positives = 50/57 (87%), Gaps = 1/57 (1%)
Frame = +2
Query: 253 STSSNAGSSQETKKLECFKDFAISLR-NNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+T+++AGSS+ +KLE KDFAISLR NNS F+Q FPRDEVAEAALLLMDLSCG+VH
Sbjct: 11 TTTTSAGSSEGVRKLEYLKDFAISLRSNNSDFEQGFPRDEVAEAALLLMDLSCGFVH 181
>TC223344
Length = 639
Score = 81.6 bits (200), Expect = 4e-16
Identities = 51/123 (41%), Positives = 66/123 (53%), Gaps = 4/123 (3%)
Frame = +3
Query: 192 GPKSLCNACGIRQRKARRAMAEAANGLAT-PINTAST---KTRVHHNKEKKPRANHFAQF 247
GPKSLCNACGIRQRK RRA+A AA T P+ + K H+K K + Q
Sbjct: 6 GPKSLCNACGIRQRKVRRAIAAAATSNGTNPVEAEKSQVKKGNTLHSKGMKSKTEGAQQM 185
Query: 248 KNKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPRDEVAEAALLLMDLSCGYV 307
K K ++ K+ F+D + L N Q+VFP+DE EAA+LLM LS G +
Sbjct: 186 KKNRKL------GARYRKRFGAFEDLTVRLSKNFALQRVFPQDE-KEAAILLMALSYGLL 344
Query: 308 HSY 310
H +
Sbjct: 345 HGF 353
>TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%)
Length = 765
Score = 73.9 bits (180), Expect = 8e-14
Identities = 33/54 (61%), Positives = 38/54 (70%)
Frame = +2
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAM 211
S SP +NN T C+DC T+ TPLWR GP GPKSLCNACGIR RK +RA+
Sbjct: 248 SGNSPSSNNEQKKT---CADCGTTKTPLWRGGPAGPKSLCNACGIRSRKKKRAI 400
>TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(27%)
Length = 1018
Score = 65.5 bits (158), Expect = 3e-11
Identities = 46/161 (28%), Positives = 71/161 (43%), Gaps = 2/161 (1%)
Frame = +1
Query: 80 SPQPVMD--NPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKAN 137
+P+P M+ +P+SS N+ + + + + A+ S + + K+ S K
Sbjct: 73 NPRPAMNLISPASSFVGENMQPNVISSKASSDSENFAE--SQLVPIFLKLASGEPKKKNK 246
Query: 138 NSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLC 197
+P++P NN N+S R C C + TP WR+GP GPK+LC
Sbjct: 247 VKVPLPVAPA---------------DNNQNASQPVRKCMHCEITKTPQWRAGPMGPKTLC 381
Query: 198 NACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKK 238
NACG+R + R + A+P S VH N KK
Sbjct: 382 NACGVRYKSGR--LFPEYRPAASPTFCPS----VHSNSHKK 486
>TC222323 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (39%)
Length = 368
Score = 65.1 bits (157), Expect = 4e-11
Identities = 26/42 (61%), Positives = 31/42 (72%)
Frame = +2
Query: 167 NSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
N + + C+DC T+ TPLWR GP GPK+LCNACGIR RK R
Sbjct: 164 NVNEKKKCCADCKTTKTPLWRGGPAGPKTLCNACGIRYRKRR 289
>TC211450 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(23%)
Length = 825
Score = 61.2 bits (147), Expect = 6e-10
Identities = 41/143 (28%), Positives = 68/143 (46%), Gaps = 4/143 (2%)
Frame = +2
Query: 129 STPATDKANNSTTIPISPRIQNQGHENRYSQR----SPRNNNNSSNTTRVCSDCNTSSTP 184
S ++D N + ++ +P+ + H+ + + S + N S R C C + TP
Sbjct: 38 SKASSDSENFAESVIKAPKQASGEHKKKKKIKVTFPSGQERNAPSQAIRKCLHCEITKTP 217
Query: 185 LWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHF 244
WR+GP GPK+LCNACG+R + R + A+P A+ +H N KK
Sbjct: 218 QWRAGPMGPKTLCNACGVRYKSGR--LFPEYRPAASPTFCAA----MHSNSHKK-----V 364
Query: 245 AQFKNKSKSTSSNAGSSQETKKL 267
+ +NK+ + S A S + +L
Sbjct: 365 LEMRNKTGTKSGFATVSAASPEL 433
>TC209371 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (40%)
Length = 857
Score = 60.8 bits (146), Expect = 7e-10
Identities = 38/137 (27%), Positives = 65/137 (46%), Gaps = 11/137 (8%)
Frame = +2
Query: 83 PVMDNPSSSTCDPN--LSFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNST 140
P N S + DPN L+F + ++ + +++++ + + PA+ ++ +
Sbjct: 107 PPPQNSPSISHDPNFFLNFPSVPSDEAVELEWLSQFVNDEATSFHNI-PPPASIGSHTTP 283
Query: 141 TIPISPRIQNQGHENRYSQRSP---------RNNNNSSNTTRVCSDCNTSSTPLWRSGPN 191
+ + R N + S SP R + + + R CS C T TP WR+GP
Sbjct: 284 FLSNNNRNDNNNEYPKSSSSSPVLAGKSRARREGSVTGDGVRRCSHCATDKTPQWRTGPL 463
Query: 192 GPKSLCNACGIRQRKAR 208
GPK+LCNACG+R + R
Sbjct: 464 GPKTLCNACGVRFKSGR 514
>TC228141 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F28P10_210
{Arabidopsis thaliana;} , partial (23%)
Length = 1143
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/76 (39%), Positives = 40/76 (52%)
Frame = +3
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPI 222
R+++ S R C C + TP WR GP GPK+LCNACG+R R R AE P
Sbjct: 726 RSSSQESVAPRKCLHCEVTKTPQWREGPMGPKTLCNACGVRYRSG-RLFAE-----YRPA 887
Query: 223 NTASTKTRVHHNKEKK 238
++ + +H N KK
Sbjct: 888 SSPTFVASLHSNSHKK 935
>TC218100 similar to UP|Q9AVU3 (Q9AVU3) GATA-1 zinc finger protein, partial
(27%)
Length = 1284
Score = 59.3 bits (142), Expect = 2e-09
Identities = 28/70 (40%), Positives = 38/70 (54%)
Frame = +2
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPI 222
R+++ S + R C C + TP WR GP GPK+LCNACG+R R R + A+P
Sbjct: 863 RSSSPESGSPRKCMHCEVTKTPQWREGPVGPKTLCNACGVRYRSGR--LFPEYRPAASPT 1036
Query: 223 NTASTKTRVH 232
AS + H
Sbjct: 1037FVASLHSNCH 1066
>TC218101 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F28P10_210
{Arabidopsis thaliana;} , partial (23%)
Length = 638
Score = 58.9 bits (141), Expect = 3e-09
Identities = 28/70 (40%), Positives = 37/70 (52%)
Frame = +1
Query: 163 RNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPI 222
R+++ S R C C + TP WR GP GPK+LCNACG+R R R + A+P
Sbjct: 181 RSSSPESGPPRKCMHCEVTKTPQWREGPMGPKTLCNACGVRYRSGR--LFPEYRPAASPT 354
Query: 223 NTASTKTRVH 232
AS + H
Sbjct: 355 FVASLHSNCH 384
>TC217662 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%)
Length = 663
Score = 58.9 bits (141), Expect = 3e-09
Identities = 33/82 (40%), Positives = 38/82 (46%), Gaps = 28/82 (34%)
Frame = +3
Query: 158 SQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPK----------------------- 194
S SP +NN T C+DC T+ TPLWR GP GPK
Sbjct: 6 SGNSPSSNNEQKKT---CADCGTTKTPLWRGGPAGPKVSSPSISQIAPRGVKPTLDLTLC 176
Query: 195 -----SLCNACGIRQRKARRAM 211
SLCNACGIR RK +RA+
Sbjct: 177 YPFFQSLCNACGIRSRKKKRAI 242
>TC230124 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-3, partial
(26%)
Length = 894
Score = 58.9 bits (141), Expect = 3e-09
Identities = 32/79 (40%), Positives = 43/79 (53%), Gaps = 4/79 (5%)
Frame = +1
Query: 164 NNNNSSNTT----RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLA 219
NNNN+ N R C+ C TP WR+GP GPK+LCNACG+R K+ R + E
Sbjct: 151 NNNNNDNVQHPIPRRCTHCLAQRTPQWRAGPLGPKTLCNACGVRY-KSGRLLPE-----Y 312
Query: 220 TPINTASTKTRVHHNKEKK 238
P + + + +H N KK
Sbjct: 313 RPAKSPTFVSYLHSNSHKK 369
>TC211646 homologue to GB|AAP37701.1|30725358|BT008342 At4g32890 {Arabidopsis
thaliana;} , partial (26%)
Length = 778
Score = 57.0 bits (136), Expect = 1e-08
Identities = 24/54 (44%), Positives = 30/54 (55%)
Frame = +2
Query: 155 NRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
N S S + + R C C T TP WR+GP GPK+LCNACG+R + R
Sbjct: 197 NTKSSSSSHSGAEGGSEGRKCLHCATDKTPQWRTGPMGPKTLCNACGVRYKSGR 358
>CA938507 homologue to PIR|T52104|T521 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 449
Score = 56.6 bits (135), Expect = 1e-08
Identities = 27/74 (36%), Positives = 38/74 (50%)
Frame = +1
Query: 135 KANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPK 194
K++ S+ IP S + + R + R CS C + TP WR+GP GPK
Sbjct: 229 KSSLSSNIPCSSAVAGKSRARR------EGSVTGDGGVRRCSHCASEKTPQWRAGPLGPK 390
Query: 195 SLCNACGIRQRKAR 208
+LCNACG+R + R
Sbjct: 391 TLCNACGVRFKSGR 432
>TC211878 weakly similar to UP|LIB5_HUMAN (O75023) Leukocyte
immunoglobulin-like receptor subfamily B member 5
precursor (Leukocyte immunoglobulin-like receptor 8)
(LIR-8) (CD85c antigen), partial (3%)
Length = 669
Score = 55.5 bits (132), Expect = 3e-08
Identities = 46/107 (42%), Positives = 54/107 (49%), Gaps = 6/107 (5%)
Frame = +3
Query: 208 RRAMAEAA-----NGLATPINTASTK-TRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSS 261
RRAMA AA +G S K ++ KEKK R AQ K K K A +S
Sbjct: 3 RRAMAAAAXXALGDGAVIVEAEKSVKGKKLQKKKEKKTRIEGAAQMKMKRK-LGVGAKAS 179
Query: 262 QETKKLECFKDFAISLRNNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
Q K F+D + LR N QVFP+DE EAA+LLM LS G VH
Sbjct: 180 QSRNKFG-FEDLTLRLRKNLAMHQVFPQDE-KEAAILLMALSYGLVH 314
>TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcription
factor-like protein (At5g66320), partial (25%)
Length = 1084
Score = 52.0 bits (123), Expect = 3e-07
Identities = 20/36 (55%), Positives = 24/36 (66%)
Frame = +3
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R CS C TP WR+GP G K+LCNACG+R + R
Sbjct: 558 RRCSHCQVQKTPQWRTGPLGAKTLCNACGVRYKSGR 665
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,534,825
Number of Sequences: 63676
Number of extensions: 244264
Number of successful extensions: 1661
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 1581
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1622
length of query: 310
length of database: 12,639,632
effective HSP length: 97
effective length of query: 213
effective length of database: 6,463,060
effective search space: 1376631780
effective search space used: 1376631780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0019a.4


