

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.18
(181 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
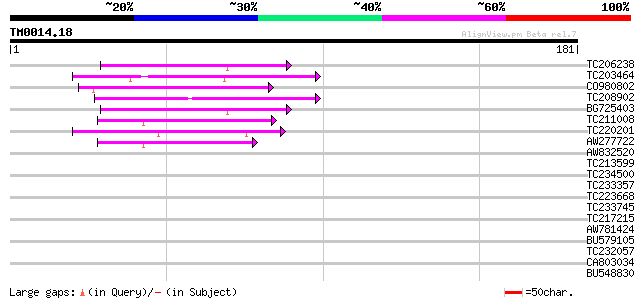
Score E
Sequences producing significant alignments: (bits) Value
TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragmen... 50 8e-07
TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leish... 48 3e-06
CO980802 47 5e-06
TC208902 46 1e-05
BG725403 45 2e-05
TC211008 44 5e-05
TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance ... 41 4e-04
AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa pr... 40 6e-04
AW832520 38 0.003
TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transpor... 35 0.015
TC234500 32 0.13
TC233357 homologue to UP|Q8VWW2 (Q8VWW2) Two-component response ... 29 1.1
TC223668 29 1.1
TC233745 28 1.8
TC217215 weakly similar to UP|Q84ZX0 (Q84ZX0) HEN4, partial (31%) 28 1.8
AW781424 28 2.4
BU579105 similar to GP|12328468|dbj P0416D03.16 {Oryza sativa (j... 28 2.4
TC232057 similar to UP|Q8L5C7 (Q8L5C7) UDP-glucuronosyltransfera... 28 3.1
CA803034 27 4.0
BU548830 27 4.0
>TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragment), partial
(9%)
Length = 1259
Score = 49.7 bits (117), Expect = 8e-07
Identities = 25/62 (40%), Positives = 37/62 (59%), Gaps = 1/62 (1%)
Frame = +3
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS-YSTLQAELWGILCGARLLQE 88
++LN GAY S+D AAC GI RD FV+ ++ +LG S + E+WGI G ++ +
Sbjct: 624 LRLNVSGAYDRSSDTAACGGIFRDNNDRFVLGFSVKLGECLSNDEGEIWGIYHGMKIARR 803
Query: 89 RD 90
D
Sbjct: 804 YD 809
>TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leishmania
major;} , partial (8%)
Length = 1386
Score = 47.8 bits (112), Expect = 3e-06
Identities = 28/83 (33%), Positives = 50/83 (59%), Gaps = 4/83 (4%)
Frame = +2
Query: 21 RWWPLDAVWVKLNSDGA---YISSADMAACRGIIRDRFSAFVIAYARRLG-SYSTLQAEL 76
RW + WVKLN DG+ Y SS+ A C G++RD + ++ +A++L +Y+ Q EL
Sbjct: 485 RWKKPEIGWVKLNVDGSRDPYKSSS--AGCGGVLRDASAKWLRGFAKKLNPTYAVHQTEL 658
Query: 77 WGILCGARLLQERDY*RVLIETN 99
IL G ++ E + ++++E++
Sbjct: 659 EAILTGLKVASEMNVKKLIVESD 727
>CO980802
Length = 803
Score = 47.0 bits (110), Expect = 5e-06
Identities = 23/64 (35%), Positives = 37/64 (56%), Gaps = 2/64 (3%)
Frame = -3
Query: 23 WPL--DAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGIL 80
WPL ++ K+N DG+Y +S ACRGII D F + + +G +++ A+LWG+
Sbjct: 429 WPLLGPLLFFKVNVDGSYKASTAKMACRGIILDSFGRVLSCFHSNIGVCNSIWAKLWGLK 250
Query: 81 CGAR 84
G +
Sbjct: 249 HGIK 238
>TC208902
Length = 838
Score = 45.8 bits (107), Expect = 1e-05
Identities = 23/72 (31%), Positives = 42/72 (57%)
Frame = +1
Query: 28 VWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQ 87
VW ++NS+ S +AAC ++R++ A V+ +A LG S QAE+W I G +L
Sbjct: 220 VWGRINSNCDGAVSGGVAACGRVLRNQAGA-VVVFAHNLGMCSISQAEIWAITMGVKLAW 396
Query: 88 ERDY*RVLIETN 99
++ + + +E++
Sbjct: 397 DKGFTNLYVESD 432
>BG725403
Length = 404
Score = 44.7 bits (104), Expect = 2e-05
Identities = 23/62 (37%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Frame = +3
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS-YSTLQAELWGILCGARLLQE 88
++LN GAY S+ A C GI D+ FV+ + LG +S +AE+WGI G ++ ++
Sbjct: 12 LRLNVSGAYDPSSGTATCGGIFSDKHRRFVLGFWVELGECWSIDEAEIWGIYHGMKIARK 191
Query: 89 RD 90
D
Sbjct: 192YD 197
>TC211008
Length = 516
Score = 43.5 bits (101), Expect = 5e-05
Identities = 21/61 (34%), Positives = 34/61 (55%), Gaps = 4/61 (6%)
Frame = +1
Query: 29 WVKLNSDGAYISS----ADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGAR 84
WV LN+DG+ + +AC G++RD ++ + LG+ S AELWG++ G +
Sbjct: 85 WVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELWGVVHGLK 264
Query: 85 L 85
L
Sbjct: 265 L 267
>TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance receptor
PtdSerR (Phosphatidylserine receptor), partial (5%)
Length = 1185
Score = 40.8 bits (94), Expect = 4e-04
Identities = 24/72 (33%), Positives = 39/72 (53%), Gaps = 4/72 (5%)
Frame = +1
Query: 21 RWWPLDAVWVKLNSDGAYISSADMAA-CRGIIRDRFSAFVIAYARRLGSYSTLQA---EL 76
+W ++ WVKLN DG+ I +A C G+IRD + + + + ++L QA EL
Sbjct: 334 KWKKPESGWVKLNVDGSRIHEEPASAGCGGVIRDEWGTWCVGFDQKLDPNICRQAHYTEL 513
Query: 77 WGILCGARLLQE 88
IL G ++ +E
Sbjct: 514 QAILTGLKVARE 549
>AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa protein 3
(Hsc70.3). [Mouse-ear cress] {Arabidopsis thaliana},
partial (8%)
Length = 432
Score = 40.0 bits (92), Expect = 6e-04
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 4/55 (7%)
Frame = +1
Query: 29 WVKLNSDGAYISS----ADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
WV LN+DG+ + +AC G++RD ++ + LG+ S AELWG+
Sbjct: 268 WVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELWGV 432
>AW832520
Length = 423
Score = 37.7 bits (86), Expect = 0.003
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Frame = -3
Query: 7 VTRVVSPVRDAP-AIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARR 65
+ R++ + P IRW P KLN DGA ++ + G++RD FVI +
Sbjct: 298 ILRLMEEITSYPNGIRWSPPPEGLFKLNYDGAVNCNSLKVSAGGVVRDSHGNFVIGFTTT 119
Query: 66 LGSYSTLQAELW 77
S + AELW
Sbjct: 118 PDSCTINIAELW 83
>TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transport exbB
protein, partial (8%)
Length = 825
Score = 35.4 bits (80), Expect = 0.015
Identities = 23/68 (33%), Positives = 35/68 (50%)
Frame = +3
Query: 40 SSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQERDY*RVLIETN 99
S DMAAC G++RD + ++ +LG+ S AEL G G L ++ +E N
Sbjct: 336 SDNDMAACGGLLRDCHGRWAASFV*KLGNTSAFVAEL*GAYLGLSLAWDKG--DKNLEVN 509
Query: 100 LSILGVVL 107
+ L VV+
Sbjct: 510 IDSLVVVV 533
>TC234500
Length = 866
Score = 32.3 bits (72), Expect = 0.13
Identities = 20/81 (24%), Positives = 37/81 (44%)
Frame = +1
Query: 29 WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQE 88
W+K N DG+Y + + + + RD ++ +A LG+ S CG LL+
Sbjct: 139 WIKANVDGSYKALQHLTSSSAVFRDSTGSWCFGFALNLGNLS----------CG*PLLKL 288
Query: 89 RDY*RVLIETNLSILGVVLVL 109
++ E ++ I V L++
Sbjct: 289 SEF-----EDSVRIFSVTLII 336
>TC233357 homologue to UP|Q8VWW2 (Q8VWW2) Two-component response
regulator-like protein, partial (8%)
Length = 811
Score = 29.3 bits (64), Expect = 1.1
Identities = 15/45 (33%), Positives = 23/45 (50%)
Frame = +2
Query: 47 CRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQERDY 91
CRG F + + A+A +G +S L A+LW G + RD+
Sbjct: 353 CRGF-EQSFYSDLFAFADGIGCFSALNAKLWAXXXGICVAWARDF 484
>TC223668
Length = 866
Score = 29.3 bits (64), Expect = 1.1
Identities = 17/55 (30%), Positives = 27/55 (48%)
Frame = -3
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGAR 84
VKLN DG+ + G+IRD ++ +AR G + L +EL+ + R
Sbjct: 555 VKLNVDGSGHYEPHVMEVGGLIRDGVGRWLFRFARHCGLGNPLLSELFALEASLR 391
>TC233745
Length = 423
Score = 28.5 bits (62), Expect = 1.8
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +1
Query: 62 YARRLGSYSTLQAELWGILCGARL 85
+A +LGS S L A LW IL G R+
Sbjct: 70 FAYKLGSCSVLNASLWVILIGVRV 141
>TC217215 weakly similar to UP|Q84ZX0 (Q84ZX0) HEN4, partial (31%)
Length = 1798
Score = 28.5 bits (62), Expect = 1.8
Identities = 20/48 (41%), Positives = 25/48 (51%)
Frame = -2
Query: 83 ARLLQERDY*RVLIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRL 130
ARLL Y + + +I VVLV + VLL LT F V+PRL
Sbjct: 1266 ARLLPFSPYTEPITSSGTAICTVVLVTMAVLLPLTSSKPPF*EVNPRL 1123
>AW781424
Length = 427
Score = 28.1 bits (61), Expect = 2.4
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = +1
Query: 18 PAIRWWPLDAVWVKLNS 34
P WW LD++W++LN+
Sbjct: 349 PPFNWWILDSLWMELNN 399
>BU579105 similar to GP|12328468|dbj P0416D03.16 {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 425
Score = 28.1 bits (61), Expect = 2.4
Identities = 16/35 (45%), Positives = 21/35 (59%), Gaps = 3/35 (8%)
Frame = +3
Query: 21 RWWPLDAVWVKLNSDGA---YISSADMAACRGIIR 52
RW + WVKLN DG+ Y SS+ A C G++R
Sbjct: 327 RWKKPEIGWVKLNVDGSRDPYKSSS--AGCGGVLR 425
>TC232057 similar to UP|Q8L5C7 (Q8L5C7) UDP-glucuronosyltransferase , partial
(34%)
Length = 1228
Score = 27.7 bits (60), Expect = 3.1
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = +3
Query: 18 PAIRWWPLDAVWVKLNS 34
P WW LD++W++LN+
Sbjct: 852 PHFNWWILDSLWMELNN 902
>CA803034
Length = 456
Score = 27.3 bits (59), Expect = 4.0
Identities = 13/31 (41%), Positives = 17/31 (53%), Gaps = 3/31 (9%)
Frame = +2
Query: 29 WVKLNSDGAYISSADMAACRGII---RDRFS 56
W+K N+DGA S +A C G+ R FS
Sbjct: 152 WIKCNTDGAAKGSPGLAGCCGLFIQ*RSEFS 244
>BU548830
Length = 712
Score = 27.3 bits (59), Expect = 4.0
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +1
Query: 68 SYSTLQAELWGILCG 82
+Y T Q +LWGI+CG
Sbjct: 448 NYKTRQIQLWGIICG 492
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.336 0.145 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,044,544
Number of Sequences: 63676
Number of extensions: 102551
Number of successful extensions: 827
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 817
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 824
length of query: 181
length of database: 12,639,632
effective HSP length: 91
effective length of query: 90
effective length of database: 6,845,116
effective search space: 616060440
effective search space used: 616060440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0014.18


