

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
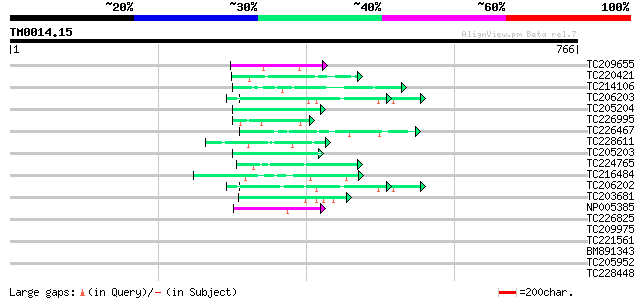
Query= TM0014.15
(766 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC209655 similar to UP|Q8L844 (Q8L844) Crp1 protein-like, partia... 49 6e-06
TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell... 47 2e-05
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 47 3e-05
TC206203 UP|Q99366 (Q99366) DNA-directed RNA polymerase (Fragme... 47 3e-05
TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein pr... 46 5e-05
TC226995 similar to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 dehyd... 46 7e-05
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 46 7e-05
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 45 9e-05
TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall pr... 45 9e-05
TC224765 similar to UP|Q09082 (Q09082) Extensin (Class I), parti... 45 1e-04
TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 44 2e-04
TC206202 UP|Q99367 (Q99367) DNA-directed RNA polymerase (Fragme... 44 2e-04
TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 44 3e-04
NP005385 DNA-directed RNA polymerase 43 6e-04
TC226825 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydra... 41 0.002
TC209975 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (9%) 41 0.002
TC221561 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate d... 41 0.002
BM891343 41 0.002
TC205952 41 0.002
TC228448 weakly similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (4%) 41 0.002
>TC209655 similar to UP|Q8L844 (Q8L844) Crp1 protein-like, partial (32%)
Length = 832
Score = 49.3 bits (116), Expect = 6e-06
Identities = 41/137 (29%), Positives = 57/137 (40%), Gaps = 6/137 (4%)
Frame = +1
Query: 299 QHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNL---LKNPTSRSEDPTSL 355
Q SS A TTSPP SP SP +P PT P L P+S S PT+
Sbjct: 13 QSSSKKAPTTSSPSTTAATTSPPSSPSSPTHRPLPTRTPTQPLPPHSAQPSSNSPSPTAP 192
Query: 356 LTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSV---PRASARPAARTTDTDSSTAFTPV 412
+ SS P P P S +++ P +S+ P++ T ++ P
Sbjct: 193 SRLHSGTRSSSPSAPPPPPSPSRTPSFHGSRNTISASPTSSSTPSSSTPSAAPKSSTKP- 369
Query: 413 SFPINVIDSPPSNTSES 429
S+ +V S PS T+ S
Sbjct: 370 SYSPSVRSSLPSPTTRS 420
>TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell wall
protein gp1 precursor (Hydroxyproline-rich glycoprotein
1), partial (13%)
Length = 1409
Score = 47.4 bits (111), Expect = 2e-05
Identities = 51/182 (28%), Positives = 67/182 (36%), Gaps = 5/182 (2%)
Frame = +3
Query: 300 HSSNPLTPIPESQPAAQTTSPPH---SPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLL 356
H +PL + QP +TSPPH +P P P P + +P S PT+
Sbjct: 156 HHPHPLWTQNKQQPLNPSTSPPHRTPAPSPPSTMPPSATPPSPSATSSP---SASPTAPP 326
Query: 357 TIPYDPVSSEPIIHEQPEPNQTEPQ-PRTSDHSVPRASARPAARTTDTDSSTAFTPVSFP 415
T P S P P P+ T P P S + P S A T +S + +P S
Sbjct: 327 TSPSPSPPSSPSPPSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASANSPASGSPTS-- 500
Query: 416 INVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPD-EPILA 474
+ PS TS S K A PS + P P P + PD P +
Sbjct: 501 -RTSPTSPSPTSTS----------KPPAPSSSSPA*PSSKPSPSPTP---ISPDPSPATS 638
Query: 475 NP 476
P
Sbjct: 639 TP 644
Score = 36.2 bits (82), Expect = 0.056
Identities = 38/153 (24%), Positives = 54/153 (34%), Gaps = 4/153 (2%)
Frame = +3
Query: 237 KKKQKKVRIVVKPTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISA 296
+ KQ+ + P PA + A SA SS A + + + P S
Sbjct: 180 QNKQQPLNPSTSPPHRTPAPSPPSTMPPSATPPSPSATSSPSASPTAPPTSPSPSPPSSP 359
Query: 297 LLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLL 356
+ +P + P A+ TSP SP SP S + + S PTS
Sbjct: 360 SPPSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASANSPASGSPTSRTSPTSPSPTSTS 539
Query: 357 TIPYDPVSSEPIIHEQPEPNQT----EPQPRTS 385
P SS +P P+ T +P P TS
Sbjct: 540 KPPAPSSSSPA*PSSKPSPSPTPISPDPSPATS 638
Score = 32.3 bits (72), Expect = 0.80
Identities = 32/125 (25%), Positives = 49/125 (38%), Gaps = 5/125 (4%)
Frame = +3
Query: 249 PTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLTPI 308
P+ P +A + AP+ + S S P+ S A SA S +P +
Sbjct: 348 PSSPSPPSAPSPSSTAPSPPSASPPTSPSPSPPSP-------ASTASANSPASGSPTSRT 506
Query: 309 PESQPAAQTTSPPHSPRS-----PFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
+ P+ +TS P +P S P +PSP+ P+ S DP+ + P
Sbjct: 507 SPTSPSPTSTSKPPAPSSSSPA*PSSKPSPSPTPI----------SPDPSPATSTPTSHT 656
Query: 364 SSEPI 368
S PI
Sbjct: 657 SISPI 671
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 47.0 bits (110), Expect = 3e-05
Identities = 63/240 (26%), Positives = 76/240 (31%), Gaps = 5/240 (2%)
Frame = +1
Query: 301 SSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPY 360
S P +P P S P + PP P P PSPT +++P S P S P
Sbjct: 1 SPPPPSPPPPSPPPPVYSPPPPPPSPP--PPSPTYC-----VRSPPPPSPPPPSPPPPP- 156
Query: 361 DPVSSEP----IIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPI 416
PV S P + P P Q P P HS P P S PV P
Sbjct: 157 PPVFSPPPPVQYYYSSPPPPQHSPPP-PPPHSPPPPPPSPPPPVYPYLSPPPPPPVHSPP 333
Query: 417 NVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANP 476
+ SPP P P P P P ++P P + P
Sbjct: 334 PPVYSPPPPPPS-----------------------PPPCIEPPPPPPPCIEPPPPPPSPP 444
Query: 477 -IQEADPLVQQAHPVPDQQEPIQPDPEPEQSVSNQSSVRSPHPLVETSDHHRGTSEPNAP 535
+E P HP P P P P P N SP P H+ P+ P
Sbjct: 445 PCEEHSPPPPSPHPAP-YHPPPSPSPPPPPVQYNSPPPPSPPPPTPVY-HYNSPPPPSFP 618
>TC206203 UP|Q99366 (Q99366) DNA-directed RNA polymerase (Fragment) , partial
(96%)
Length = 2604
Score = 47.0 bits (110), Expect = 3e-05
Identities = 67/262 (25%), Positives = 91/262 (34%), Gaps = 10/262 (3%)
Frame = +1
Query: 311 SQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPII 369
S P TSP +SP SP + P SPT +P + +PTS + PTS P P S
Sbjct: 1138 SSPGYSPTSPGYSPTSPGYSPTSPTYSPS-SPGYSPTSPAYSPTSPSYSPTSPSYSPTSP 1314
Query: 370 HEQP-EPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSE 428
P P+ + P S S + PA T S A++P S + S TS
Sbjct: 1315 SYSPTSPSYSPTSPSYSPTSPAYSPTSPAYSPT----SPAYSPTSPSYSPTSPSYSPTSP 1482
Query: 429 SIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQAH 488
S Y T PS Y P P P + P
Sbjct: 1483 S-----------------YSPTSPS---YSPTSPS--YSPTSPAYSPTSPGYSPTSPSYS 1596
Query: 489 PVPDQQEPIQPDPEPEQSVSNQSSVRSP--------HPLVETSDHHRGTSEPNAPMINIG 540
P P P P+ + + S SP P TS ++ TS +P
Sbjct: 1597 PTSPSYSPTSPSYNPQSAKYSPSLAYSPSSPRLSPSSPYSPTSPNYSPTSPSYSPTSPSY 1776
Query: 541 SPQGASEAHSSNHQASPEPNLS 562
SP + + SS + + P+ S
Sbjct: 1777 SPSSPTYSPSSPYNSGVSPDYS 1842
Score = 43.9 bits (102), Expect = 3e-04
Identities = 60/236 (25%), Positives = 85/236 (35%), Gaps = 12/236 (5%)
Frame = +1
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSED 351
P S SS +P + PA TSP +SP SP + P SP+ +P + +PTS S
Sbjct: 1198 PTSPTYSPSSPGYSP---TSPAYSPTSPSYSPTSPSYSPTSPSYSPT-SPSYSPTSPSYS 1365
Query: 352 PTSLLTIPYDPVSSEPIIHEQP-EPNQTEPQPRTSDHSVPRASARPAARTTD---TDSST 407
PTS P P S P P+ + P S S + P+ T + +S
Sbjct: 1366 PTSPAYSPTSPAYSPTSPAYSPTSPSYSPTSPSYSPTSPSYSPTSPSYSPTSPSYSPTSP 1545
Query: 408 AFTPVS---FPINVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPER 464
A++P S P + SP S + + K S Y PS R P
Sbjct: 1546 AYSPTSPGYSPTSPSYSPTSPSYSPTSPSYNPQSAKYSPSLAY---SPSSPRLSPSSPYS 1716
Query: 465 LVDPDEPILANPIQEADPLVQQAHPVPDQQEP----IQPDPEPEQSVSNQSSVRSP 516
P+ + P + P P + PD P + S+ SP
Sbjct: 1717 PTSPNYSPTSPSYSPTSPSYSPSSPTYSPSSPYNSGVSPDYSPSSPQFSPSTGYSP 1884
Score = 32.7 bits (73), Expect = 0.61
Identities = 25/92 (27%), Positives = 38/92 (41%), Gaps = 1/92 (1%)
Frame = +1
Query: 303 NPLTPIPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYD 361
+P +P + P TSP +SP SP + P SPT +P +P + P Y
Sbjct: 1696 SPSSPYSPTSPNYSPTSPSYSPTSPSYSPSSPTYSP-----SSPYNSGVSP------DYS 1842
Query: 362 PVSSEPIIHEQPEPNQTEPQPRTSDHSVPRAS 393
P S + P+Q P ++ P+ S
Sbjct: 1843 PSSPQFSPSTGYSPSQPGYSPSSTSQYTPQTS 1938
>TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (44%)
Length = 996
Score = 46.2 bits (108), Expect = 5e-05
Identities = 33/128 (25%), Positives = 47/128 (35%), Gaps = 2/128 (1%)
Frame = +1
Query: 301 SSNPLTPIPESQPAAQTTSPPHSPRS--PFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTI 358
S++P P P+S P +SPP P S P P P P L +P S P+S
Sbjct: 187 STSPAPPTPQSSPPPIQSSPPPVPSSPPPAQSPPPASTPPPAPLSSPPPASPPPSSPPPA 366
Query: 359 PYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINV 418
P S P P P P + + P ++ P A P +
Sbjct: 367 SPPPASPPPASPPPASPPPASPPPASPPPATPPPASPPPFSPPPATPPPATPPPALTPTP 546
Query: 419 IDSPPSNT 426
+ SPP+ +
Sbjct: 547 LSSPPATS 570
>TC226995 similar to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 dehydratase
(GDP-D-mannose dehydratase) (GMD) , partial (59%)
Length = 679
Score = 45.8 bits (107), Expect = 7e-05
Identities = 42/133 (31%), Positives = 53/133 (39%), Gaps = 22/133 (16%)
Frame = +3
Query: 301 SSNPLTPIPE--------SQPAAQTTSPPHSPRSPFFQPSPTEAPL--------WNLLKN 344
S+ P++P P S P TTSPP SP SP SPT P ++
Sbjct: 213 STTPISPTPPPSAAGSTPSSPTRSTTSPP-SPTSPTHSRSPTTPPTSSPPAPSAYSRPIA 389
Query: 345 PTS-RSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPR-----ASARPAA 398
PTS + PTS T P P S P + +P P P T + P +ARP
Sbjct: 390 PTSPTTAAPTSATTKPAPPRCSAPPLRRNRKPLACTPAPHTPAANAPTTGTP*TTARPTP 569
Query: 399 RTTDTDSSTAFTP 411
+ T SS P
Sbjct: 570 SSPATASSLTNDP 608
Score = 28.9 bits (63), Expect = 8.9
Identities = 30/103 (29%), Positives = 41/103 (39%), Gaps = 8/103 (7%)
Frame = +1
Query: 302 SNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPL----WNLLKNPTSRSEDPTSLL- 356
+ P P+ E Q A T PH SP P EAPL LL P +R P L
Sbjct: 118 ARPDPPLVELQHA---THRPHIRGSPQRPQGPHEAPLRRSLRRLLPPPLARHHPPRRGLQ 288
Query: 357 ---TIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARP 396
+P + +P +H + ++ P T S+P RP
Sbjct: 289 PRRPVPRRRLIRDPRLHRRRRRHRRPPP--TRGRSLPHLRLRP 411
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 45.8 bits (107), Expect = 7e-05
Identities = 61/259 (23%), Positives = 93/259 (35%), Gaps = 15/259 (5%)
Frame = +3
Query: 311 SQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEP--I 368
S A T+P +SP + SP+ + PT + PT+ P +S+P I
Sbjct: 39 SNAQAPATAPANSPPTTPAATSPSSGTPLAVTLPPTVVASPPTTSTP----PTTSQPPAI 206
Query: 369 IHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSE 428
+ +P P T P P+ + S P+A + S TP S P I PP +T
Sbjct: 207 VAPKPAP-VTPPAPKVAPASSPKAPPPQTPQPQPPKVSPVVTPTSPP--PIPPPPVSTPP 377
Query: 429 SIRKFMEVRKEKVSALEEYYLTCPSPRRY---PGPRPERLVDPDEPILANPIQEADPLVQ 485
+ K++ P+P + P P P + P+ P P+V+
Sbjct: 378 PLPP------PKIAPTPAKTPPAPAPAKATPAPAPAPTKPAPTPAPVPPPPTLAPTPVVE 539
Query: 486 QAHPVPDQQEPIQ----------PDPEPEQSVSNQSSVRSPHPLVETSDHHRGTSEPNAP 535
P P + + P P P + VR P S T+ +P
Sbjct: 540 VPAPAPSSHKHRRHRHRHRRHQAPAPAP-------TIVRKSPPAPPDSTAESDTTPAPSP 698
Query: 536 MINIGSPQGASEAHSSNHQ 554
+N+ A SSNHQ
Sbjct: 699 SLNL-------NAASSNHQ 734
Score = 29.6 bits (65), Expect = 5.2
Identities = 41/205 (20%), Positives = 59/205 (28%), Gaps = 24/205 (11%)
Frame = +3
Query: 249 PTRVEPAAAVVRRTEAPARVTRSSAHSSKPALASDDDLNLFD----ALPISALLQHSSNP 304
P P++ P V S +S P S + P + + +S+P
Sbjct: 90 PAATSPSSGTPLAVTLPPTVVASPPTTSTPPTTSQPPAIVAPKPAPVTPPAPKVAPASSP 269
Query: 305 LTPIPES--------QPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLL 356
P P++ P TSPP P P P P P + P P
Sbjct: 270 KAPPPQTPQPQPPKVSPVVTPTSPPPIPPPPVSTPPPLPPP--KIAPTPAKTPPAPAPAK 443
Query: 357 TIPYD------------PVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTD 404
P PV P + P P P + H R R
Sbjct: 444 ATPAPAPAPTKPAPTPAPVPPPPTLAPTPVVEVPAPAPSSHKHRRHRHRHRRHQAPAPAP 623
Query: 405 SSTAFTPVSFPINVIDSPPSNTSES 429
+ +P +PP +T+ES
Sbjct: 624 TIVRKSP--------PAPPDSTAES 674
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 45.4 bits (106), Expect = 9e-05
Identities = 50/189 (26%), Positives = 73/189 (38%), Gaps = 20/189 (10%)
Frame = +3
Query: 265 PARVTRSSAHSSKPALASDDDLNLFDALPISALLQHSSNPLTPIPESQPAAQTTSPP--- 321
P++ +RS A P ++ A +A+ SS+P P P +TSP
Sbjct: 147 PSQASRSPAPPPPPPRSASSSTT---ASSPTAVTPPSSSPPRPSPGPSATPTSTSPSPAT 317
Query: 322 -----HSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPIIHEQPEPN 376
+PR P P+PT +P P + S P S P P SS E P P
Sbjct: 318 PSTSSSAPRPPLSSPAPTPSPAPTFSPPPPATS-SPCS--AAPSSPSSSAAATAEPPSPP 488
Query: 377 QTEP-----------QPRTSDHSV-PRASARPAARTTDTDSSTAFTPVSFPINVIDSPPS 424
P P +S ++ P ++ P+ T +DS TA P S + PP
Sbjct: 489 PYPPTSPPPPCPSLKSPNSSPNAASPSKNSSPSVGPTPSDSLTA--PSSSRTSPTTLPPP 662
Query: 425 NTSESIRKF 433
T ++R F
Sbjct: 663 TTLATLRGF 689
>TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall protein,
partial (48%)
Length = 1076
Score = 45.4 bits (106), Expect = 9e-05
Identities = 34/125 (27%), Positives = 48/125 (38%), Gaps = 2/125 (1%)
Frame = +3
Query: 301 SSNPLTPIPESQPAAQTTSPPHSPRS--PFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTI 358
S++P P P++ P +SPP P S P P P P + L +P S P+S
Sbjct: 279 STSPAPPTPQASPPPVQSSPPPVPSSPPPSQSPPPASTPPPSPLSSPPPSSPPPSSPPPS 458
Query: 359 PYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINV 418
P S P P + P P + P + P A + TP P++
Sbjct: 459 SPPPASPPPSSPPPASPPPSSPPPSSPPPFSPPPATPPPATPPPAVPPPSLTPTVTPLS- 635
Query: 419 IDSPP 423
SPP
Sbjct: 636 --SPP 644
>TC224765 similar to UP|Q09082 (Q09082) Extensin (Class I), partial (46%)
Length = 708
Score = 45.1 bits (105), Expect = 1e-04
Identities = 46/175 (26%), Positives = 65/175 (36%), Gaps = 5/175 (2%)
Frame = +3
Query: 307 PIPESQPAAQTTSPPHSPRSP---FFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
P P P + PP SP P +++ P +P++ + K+P S PT + P PV
Sbjct: 15 PPPSPTPYYYHSPPPPSPSPPSPYYYKSPPPPSPVY-IYKSPPPPS--PTYVYKSPPPPV 185
Query: 364 SSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDS-- 421
S P ++ P P +P+P S P S +P S +PV P S
Sbjct: 186 KSPPYYYQSPPPPSPKPKPPYYYKSPPPPSPKPKP-PYYYQSPPPPSPVPKPPYYYKSPP 362
Query: 422 PPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANP 476
PPS + + S YY P P P P P P + P
Sbjct: 363 PPSPVPKPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPP 527
Score = 32.7 bits (73), Expect = 0.61
Identities = 27/99 (27%), Positives = 37/99 (37%), Gaps = 11/99 (11%)
Frame = +3
Query: 303 NPLTPIPESQPAAQTTSPPHS---PRSPFFQPSPT------EAPLWNLLKNPTSRSEDPT 353
+P P P+ +P SPP P+ P++ SP + P + P S S P
Sbjct: 258 SPPPPSPKPKPPYYYQSPPPPSPVPKPPYYYKSPPPPSPVPKPPYYYHSPPPPSPSPPPP 437
Query: 354 SLLTIPYDPVSSEP--IIHEQPEPNQTEPQPRTSDHSVP 390
P P S P ++ P P P P HS P
Sbjct: 438 YYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYHSPP 554
>TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (58%)
Length = 1213
Score = 44.3 bits (103), Expect = 2e-04
Identities = 57/245 (23%), Positives = 90/245 (36%), Gaps = 15/245 (6%)
Frame = +1
Query: 249 PTRVEPAAAVVRRTEAPARVTRSSAHSSKP-ALASDDDLNLFDALPISALLQHSSNPLTP 307
PTR A+ +P T S + +S P + A + P S+ P TP
Sbjct: 103 PTRTTSPASSQSTPSSPPSTTTSPSPTSPPKSTAKPPSPSAPSTTPPCRTFSRSTPPSTP 282
Query: 308 IPESQPAAQ---TTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVS 364
S P+ T++P S RSP PSP + P + DP + T P S
Sbjct: 283 SRTSSPSTSSSTTSAPRSSTRSPTAPPSPPQC------TKPPAPPPDPPASST---SPTS 435
Query: 365 SEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDS---STAFTPVSFPINVIDS 421
+ H P+ T+ HS P+ S+ P ++ T S F+P+ +
Sbjct: 436 TAEKSHSPPK--------TTTAHS-PQRSSNP*KKSLTTSQ*SRSAKFSPLPQRKPLHPH 588
Query: 422 PPSNTSESIRKFMEVRKEKVSALEEYYLTCPS--------PRRYPGPRPERLVDPDEPIL 473
PP+ TS ++ + + R +L PS P P + P+
Sbjct: 589 PPNKTSPTLCQNTDARSSPTLSLLNPTRLTPSMTTSMGA*PSFAPSTTRSKRFSPNSKT* 768
Query: 474 ANPIQ 478
NP++
Sbjct: 769 PNPVR 783
Score = 32.0 bits (71), Expect = 1.0
Identities = 16/49 (32%), Positives = 24/49 (48%)
Frame = +2
Query: 185 PPAPEFPSPPKKKTKKKMVLEESSEESDVPLVKKTKSKPDDGDDDDSED 233
P A + P+P K KKK + +++SD P PDD D ++D
Sbjct: 1067 PSAADAPAPAKNGKKKKKAADAPADDSDAP-----ADSPDDAADVTADD 1198
>TC206202 UP|Q99367 (Q99367) DNA-directed RNA polymerase (Fragment) ,
complete
Length = 1826
Score = 44.3 bits (103), Expect = 2e-04
Identities = 65/262 (24%), Positives = 89/262 (33%), Gaps = 10/262 (3%)
Frame = +3
Query: 311 SQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEPII 369
S P TSP +SP SP + P SP +P + +P+S PTS P P S
Sbjct: 660 SSPGYSPTSPGYSPTSPGYSPTSPGYSPT-SPTYSPSSPGYSPTSPAYSPTSPSYSPTSP 836
Query: 370 HEQP-EPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSE 428
P P+ + P S S + PA T S A++P S + S TS
Sbjct: 837 SYSPTSPSYSPTSPSYSPTSPSYSPTSPAYSPT----SPAYSPTSPSYSPTSPSYSPTSP 1004
Query: 429 SIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQAH 488
S Y T PS Y P P P + P
Sbjct: 1005S-----------------YSPTSPS---YSPTSPS--YSPTSPAYSPTSPGYSPTSPSYS 1118
Query: 489 PVPDQQEPIQPDPEPEQSVSNQSSVRSP--------HPLVETSDHHRGTSEPNAPMINIG 540
P P P P+ + + S SP P TS ++ TS +P
Sbjct: 1119PTSPSYSPTSPSYNPQSAKYSPSLAYSPSSPRLSPSSPYSPTSPNYSPTSPSYSPTSPSY 1298
Query: 541 SPQGASEAHSSNHQASPEPNLS 562
SP + + SS + + P+ S
Sbjct: 1299SPSSPTYSPSSPYNSGVSPDYS 1364
Score = 43.5 bits (101), Expect = 3e-04
Identities = 60/234 (25%), Positives = 86/234 (36%), Gaps = 10/234 (4%)
Frame = +3
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSED 351
P S SS +P + PA TSP +SP SP + P SP+ +P + +PTS S
Sbjct: 741 PTSPTYSPSSPGYSP---TSPAYSPTSPSYSPTSPSYSPTSPSYSPT-SPSYSPTSPSYS 908
Query: 352 PTSLLTIPYDPVSSEPIIHEQPEPNQTEP--QPRTSDHSVPRASARPAARTTDTDSSTAF 409
PTS P P S P + T P P + +S S P + + + +S A+
Sbjct: 909 PTSPAYSPTSPAYSP----TSPSYSPTSPSYSPTSPSYSPTSPSYSPTS-PSYSPTSPAY 1073
Query: 410 TPVS---FPINVIDSPPSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLV 466
+P S P + SP S + + K S Y PS R P
Sbjct: 1074SPTSPGYSPTSPSYSPTSPSYSPTSPSYNPQSAKYSPSLAY---SPSSPRLSPSSPYSPT 1244
Query: 467 DPDEPILANPIQEADPLVQQAHPVPDQQEP----IQPDPEPEQSVSNQSSVRSP 516
P+ + P + P P + PD P + S+ SP
Sbjct: 1245SPNYSPTSPSYSPTSPSYSPSSPTYSPSSPYNSGVSPDYSPSSPQFSPSTGYSP 1406
Score = 32.7 bits (73), Expect = 0.61
Identities = 25/92 (27%), Positives = 38/92 (41%), Gaps = 1/92 (1%)
Frame = +3
Query: 303 NPLTPIPESQPAAQTTSPPHSPRSPFFQP-SPTEAPLWNLLKNPTSRSEDPTSLLTIPYD 361
+P +P + P TSP +SP SP + P SPT +P +P + P Y
Sbjct: 1218 SPSSPYSPTSPNYSPTSPSYSPTSPSYSPSSPTYSP-----SSPYNSGVSP------DYS 1364
Query: 362 PVSSEPIIHEQPEPNQTEPQPRTSDHSVPRAS 393
P S + P+Q P ++ P+ S
Sbjct: 1365 PSSPQFSPSTGYSPSQPGYSPSSTSQYTPQTS 1460
>TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 913
Score = 43.9 bits (102), Expect = 3e-04
Identities = 45/170 (26%), Positives = 67/170 (38%), Gaps = 17/170 (10%)
Frame = +3
Query: 310 ESQPAAQTTSPPHSPRSPFFQP--SPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPVSSEP 367
+S A+ TTSPP + +PF P +P++ + +PTS S +S P +S P
Sbjct: 75 QSPAASPTTSPPAATTTPFASPATAPSKPKSPAPVASPTSSSPPASSPNAATATPPASSP 254
Query: 368 IIHEQPEPNQTEPQPRTSDHSVPRASARPAA---RTTDTDSSTAFTPVS---FPINVIDS 421
+ P T + P A+ PAA T S A PVS P+ V
Sbjct: 255 TVASPPSKAAAPAPVATPPAATPPAATPPAATPPAVTPVSSPPAPVPVSSPPAPVPVSSP 434
Query: 422 P---PSNTSESIRKFMEV------RKEKVSALEEYYLTCPSPRRYPGPRP 462
P P+ + + EV K+K +++ PSP P P
Sbjct: 435 PALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHTAPAPSPSLLGPPAP 584
>NP005385 DNA-directed RNA polymerase
Length = 2934
Score = 42.7 bits (99), Expect = 6e-04
Identities = 38/131 (29%), Positives = 56/131 (42%), Gaps = 7/131 (5%)
Frame = +1
Query: 303 NPLTPIPES-QPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYD 361
+P +P S PA TSP +SP SP + P+ L + +PTS S PTS P
Sbjct: 2383 SPTSPAYSSTSPAYSPTSPSYSPTSPAYSPTSPSYSLTSPSYSPTSPSYSPTSPSYSPTS 2562
Query: 362 PVSSEPIIHEQP-----EPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSF-P 415
P S P P P+++ +S A + + R + T + + T S+ P
Sbjct: 2563 PAYSPTSPSYSPTSPSYSPTSPSYNPQSAKYSPSLAYSPSSPRLSPTSPNYSPTSPSYSP 2742
Query: 416 INVIDSPPSNT 426
+ SP S T
Sbjct: 2743 TSPSYSPSSPT 2775
Score = 40.0 bits (92), Expect = 0.004
Identities = 48/210 (22%), Positives = 70/210 (32%), Gaps = 4/210 (1%)
Frame = +1
Query: 311 SQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV-SSEPII 369
+ P TSP +SP SP + PS + +PTS S PTS P P SS
Sbjct: 2242 TSPGYSPTSPGYSPTSPTYSPSSPGYSPTSPAYSPTSPSYSPTSPAYSPTSPAYSSTSPA 2421
Query: 370 HEQPEPNQTEPQPRTSDHSVPRASARPAARTTD---TDSSTAFTPVSFPINVIDSPPSNT 426
+ P+ + P S S + P+ T + +S +++P S + S T
Sbjct: 2422 YSPTSPSYSPTSPAYSPTSPSYSLTSPSYSPTSPSYSPTSPSYSPTSPAYSPTSPSYSPT 2601
Query: 427 SESIRKFMEVRKEKVSALEEYYLTCPSPRRYPGPRPERLVDPDEPILANPIQEADPLVQQ 486
S S + + PS R P P P + P
Sbjct: 2602 SPSYSPTSPSYNPQSAKYSPSLAYSPSSPRLSPTSPN--YSPTSPSYSPTSPSYSPSSPT 2775
Query: 487 AHPVPDQQEPIQPDPEPEQSVSNQSSVRSP 516
P + PD P + S+ SP
Sbjct: 2776 YSPSSPYNSGVSPDYSPTSPQFSPSAGYSP 2865
Score = 38.9 bits (89), Expect = 0.009
Identities = 33/121 (27%), Positives = 50/121 (41%)
Frame = +1
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDP 352
P S SS +P + PA TSP +SP SP + P+ + +PTS S P
Sbjct: 2281 PTSPTYSPSSPGYSP---TSPAYSPTSPSYSPTSPAYSPTSPAYSSTSPAYSPTSPSYSP 2451
Query: 353 TSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPV 412
TS P P S + P P + +S P + A + + +S +++P
Sbjct: 2452 TSPAYSPTSP--SYSLTSPSYSPTSPSYSPTSPSYS-PTSPAYSPTSPSYSPTSPSYSPT 2622
Query: 413 S 413
S
Sbjct: 2623 S 2625
>TC226825 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydratase, small
subunit, partial (71%)
Length = 1121
Score = 41.2 bits (95), Expect = 0.002
Identities = 39/138 (28%), Positives = 59/138 (42%), Gaps = 7/138 (5%)
Frame = +3
Query: 294 ISALLQHSSNPLTPIPESQP---AAQTTSPPHS--PRSPFFQPSPTEAPLWN--LLKNPT 346
++ L + ++ L+ +P Q A+Q+ S PH+ P P P P+ A + + PT
Sbjct: 81 LTLTLFYRASFLSQLPSHQTLATASQSLSKPHALNPPRPLLPPPPSTASATSSATISTPT 260
Query: 347 SRSEDPTSLLTIPYDPVSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSS 406
TS ++P P S+ P P+ P P PR S+ PA T SS
Sbjct: 261 RSFPPSTSPSSLP-SPTSTRS---SAPTPSSASPPP------TPRVSSNPARSKPSTPSS 410
Query: 407 TAFTPVSFPINVIDSPPS 424
+A P S +PPS
Sbjct: 411 SA-VPTSVAAPPASTPPS 461
>TC209975 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (9%)
Length = 537
Score = 41.2 bits (95), Expect = 0.002
Identities = 37/124 (29%), Positives = 47/124 (37%)
Frame = +1
Query: 304 PLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPYDPV 363
P TP P P T PP +P SP QP T P K P+ S+ P + P P+
Sbjct: 82 PKTPSPSYSPPNVPT-PPKTP-SPSSQPPFTPTPP----KTPSPTSQPP--YIPTPPSPI 237
Query: 364 SSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPP 423
S P I P QP P S P+ + + SS P S+ P
Sbjct: 238 SQPPSIATPPNTLSPTSQPPYPSTIPPSTSLSPSYPPSPSPSSAPTYPPSYLAPATSPPS 417
Query: 424 SNTS 427
+TS
Sbjct: 418 PSTS 429
Score = 37.7 bits (86), Expect = 0.019
Identities = 33/106 (31%), Positives = 41/106 (38%), Gaps = 1/106 (0%)
Frame = +1
Query: 293 PISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDP 352
PIS ++ P T P SQP +T PP + SP + PSP +P+S P
Sbjct: 232 PISQPPSIATPPNTLSPTSQPPYPSTIPPSTSLSPSYPPSP----------SPSSAPTYP 381
Query: 353 TSLLTIPYDPVS-SEPIIHEQPEPNQTEPQPRTSDHSVPRASARPA 397
S L P S S P T P P +S S PA
Sbjct: 382 PSYLAPATSPPSPSTSATTPTISPAATTPSPSSSSSSGASNETTPA 519
>TC221561 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate dehydrogenase,
complete
Length = 2001
Score = 40.8 bits (94), Expect = 0.002
Identities = 39/148 (26%), Positives = 63/148 (42%), Gaps = 4/148 (2%)
Frame = -1
Query: 366 EPIIHEQPEPNQTEPQPRT-SDHSVP--RASARPAARTTDTDSSTAFTPVSFPINVIDSP 422
+P H QP+P +++ QP T S HS +A +P++ T+ + + +P ++P +
Sbjct: 1122 QPSTH-QPQP-ESDSQPGT*SQHSPQSHQAQGKPSSHTSPSLTDKLSSPENYPTLQHSTQ 949
Query: 423 PSNTSESIRKFMEVRKEKVSALEEYYLTCPSPRRYP-GPRPERLVDPDEPILANPIQEAD 481
PS + + Y+L PR P PRP P P+ NP
Sbjct: 948 PSPNAPT---------------SPYHLPSSPPRPPP*TPRPPP-PSPAPPVSGNPSPSPP 817
Query: 482 PLVQQAHPVPDQQEPIQPDPEPEQSVSN 509
P P P Q+P P P+P+ S+
Sbjct: 816 P------PSPPNQQPHHPSPKPQHHFSH 751
Score = 29.3 bits (64), Expect = 6.8
Identities = 18/60 (30%), Positives = 28/60 (46%)
Frame = +2
Query: 289 FDALPISALLQHSSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSR 348
F A P+ + + + P P + P++ TSPP S R + SP +P +PT R
Sbjct: 359 FVASPMDTVSESAWPPPWPPSAASPSSTPTSPPPSRRPSSAKRSPAASPSSPTPPSPTFR 538
>BM891343
Length = 418
Score = 40.8 bits (94), Expect = 0.002
Identities = 35/134 (26%), Positives = 51/134 (37%), Gaps = 11/134 (8%)
Frame = +2
Query: 301 SSNPLTPIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRSEDPTSLLTIPY 360
S++ TP S P++ T P +P SP PSP+ + W N TSR +
Sbjct: 2 SASKTTPKSPS-PSSTTPKPSPTPLSPT-PPSPSSSTPWLRFANSTSRGNSSSKWTNATT 175
Query: 361 DP-----------VSSEPIIHEQPEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAF 409
P SS P +P T P P+ SA + R+ T +S +
Sbjct: 176 SPQPLPLS*P*FAASSAPASRAKPYAPSTTSTPSQKPKRHPKTSAFCSTRSASTATSDSP 355
Query: 410 TPVSFPINVIDSPP 423
+ S N + PP
Sbjct: 356 SRFSTKTNTLSPPP 397
>TC205952
Length = 1414
Score = 40.8 bits (94), Expect = 0.002
Identities = 30/140 (21%), Positives = 58/140 (41%), Gaps = 7/140 (5%)
Frame = +1
Query: 373 PEPNQTEPQPRTSDHSVPRASARPAARTTDTDSSTAFTPVSFPINVIDSPPSNTSESIRK 432
P+ + E + + VP + +A T + F P + + ++ +P ++ S + +
Sbjct: 586 PDASSIEERNALALAIVPTETGTTSAFNTTAAQTKDFDPTGWELALVSTPSTDISAANER 765
Query: 433 FMEVRKEKVSALEEY----YLTCPSPRRYPGPRPERLVDP---DEPILANPIQEADPLVQ 485
+ + ++ Y Y + P P P + DP I P + + Q
Sbjct: 766 QLAGGLDSLTLNSLYDEAAYRSQQPVYGAPAPNPFEMQDPFALSSSIPPPPAVQLAAMQQ 945
Query: 486 QAHPVPDQQEPIQPDPEPEQ 505
QA+P Q+P QP P+P+Q
Sbjct: 946 QANPFGPYQQPFQPQPQPQQ 1005
>TC228448 weakly similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (4%)
Length = 458
Score = 40.8 bits (94), Expect = 0.002
Identities = 42/130 (32%), Positives = 60/130 (45%), Gaps = 14/130 (10%)
Frame = +1
Query: 298 LQHSSNPLT--------PIPESQPAAQTTSPPHSPRSPFFQPSPTEAPLWNLLKNPTSRS 349
LQH +P + P P S A+ + P+ P SP +PSP AP +P SRS
Sbjct: 82 LQHPHHPPSSPSPLKP*PYPHSNLASPSGPTPNPPFSPTSKPSP--AP----RSSPPSRS 243
Query: 350 EDP---TSLLTIPYDPVSSEPIIHEQP-EPNQTEPQPRTSDHSVPRAS-ARPAARTT-DT 403
P L + P ++ P P ++P PR S S P +S + +ARTT +
Sbjct: 244 NSPPPHVPPLRLLSKPTTASTSRSPSPLSPTSSKPTPRPSPRSSPSSSRSSSSARTTRRS 423
Query: 404 DSSTAFTPVS 413
SS+A +P S
Sbjct: 424 PSSSASSPRS 453
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,107,488
Number of Sequences: 63676
Number of extensions: 563732
Number of successful extensions: 6812
Number of sequences better than 10.0: 505
Number of HSP's better than 10.0 without gapping: 5090
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6193
length of query: 766
length of database: 12,639,632
effective HSP length: 105
effective length of query: 661
effective length of database: 5,953,652
effective search space: 3935363972
effective search space used: 3935363972
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0014.15


