

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0016.15
(1469 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
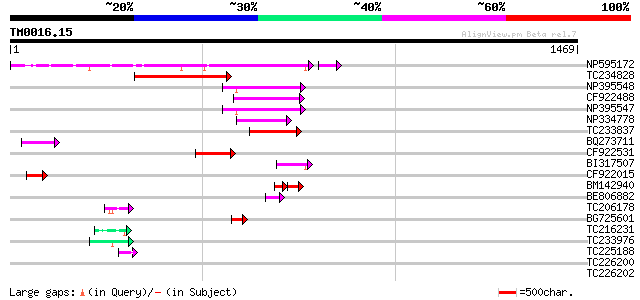
NP595172 polyprotein [Glycine max] 286 4e-77
TC234828 240 3e-63
NP395548 reverse transcriptase [Glycine max] 135 1e-31
CF922488 129 7e-30
NP395547 reverse transcriptase [Glycine max] 129 1e-29
NP334778 reverse transcriptase [Glycine max] 105 1e-22
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 99 1e-20
BQ273711 76 1e-13
CF922531 73 1e-12
BI317507 71 4e-12
CF922015 58 3e-08
BM142940 44 3e-08
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 47 5e-05
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 46 1e-04
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 45 2e-04
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 44 4e-04
TC233976 43 9e-04
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 43 9e-04
TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 43 0.001
TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 43 0.001
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 286 bits (732), Expect(2) = 4e-77
Identities = 245/833 (29%), Positives = 378/833 (44%), Gaps = 46/833 (5%)
Frame = +1
Query: 1 EMMATMAQAVTVQANDSAMRRAAEEAREQHQRQ----REVTLDQNKGLNDFRRHDPPKFS 56
++ A M+Q + N A ++++ QR R V LD P+F
Sbjct: 175 QLNAVMSQVLQRLQNIPMSSHGASNSQKEQQRSSFQVRSVKLDF------------PRFD 318
Query: 57 GGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEEIN 116
G D WI + E+ F T + +L +A+ L D W++ +++ +
Sbjct: 319 GKNVMD----WIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQ----MLQKTEPFSS 474
Query: 117 *NSFRMAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERY 176
+F A LE F SA D + L Q +V E+ + +L D ++
Sbjct: 475 WQAFTRA-LELDFGPSAYDCPRATLFKLNQSA-TVNEYYMQFTALVNRV----DGLSAEA 636
Query: 177 MCKRFVNGLRPDIEDSVRPLGIMRFQSLV-------EKATEVELMKN----RRLNRAGTG 225
+ FV+GL+ +I V+ + V EK T K R + T
Sbjct: 637 ILDCFVSGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTS 816
Query: 226 GPMRSGSQNFQNRGKFQNRRPY-QRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKC 284
+ N +N N P P+ + F + + + Q + + C+ C
Sbjct: 817 ATQKYPPTNQKNDNPKPNLPPLLPTPSTKPFNLRNQN--IKKISPAEIQLRREKNLCYFC 990
Query: 285 GKPGHFARACPDTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTMDGDD 344
+ A CP+ + + +V E A + M G +
Sbjct: 991 DEKFSPAHKCPNRQVMLLQLEETDEDQTDEQVMVTEE----ANMDDDTHHLSLNAMRGSN 1158
Query: 345 TEGVDGLIRGDCEIDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTL 404
GV G IR ++ G + +L D G++ +FI A L+LP+ ++ + L
Sbjct: 1159--GV-GTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLVGNGQIL 1329
Query: 405 LANTACMYCSVTYNNRVYHANLVCLTLKNLDVILGMDWLS-------HYHCL---LDCN* 454
A + + + L + DVILG WL+ Y L N
Sbjct: 1330SAEGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALTLKFFQND 1509
Query: 455 KRVVFPDSVLSEYLFA--NHISVSLNGGSQEYSVLLSLESKGNPE--VDSIP-------- 502
K + SE A +H N S E + L K PE + +P
Sbjct: 1510KFITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPTNIDPELA 1689
Query: 503 -VVKEFTDVFPGDVPG-LPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLL 560
++ + VF VP LPP R+ + AI + G+GP+ + PYR + +++ ++++L
Sbjct: 1690ILLHTYAQVFA--VPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEML 1863
Query: 561 SKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAA 620
+G I+PS SP+ P+LLVKKKDG R C DYR LN +TVK+ +P+P +D+L+D+L GA
Sbjct: 1864VQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQ 2043
Query: 621 IFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTF 680
FSK+DL+SGYHQI V+ ED +KTAFRT +GHYE+LVMPFG+TNAPA F MN+ F
Sbjct: 2044YFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFA 2223
Query: 681 LDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVIS 740
L +FV+VF DDILIYS + +H HL VLQ L++ +L+A SKC F EV +LGH +S
Sbjct: 2224LRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVS 2403
Query: 741 KEGIAVDPAKVETVLSWERPKTV------LGVSLVWRDIIVALSRISPRSLDL 787
G++++ KV+ VL W P V LG++ +R I + + I+ DL
Sbjct: 2404GLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDL 2562
Score = 22.3 bits (46), Expect(2) = 4e-77
Identities = 15/60 (25%), Positives = 27/60 (45%)
Frame = +2
Query: 799 HGLKRAKLVFRR*RNV*RRHLYSFYRNQRNRMKCIVTLRIKVWDVC*CNIGR*LLMLHDN 858
+G+ + K + *RR Y Y+ N + +T ++ VW+ C + LL+ N
Sbjct: 2582 YGIMKQKQHLSSSKKP*RRLQYYHYQIFLNHLFWKLTHQVLVWEQCWDRMATQLLIFQKN 2761
>TC234828
Length = 857
Score = 240 bits (613), Expect = 3e-63
Identities = 123/253 (48%), Positives = 159/253 (62%), Gaps = 1/253 (0%)
Frame = +3
Query: 324 NTARGKRPAAKARVYTMDGDDTEGVDGLIRGDCEIDGNLLSVLFDSGATHSFISRECAIR 383
N G RP +RV+ M G + D LIRG C I LL VL+DSGATHSFIS C R
Sbjct: 96 NNISGGRPKVPSRVFAMSGSEAAASDDLIRGKCLIADKLLDVLYDSGATHSFISHACVER 275
Query: 384 LRLPITALSFDLIVTTP-AKTLLANTACMYCSVTYNNRVYHANLVCLTLKNLDVILGMDW 442
L L T L +D++V+TP ++ + + C+ C + R + A+L+CL L +LDVILGMDW
Sbjct: 276 LGLCATELPYDMVVSTPTSEPVTTSRVCLKCPIIVEGRSFMADLICLPLAHLDVILGMDW 455
Query: 443 LSHYHCLLDCN*KRVVFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIP 502
LS H LDC K +VF V+ + + Y VL S+ + + EV IP
Sbjct: 456 LSTNHIFLDCKEKMLVFGGDVVPSEPLKEDAANEETEDVRTYMVLFSMYVEEDAEVSCIP 635
Query: 503 VVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSK 562
VV EF +VFP DV LPP R++EF ID+VPG P+SIAPYRM+P EL E+K+Q++DLLSK
Sbjct: 636 VVSEFPEVFPDDVCELPPEREVEFIIDVVPGANPVSIAPYRMSPVELAEVKAQVQDLLSK 815
Query: 563 GFIRPSVSPWGAP 575
F+RPS SPWGAP
Sbjct: 816 QFVRPSASPWGAP 854
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 135 bits (340), Expect = 1e-31
Identities = 80/234 (34%), Positives = 122/234 (51%), Gaps = 19/234 (8%)
Frame = +1
Query: 552 LKSQLEDLLSKGFIRP-SVSPWGAPVLLVKKKDGKS------------------RLCVDY 592
++ ++ LL G I P S S W +PVL+V KK+G + +LC+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
R+LN+ T K+ +PLP +D ++++L G A + +D GY+QI V +D +K AF +G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
+ Y +PFG+ NAP F M F +++ + VF+DD ++ ++ L VLQ
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 713 LREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPKTVLGV 766
E L N KC F + E LGH IS GI VD K++ + P V G+
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGI 702
>CF922488
Length = 741
Score = 129 bits (325), Expect = 7e-30
Identities = 70/184 (38%), Positives = 110/184 (59%)
Frame = +3
Query: 579 VKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKT 638
V K+DGK +CVDYR LN + K+++PLP I+ L+D + FS +D SGY+QI++
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 639 EDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKN 698
ED++KT F T +G + Y M FG+ N A + M F + + + V++DD+++ S+
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 699 VVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWE 758
EH +LR++ + LR+ L NP+KC F ++ K L + S GI VD KV+ +L
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 759 RPKT 762
+P T
Sbjct: 543 KPHT 554
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 129 bits (323), Expect = 1e-29
Identities = 78/234 (33%), Positives = 119/234 (50%), Gaps = 19/234 (8%)
Frame = +1
Query: 552 LKSQLEDLLSKGFIRP-SVSPWGAPVLLVKKKDGKS------------------RLCVDY 592
++ ++ LL G I P S S W +PV +V KK G + R+C+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 593 RQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGH 652
R+LN+ T K+ YPLP +D ++ +L + + +D SGY+QI V +D +KTAF +
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 653 YEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
+ Y MPFG+ NA F M F +++ + VF+DD + + +L +VLQ
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 713 LREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPKTVLGV 766
+ L N KC F ++E LGH ISK GI V K++ + P V G+
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGI 702
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 105 bits (263), Expect = 1e-22
Identities = 55/143 (38%), Positives = 85/143 (58%)
Frame = +3
Query: 587 RLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAF 646
R+CVDYR LN+ + K+ +PLP ID LM + A+FS +D SGY+QI++ ED++KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 647 RTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHL 706
T +G + Y VM FG+ N A + M F + + + ++D+++ S+ EH +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 707 RQVLQVLREKELYANPSKCEFWL 729
+ + LR+ L NP KC F L
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 99.0 bits (245), Expect = 1e-20
Identities = 49/133 (36%), Positives = 82/133 (60%)
Frame = +2
Query: 622 FSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFL 681
FS +D SGY+QI + ED++KT F T +G + Y VM FG+ N A + M FH +
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 682 DRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISK 741
+ + V++DD++ S+ EH +L ++ L++ +L NP+KC F ++ K LG ++S+
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 742 EGIAVDPAKVETV 754
+GI +DP KV+ +
Sbjct: 362 KGIEIDPEKVKAL 400
>BQ273711
Length = 409
Score = 76.3 bits (186), Expect = 1e-13
Identities = 40/97 (41%), Positives = 57/97 (58%)
Frame = -2
Query: 32 RQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATY 91
++R+V + +GL FR++ PPKFSG DP+ A LW+ E EKIF + E K+ AT+
Sbjct: 291 QERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATF 112
Query: 92 LLLGDAEYWWKGTRGIMEDNHEEIN*NSFRMAFLEKY 128
+L G+AE WWK + I N+F+ FLE Y
Sbjct: 111 MLQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922531
Length = 602
Score = 72.8 bits (177), Expect = 1e-12
Identities = 44/112 (39%), Positives = 69/112 (61%), Gaps = 10/112 (8%)
Frame = -2
Query: 482 QEYSVLLSLESKGN----PEVDSIP-----VVKEFTDVFPGDVP-GLPPVRDIEFAIDIV 531
Q + +LLS E+ + P ++++P ++ EF D+FP ++P GLPP+R IE ID+V
Sbjct: 337 QSFYLLLSRETSLSTVTIPTIETLPPKVQELLHEFGDIFPKEIPLGLPPLRGIEHQIDLV 158
Query: 532 PGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKD 583
P + YR P E E++SQ+++LL KG+++ S+S VLLV KKD
Sbjct: 157 PRASLPNRPTYRTNPQETKEIESQVKELLEKGWVQESLSLCVVLVLLVPKKD 2
>BI317507
Length = 359
Score = 70.9 bits (172), Expect = 4e-12
Identities = 40/101 (39%), Positives = 60/101 (58%), Gaps = 6/101 (5%)
Frame = -1
Query: 691 DILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAK 750
+ILIYS + H HL VL VL+++ L AN KC F +++LGHVISK+ +A+D K
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 751 VETVLSWERPKTV------LGVSLVWRDIIVALSRISPRSL 785
V++V+ W PK V L ++ +R I +++PR L
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPL 51
>CF922015
Length = 172
Score = 58.2 bits (139), Expect = 3e-08
Identities = 25/55 (45%), Positives = 40/55 (72%)
Frame = -3
Query: 43 GLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDA 97
GL+ F+R++PP F GG DP+ A+ W++EIEKIF V++ + K+ AT++L +A
Sbjct: 167 GLDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>BM142940
Length = 422
Score = 43.9 bits (102), Expect(2) = 3e-08
Identities = 17/42 (40%), Positives = 29/42 (68%)
Frame = -3
Query: 720 ANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPK 761
AN KC F ++ ++L H+IS +G++ DP K+E +++W PK
Sbjct: 156 ANRKKCSFGCKKFEYLVHMISVKGVSADPKKLEVMVAWPIPK 31
Score = 33.9 bits (76), Expect(2) = 3e-08
Identities = 17/34 (50%), Positives = 22/34 (64%)
Frame = -1
Query: 685 VVVFIDDILIYSKNVVEHEGHLRQVLQVLREKEL 718
+ F DILIYS N +HE HL+ VLQ +E +L
Sbjct: 260 ICFFSYDILIYSINEGKHEKHLKIVLQTFKEAKL 159
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 47.4 bits (111), Expect = 5e-05
Identities = 24/51 (47%), Positives = 30/51 (58%)
Frame = -1
Query: 662 VTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQV 712
+TNAP F MN F L ++V+VF DDIL+YS EH HL V +V
Sbjct: 153 LTNAPTSFQCLMNHIFQHALRKYVLVFFDDILVYSSTWHEHLCHLEVVFKV 1
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 45.8 bits (107), Expect = 1e-04
Identities = 30/95 (31%), Positives = 41/95 (42%), Gaps = 19/95 (20%)
Frame = +2
Query: 246 PYQRPAGRGFAS----------GSYR---PMV------GAAGGSGDQTQNRELTCFKCGK 286
PY+R + RGF+ G Y P V G G + + L C+ C +
Sbjct: 299 PYRRDSRRGFSRDNLCKNCKRPGHYARECPNVAICHNCGLPGHIASECTTKSL-CWNCKE 475
Query: 287 PGHFARACPDTRPQCYNCNKLGHTAAQCRVPKAEP 321
PGH A +CP+ C+ C K GH A +C P P
Sbjct: 476 PGHMASSCPN-EGICHTCGKAGHRARECSAPPMPP 577
Score = 42.4 bits (98), Expect = 0.002
Identities = 22/42 (52%), Positives = 23/42 (54%)
Frame = +2
Query: 278 ELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVPKA 319
E C C K GH AR CP+ P C CN GH A QC PKA
Sbjct: 641 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQC--PKA 757
Score = 41.2 bits (95), Expect = 0.003
Identities = 21/45 (46%), Positives = 22/45 (48%), Gaps = 6/45 (13%)
Frame = +2
Query: 281 CFKCGKPGHFARAC------PDTRPQCYNCNKLGHTAAQCRVPKA 319
C CGK GH AR C P C NC K GH AA+C KA
Sbjct: 515 CHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEKA 649
Score = 40.0 bits (92), Expect = 0.008
Identities = 19/50 (38%), Positives = 25/50 (50%)
Frame = +2
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQC 314
GA GG G R++ C C + GH +R C C+NC GH A +C
Sbjct: 800 GARGGGGGGY--RDVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 943
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 45.4 bits (106), Expect = 2e-04
Identities = 21/41 (51%), Positives = 27/41 (65%)
Frame = -3
Query: 576 VLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQL 616
V++VKK +GK R+C DY LN K+ YPLP ID + D L
Sbjct: 124 VVMVKKPNGKWRICTDYIDLN*ACPKDAYPLPNIDHMTDGL 2
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 44.3 bits (103), Expect = 4e-04
Identities = 33/107 (30%), Positives = 38/107 (34%), Gaps = 13/107 (12%)
Frame = +3
Query: 221 RAGTGGPMRSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELT 280
R G GG R G RG + G G+ G Y G GG G
Sbjct: 102 RGGYGGGGRGG------RGGYGG-------GGGGYGGGGYGGGGGYGGGGGGGGGG---- 230
Query: 281 CFKCGKPGHFARACPD-------------TRPQCYNCNKLGHTAAQC 314
C+KCG+ GH AR C CYNC + GH A C
Sbjct: 231 CYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGESGHFARDC 371
Score = 34.3 bits (77), Expect = 0.42
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +3
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARACPDT 297
G GG G +C+ CG+ GHFAR CP +
Sbjct: 306 GGGGGGG--------SCYNCGESGHFARDCPSS 380
>TC233976
Length = 763
Score = 43.1 bits (100), Expect = 9e-04
Identities = 30/123 (24%), Positives = 47/123 (37%), Gaps = 8/123 (6%)
Frame = +1
Query: 206 EKATEVELMKNRRLNRAGTGGPM-RSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYRPMV 264
++ +V KN + PM S ++ + +++N QR +G +G +V
Sbjct: 22 DQQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLV 201
Query: 265 -------GAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAAQCRVP 317
G G L C KCG+ GH A C C+N GH + C P
Sbjct: 202 NRVSQPAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNCPHP 381
Query: 318 KAE 320
+ E
Sbjct: 382 RKE 390
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 43.1 bits (100), Expect = 9e-04
Identities = 21/52 (40%), Positives = 27/52 (51%), Gaps = 2/52 (3%)
Frame = +1
Query: 281 CFKCGKPGHFARACP--DTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKR 330
CF CG GH+AR C D + +CY C + GH C K P + RG+R
Sbjct: 364 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNC---KNSPKKLSQRGRR 510
>TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (89%)
Length = 936
Score = 42.7 bits (99), Expect = 0.001
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +2
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
G GG G ++ +L C++CG+PGHFAR C
Sbjct: 290 GGGGGRGGRSGGSDLKCYECGEPGHFAREC 379
>TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (72%)
Length = 1003
Score = 42.7 bits (99), Expect = 0.001
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +3
Query: 265 GAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
G GG G ++ +L C++CG+PGHFAR C
Sbjct: 786 GGGGGRGGRSGGSDLKCYECGEPGHFAREC 875
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.345 0.152 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,429,309
Number of Sequences: 63676
Number of extensions: 991710
Number of successful extensions: 8571
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 6197
Number of HSP's successfully gapped in prelim test: 253
Number of HSP's that attempted gapping in prelim test: 2085
Number of HSP's gapped (non-prelim): 6791
length of query: 1469
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1360
effective length of database: 5,698,948
effective search space: 7750569280
effective search space used: 7750569280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0016.15


