

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0011a.2
(391 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
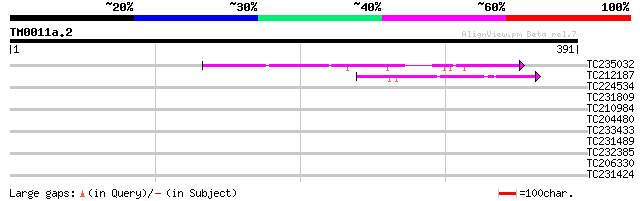
Score E
Sequences producing significant alignments: (bits) Value
TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarit... 102 4e-22
TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragm... 42 5e-04
TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), p... 35 0.076
TC231809 34 0.099
TC210984 34 0.13
TC204480 similar to UP|Q947C3 (Q947C3) 1-deoxy-D-xylulose-5-phos... 30 1.9
TC233433 28 5.5
TC231489 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F1... 28 5.5
TC232385 28 7.1
TC206330 weakly similar to UP|Q93W03 (Q93W03) AT3g56130/F18O21_9... 28 7.1
TC231424 28 9.3
>TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarity to
ribosomal protein, partial (24%)
Length = 689
Score = 102 bits (253), Expect = 4e-22
Identities = 86/240 (35%), Positives = 115/240 (47%), Gaps = 18/240 (7%)
Frame = +3
Query: 134 KDIARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQDL 193
+ I LM++ + RGF MLGSID +HW + AWKGQYC GD G PTI+L+ + SQ L
Sbjct: 6 RGIQYLMQICEIRGFQSMLGSIDYLHWK*KTCLVAWKGQYCHGD-GPPTIMLKVVVSQYL 182
Query: 194 WIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQF----------TLNGTTYNMGYY 243
I HA+ GV GSNN+INVLN+S V G++P L+ Y
Sbjct: 183 RI*HAYLGVEGSNNNINVLNESCV*RR-YGGRSPYCTICS**NPI*HGVLSDIWYLSRLD 359
Query: 244 LADEIYPEWATFVKTI--SMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGP--A 299
A E +P+ K I S+ + K+ ++ F +VR +
Sbjct: 360 DACEDHPKAIRS*KKIVYSISRSNKKGSHLKY------------------FNLVRNYTWS 485
Query: 300 RAW--HLHILKDIIY--ACIILHNMIVEDEREMCDGNIDFSYDQLENDASTPEVFSGPHP 355
W H HI I Y CIILHNMIV+DE + NID Y N+ ++ G HP
Sbjct: 486 CVWLEHEHI-SSIYYMYVCIILHNMIVKDEPHTYNENIDRQYKYPTNNVLIIDIHIGLHP 662
>TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment),
partial (45%)
Length = 973
Score = 42.0 bits (97), Expect = 5e-04
Identities = 36/139 (25%), Positives = 63/139 (44%), Gaps = 12/139 (8%)
Frame = +1
Query: 240 MGYYLADEIYPEWATFVKTIS-----MPQGE-------KRKLFAQHQEGARKDVERAFGV 287
+ +YL D YP + F+ + + +PQ + ++F + + +ERAFG+
Sbjct: 508 VNFYLVDSCYPTFMGFLGSYNKTRYHLPQFRIGPRIRGRVEVFNYYHSSLQSTIERAFGL 687
Query: 288 L*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCDGNIDFSYDQLENDASTP 347
+R+ I+ G + L II AC+ +HN I +++ DG D S D+ T
Sbjct: 688 CKARWKIL-GNMSPFALKTQNQIIVACMTIHNFIQRNDKS--DGEFD-SLDEDNEYIDTD 855
Query: 348 EVFSGPHPDFATYLQKRAQ 366
+ S P T+ + AQ
Sbjct: 856 KDESEVGPSTTTWEESDAQ 912
>TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), partial (50%)
Length = 968
Score = 34.7 bits (78), Expect = 0.076
Identities = 22/65 (33%), Positives = 37/65 (56%)
Frame = +1
Query: 262 PQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMI 321
PQ K +LF A+ +ER+FGVL R++I+R P+ + + II C +L N I
Sbjct: 577 PQNYK-ELFNLRHASAKNVIERSFGVLKKRWSILRTPS-FFDIKTQIRIINTCFMLRNFI 750
Query: 322 VEDER 326
++++
Sbjct: 751 RDEQQ 765
>TC231809
Length = 760
Score = 34.3 bits (77), Expect = 0.099
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = +3
Query: 267 RKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDER 326
++LF R + R+F VL +RF I++ A + I +DI+ A +LHN I +ER
Sbjct: 207 KELFNHRHCFLRNAILRSFNVLKTRFPILK-LAPQYSFQIQRDIVIAGCVLHNFIRREER 383
>TC210984
Length = 1085
Score = 33.9 bits (76), Expect = 0.13
Identities = 27/104 (25%), Positives = 46/104 (43%), Gaps = 1/104 (0%)
Frame = +3
Query: 267 RKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDER 326
++LF R +ER FG+ SRF I + A + +++ AC LHN + R
Sbjct: 423 KELFNLRHASLRNVIERIFGIFKSRFTIFKS-APPFLFKTQAELVLACAALHNFL----R 587
Query: 327 EMCDGNIDFSYDQLENDASTPEVFSG-PHPDFATYLQKRAQVRE 369
+ C + +F + + +S+ V D +Q + Q RE
Sbjct: 588 KECRSD-EFPVEPTDESSSSSSVLPNYEDNDHEPIVQTQEQERE 716
>TC204480 similar to UP|Q947C3 (Q947C3) 1-deoxy-D-xylulose-5-phosphate
reductoisomerase, partial (92%)
Length = 2110
Score = 30.0 bits (66), Expect = 1.9
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +3
Query: 367 VREKTVHLHLQADLVEHVWEHFRR 390
+REK+V LHL+ + V W H R
Sbjct: 387 LREKSVELHLEGEFVAQCWHHHHR 458
>TC233433
Length = 754
Score = 28.5 bits (62), Expect = 5.5
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = +1
Query: 126 EYLRKTNNKDIARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGK 180
E L+ +KD + E +G G+ GS+DC H I +G +G +GD+ +
Sbjct: 475 EDLQGDGDKDDRAIGEAS*RKGSGGVKGSVDCAHEIRVHGELPSRGS-LQGDYNR 636
>TC231489 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;} , partial (22%)
Length = 634
Score = 28.5 bits (62), Expect = 5.5
Identities = 23/93 (24%), Positives = 37/93 (39%)
Frame = +3
Query: 30 NPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATGRMGLSPLQKQTAAIRMLA 89
+P + EE+F+ +RM K F I + + D + + R + Q+ I LA
Sbjct: 360 SPDFPEEEFRLHFRMSKATFDMICQHL----DSAVTKKNTMLREAIPVRQRVAVCIWRLA 527
Query: 90 YGSSTDSVDEYVCIGESTARKCMARFVKGINEV 122
G V + +G ST K + I V
Sbjct: 528 TGDPLREVSKRFGLGISTCHKLVLEVCSAIKSV 626
>TC232385
Length = 572
Score = 28.1 bits (61), Expect = 7.1
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +3
Query: 183 IILEAIASQDLWIWHAFFGVA 203
+I ++ + DLW W FFG A
Sbjct: 414 LIFSSVLALDLWFWELFFGTA 476
>TC206330 weakly similar to UP|Q93W03 (Q93W03) AT3g56130/F18O21_90, partial
(65%)
Length = 1307
Score = 28.1 bits (61), Expect = 7.1
Identities = 18/79 (22%), Positives = 36/79 (44%)
Frame = +3
Query: 311 IYACIILHNMIVEDEREMCDGNIDFSYDQLENDASTPEVFSGPHPDFATYLQKRAQVREK 370
I CI+ + E +R +C ++ QL + ++ HP A +L + ++++
Sbjct: 582 ILKCILSETL--EQQRFLC-----LTFHQLHHHLFQVNLWMNQHP-VACHLHHQNHLQKR 737
Query: 371 TVHLHLQADLVEHVWEHFR 389
T HL + W+H+R
Sbjct: 738 TTHLQMFPKRNHQDWQHWR 794
>TC231424
Length = 1075
Score = 27.7 bits (60), Expect = 9.3
Identities = 13/44 (29%), Positives = 20/44 (44%)
Frame = +2
Query: 209 INVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEW 252
+ +LN +D ND + PE + + Y D +YPEW
Sbjct: 551 VGLLNFNDSENDQWKELIPEAEHVVLHLNYTSSNITWDVLYPEW 682
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,196,773
Number of Sequences: 63676
Number of extensions: 231288
Number of successful extensions: 1121
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1115
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1119
length of query: 391
length of database: 12,639,632
effective HSP length: 99
effective length of query: 292
effective length of database: 6,335,708
effective search space: 1850026736
effective search space used: 1850026736
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0011a.2


