

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
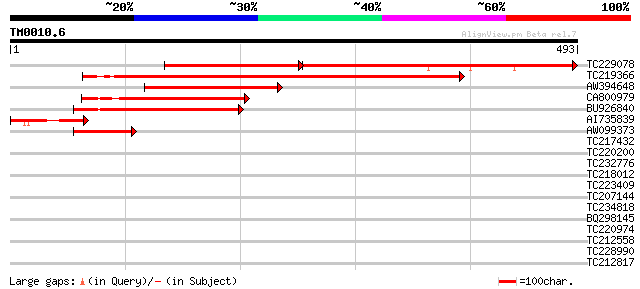
TC229078 similar to UP|O82458 (O82458) Rac GTPase activating pro... 358 e-158
TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating pro... 397 e-111
AW394648 similar to GP|3695059|gb| rac GTPase activating protein... 224 9e-59
CA800979 similar to GP|3695059|gb| rac GTPase activating protein... 211 5e-55
BU926840 similar to PIR|T01383|T01 GTPase-activating protein hom... 195 3e-50
AI735839 60 2e-09
AW099373 similar to GP|3695063|gb|A rac GTPase activating protei... 49 4e-06
TC217432 similar to UP|O52943 (O52943) Plasmid PLS32 DNA for thy... 33 0.38
TC220200 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific... 32 0.65
TC232776 weakly similar to UP|SFR6_HUMAN (Q13247) Splicing facto... 32 0.85
TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 31 1.1
TC223409 similar to UP|Q6T4P5 (Q6T4P5) Plasticity related gene 2... 30 1.9
TC207144 30 1.9
TC234818 similar to UP|O22082 (O22082) C2H2 zinc-finger protein ... 30 2.5
BQ298145 similar to GP|16326135|db Myb {Nicotiana tabacum}, part... 30 2.5
TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus pro... 30 3.2
TC212558 30 3.2
TC228990 UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1, comp... 30 3.2
TC212817 weakly similar to UP|Q9FFD5 (Q9FFD5) Gb|AAC80617.1 (At5... 30 3.2
>TC229078 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (70%)
Length = 1579
Score = 358 bits (920), Expect(2) = e-158
Identities = 185/248 (74%), Positives = 211/248 (84%), Gaps = 9/248 (3%)
Frame = +2
Query: 255 EAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTL 314
++ALLDWAINLMADVAQME+ NKMNARNIAMVFAPNMT MADPLTALMYAVQVM+FLKTL
Sbjct: 422 KSALLDWAINLMADVAQMENLNKMNARNIAMVFAPNMTQMADPLTALMYAVQVMNFLKTL 601
Query: 315 VVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCS--DEDTVFVSAE 372
VVK LREREESIVKSNPVP+LNSFDDDG+ S+S++L K+ SENGNDCS DEDTVFV+AE
Sbjct: 602 VVKALREREESIVKSNPVPDLNSFDDDGNHSNSEMLDKEDSENGNDCSDDDEDTVFVTAE 781
Query: 373 PSQPSPTHHTEDGCETESGSETSPTPA--ENFLSSGSRLLIDSCPCNVVSQLCSFAIGLQ 430
PSQ SP+H TEDGCETE+ +++ PA EN++SSG+RLL+DSCPCN+VSQ+CS AIGLQ
Sbjct: 782 PSQQSPSHLTEDGCETETATKSKSLPASTENYISSGNRLLVDSCPCNLVSQICSLAIGLQ 961
Query: 431 DSSIATG-----QAKISRSKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGTAIIGRINSR 485
D +ATG QAKI RSKSLQ+ST D DK + VI+ PV G AEKN G AII RINSR
Sbjct: 962 DCGLATGQTKGDQAKICRSKSLQLSTYDTDKCSRKVIQLPVTGAAEKNLGMAIIERINSR 1141
Query: 486 TELTEAWR 493
TEL EAWR
Sbjct: 1142TELAEAWR 1165
Score = 218 bits (556), Expect(2) = e-158
Identities = 107/121 (88%), Positives = 115/121 (94%)
Frame = +1
Query: 135 ASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVR 194
+S +VFGVSTESMQLSFDARGNSVPTILLLMQRHLYA+GGLQAEGIFRI+AEN QEE VR
Sbjct: 61 SSANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQAEGIFRIDAENGQEEFVR 240
Query: 195 EQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPT 254
E LNRGVVP+G+DVHCLAGLIKAWFRELPTG+LDP SPE+VMQSQSEEEC QLVRLLPPT
Sbjct: 241 E*LNRGVVPDGIDVHCLAGLIKAWFRELPTGVLDPFSPEQVMQSQSEEECAQLVRLLPPT 420
Query: 255 E 255
E
Sbjct: 421 E 423
>TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating protein,
partial (52%)
Length = 1397
Score = 397 bits (1019), Expect = e-111
Identities = 210/332 (63%), Positives = 250/332 (75%)
Frame = +1
Query: 64 LLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVE 123
LL LL+ RKS PR + S M+IG P+NVRHVAHVTFDRF+GFLGLPVE
Sbjct: 7 LLALLLTLLRKSF-----QLPRKN---IGSIMDIGSPTNVRHVAHVTFDRFNGFLGLPVE 162
Query: 124 FEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRI 183
FEPEVPRRPPSAS SVFGVSTESMQLS+D+RGNSVPTILLLMQRHLY +GGLQ EGIFRI
Sbjct: 163 FEPEVPRRPPSASASVFGVSTESMQLSYDSRGNSVPTILLLMQRHLYVQGGLQVEGIFRI 342
Query: 184 NAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEE 243
NA+N QEE R+QLN GVVP G+DVHCLAGLIKAWFRELPTGILD LSPE+VMQ Q+E+E
Sbjct: 343 NADNGQEEHARDQLNLGVVPEGIDVHCLAGLIKAWFRELPTGILDSLSPEQVMQCQTEDE 522
Query: 244 CDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMY 303
C +LVR LP TEA+LLDWAINLMADV Q E+ NKMNA NIAMVFAPNMT MADP++ALMY
Sbjct: 523 CAELVRHLPHTEASLLDWAINLMADVVQHENVNKMNAHNIAMVFAPNMTQMADPISALMY 702
Query: 304 AVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSD 363
AVQVM+FLKTL+++T+RER++S+V+S P L D+ ++ + +D + +
Sbjct: 703 AVQVMNFLKTLILRTVRERKDSVVESCPRFYLQPSVDNENRRILESFRQDTPAENEEAQE 882
Query: 364 EDTVFVSAEPSQPSPTHHTEDGCETESGSETS 395
+ +A P + G E S + +S
Sbjct: 883 NFVLEKTALDRSPESLQNNSTGGEPGSLTNSS 978
>AW394648 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (24%)
Length = 361
Score = 224 bits (570), Expect = 9e-59
Identities = 108/120 (90%), Positives = 115/120 (95%)
Frame = +2
Query: 118 LGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQA 177
LGLPVEFEPEVPRRPPSAS +VFGVSTESMQLSFDARGNSVPTILLLMQRHLYA+GGLQA
Sbjct: 2 LGLPVEFEPEVPRRPPSASANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQA 181
Query: 178 EGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ 237
EGIFRINAEN QEE VREQLNRG+VP+G+DVHCLAGLIKAWFRELP +LDPLSPE+VMQ
Sbjct: 182 EGIFRINAENGQEEFVREQLNRGIVPDGIDVHCLAGLIKAWFRELPNWVLDPLSPEQVMQ 361
>CA800979 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (25%)
Length = 421
Score = 211 bits (538), Expect = 5e-55
Identities = 109/146 (74%), Positives = 121/146 (82%)
Frame = +2
Query: 63 SLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPV 122
SLL +L+ RKSLI +C+ S GA MEIGWP+NVRHVAHVTFDRF+GFLGLP
Sbjct: 2 SLLAILVTLLRKSLI-ACNKSEEGHGA-----MEIGWPTNVRHVAHVTFDRFNGFLGLPR 163
Query: 123 EFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFR 182
EFEPEV RPPSAS +VFGVSTESMQLS+D RGNSVPTILLLMQRHLYA GGLQ EGIFR
Sbjct: 164 EFEPEVSTRPPSASATVFGVSTESMQLSYDTRGNSVPTILLLMQRHLYALGGLQEEGIFR 343
Query: 183 INAENSQEELVREQLNRGVVPNGVDV 208
INA+NSQEE VR+QLNRG+VP VD+
Sbjct: 344 INADNSQEESVRDQLNRGLVPEDVDI 421
>BU926840 similar to PIR|T01383|T01 GTPase-activating protein homolog T4I9.2
- Arabidopsis thaliana, partial (32%)
Length = 443
Score = 195 bits (496), Expect = 3e-50
Identities = 96/148 (64%), Positives = 118/148 (78%)
Frame = +3
Query: 56 DREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFH 115
+ + +QLSL+ LL+A RKS++ SC P + + MEIGWP+NV+H+ HVTFDRF+
Sbjct: 3 EEQQNQLSLVALLLAAIRKSMV-SCRVDPPEDVISTVHHMEIGWPTNVQHITHVTFDRFN 179
Query: 116 GFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGL 175
GFLGLP EF+ E+P R PSAS SVFGVS ESMQ S+D +GNSVPTILLLMQ LY++GGL
Sbjct: 180 GFLGLPYEFQVEIPARVPSASVSVFGVSAESMQCSYDPKGNSVPTILLLMQDRLYSQGGL 359
Query: 176 QAEGIFRINAENSQEELVREQLNRGVVP 203
+AEGIFRIN ENSQEE VR+QLNRG+VP
Sbjct: 360 KAEGIFRINPENSQEEHVRDQLNRGIVP 443
>AI735839
Length = 425
Score = 60.5 bits (145), Expect = 2e-09
Identities = 40/76 (52%), Positives = 47/76 (61%), Gaps = 8/76 (10%)
Frame = +2
Query: 1 MTEVLQLPSPS----RRCS----NDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEE 52
MTEVLQLPS S R C NDGSH +LIN +VE+ VE +EEEE
Sbjct: 224 MTEVLQLPSSSSCSRRPCGSLTPNDGSHPISLINAPPTVED---------QRVEIEEEEE 376
Query: 53 KEKDREGDQLSLLTLL 68
KE++RE DQLS+LTLL
Sbjct: 377 KERERERDQLSILTLL 424
>AW099373 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 416
Score = 49.3 bits (116), Expect = 4e-06
Identities = 23/55 (41%), Positives = 34/55 (61%)
Frame = +2
Query: 56 DREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVT 110
+ E + S + +L+A RKS++ SP D + MEIGWP+NV+HV+HVT
Sbjct: 251 EEEQNPASPVAVLLAALRKSMVACSVDSPDDVISAVHHPMEIGWPTNVKHVSHVT 415
>TC217432 similar to UP|O52943 (O52943) Plasmid PLS32 DNA for thymidine
kinase, replication protein, complete cds, partial (14%)
Length = 1041
Score = 32.7 bits (73), Expect = 0.38
Identities = 18/96 (18%), Positives = 39/96 (39%), Gaps = 5/96 (5%)
Frame = +3
Query: 16 NDGSHQTALINNSSSVEEGVLQLQTH-----LDLVEEDEEEEKEKDREGDQLSLLTLLIA 70
N H+ + + + +L+ H +L++ E ++ + LS +T +A
Sbjct: 171 NPSKHKVYAVLKDQAKSDDILKAAFHAHVLLFNLIKSWNESNASSLKQREDLSNMTHTVA 350
Query: 71 TFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHV 106
+ G+C T G +++ E GW + H+
Sbjct: 351 DIEACMAGTCKTVADSYGFFKNNAKEQGWTMSESHL 458
>TC220200 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein T2
precursor, partial (40%)
Length = 506
Score = 32.0 bits (71), Expect = 0.65
Identities = 31/115 (26%), Positives = 45/115 (38%), Gaps = 1/115 (0%)
Frame = +2
Query: 286 VFAPNMTH-MADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQ 344
+F+PN TH MA+P +A V+ L +L+V + P P D
Sbjct: 92 LFSPNSTHSMANPASARF---SVLILLVSLLVNIAATAATAAESPAPTPT-----PDFPP 247
Query: 345 SDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPA 399
S S P + S+ T P+ P+P+HH+ S SP PA
Sbjct: 248 SSSPPFP-------SPASNPSTANFPPSPT-PAPSHHSPPAPSPSSNPSPSPAPA 388
>TC232776 weakly similar to UP|SFR6_HUMAN (Q13247) Splicing factor,
arginine/serine-rich 6 (Pre-mRNA splicing factor SRP55),
partial (7%)
Length = 604
Score = 31.6 bits (70), Expect = 0.85
Identities = 17/52 (32%), Positives = 31/52 (58%)
Frame = +2
Query: 8 PSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREG 59
P+P+ RC ++ SH+ + S +V + Q+ + +VE+ EEEE E++ G
Sbjct: 128 PNPATRCVSNRSHKPPHLARSDAVS*KMGASQS-VQVVEDSEEEEDERNLNG 280
>TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(54%)
Length = 1142
Score = 31.2 bits (69), Expect = 1.1
Identities = 21/72 (29%), Positives = 30/72 (41%)
Frame = -1
Query: 342 GHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAEN 401
GH D +L E D DED E ++ H E+G + G E T +
Sbjct: 785 GHLFDGDILSLGQQEKHEDRHDEDPAREEQEDAKLEVAEHREEGLCNDEGEEQVNTNGDT 606
Query: 402 FLSSGSRLLIDS 413
L+SG+ L +S
Sbjct: 605 -LTSGTSLQWES 573
>TC223409 similar to UP|Q6T4P5 (Q6T4P5) Plasticity related gene 2a, partial
(3%)
Length = 439
Score = 30.4 bits (67), Expect = 1.9
Identities = 14/31 (45%), Positives = 21/31 (67%)
Frame = +1
Query: 46 EEDEEEEKEKDREGDQLSLLTLLIATFRKSL 76
EEDEEE++E+D E D+ L + L F+ +L
Sbjct: 49 EEDEEEDEEEDEEEDEAPLSSSLEIKFKTAL 141
>TC207144
Length = 1174
Score = 30.4 bits (67), Expect = 1.9
Identities = 14/58 (24%), Positives = 30/58 (51%), Gaps = 3/58 (5%)
Frame = +1
Query: 339 DDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTE---DGCETESGSE 393
++DG +++++ + + SE + +E + +P QP PT + E E+GS+
Sbjct: 268 ENDGAEAETEPVTVETSEGSTEEEEEKVIEPKEQPQQPGPTREPNSDPESMERENGSK 441
>TC234818 similar to UP|O22082 (O22082) C2H2 zinc-finger protein ZPT2-10,
partial (19%)
Length = 684
Score = 30.0 bits (66), Expect = 2.5
Identities = 21/50 (42%), Positives = 27/50 (54%)
Frame = +3
Query: 43 DLVEEDEEEEKEKDREGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSS 92
D+VEE+EEEE E++ EG + L L K SC + R S AL S
Sbjct: 123 DVVEEEEEEE-EEEEEGGRSVLEIKLRRVRGKHQCQSCGKTFRSSRALGS 269
>BQ298145 similar to GP|16326135|db Myb {Nicotiana tabacum}, partial (7%)
Length = 422
Score = 30.0 bits (66), Expect = 2.5
Identities = 25/71 (35%), Positives = 30/71 (42%), Gaps = 8/71 (11%)
Frame = +1
Query: 348 QVLPKDGSENGNDCSDEDTVFV------SAEPSQPSPTHHTED--GCETESGSETSPTPA 399
+ LPKD SEN D E+ V S PS T D CET E S + A
Sbjct: 103 KALPKDASENDKDKDHENNYPVNQSGNHSQLPSNALIERRTLDFSECETPGRDENSKSSA 282
Query: 400 ENFLSSGSRLL 410
+F S S L+
Sbjct: 283 MSFSSPSSYLM 315
>TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus protein p62,
partial (7%)
Length = 436
Score = 29.6 bits (65), Expect = 3.2
Identities = 23/87 (26%), Positives = 34/87 (38%), Gaps = 11/87 (12%)
Frame = +1
Query: 323 EESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDED-----------TVFVSA 371
EE + P N ++D S + +D E S+E+ +
Sbjct: 133 EEEPQQQQPSSQKNKEEEDEEVSSGEEEEEDEEEEEAASSEEEEEDEDLPPPPVSKNPPP 312
Query: 372 EPSQPSPTHHTEDGCETESGSETSPTP 398
P+ P P H + + ETESGSET P
Sbjct: 313 PPANPQPQHSSSES-ETESGSETESEP 390
>TC212558
Length = 620
Score = 29.6 bits (65), Expect = 3.2
Identities = 14/22 (63%), Positives = 17/22 (76%)
Frame = -3
Query: 43 DLVEEDEEEEKEKDREGDQLSL 64
DLV E EEEEKEK ++G+ L L
Sbjct: 300 DLVFESEEEEKEKKKKGELLFL 235
>TC228990 UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1, complete
Length = 3346
Score = 29.6 bits (65), Expect = 3.2
Identities = 21/93 (22%), Positives = 39/93 (41%)
Frame = +2
Query: 323 EESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHT 382
E + S P P+ +F S ++ LP + + S ++ P+ P+ T
Sbjct: 542 ENFLSPSLPCPSSATFTSAATSSPARSLPSTAPGSTSSTSPSPA---TSSPAPSRPSSET 712
Query: 383 EDGCETESGSETSPTPAENFLSSGSRLLIDSCP 415
+ + + T+PTPA SR +++CP
Sbjct: 713 SPPSASSTSATTTPTPA------ASRPRLETCP 793
>TC212817 weakly similar to UP|Q9FFD5 (Q9FFD5) Gb|AAC80617.1 (At5g16490),
partial (18%)
Length = 800
Score = 29.6 bits (65), Expect = 3.2
Identities = 11/18 (61%), Positives = 15/18 (83%)
Frame = +2
Query: 95 MEIGWPSNVRHVAHVTFD 112
MEIG P+NV+HV H+ +D
Sbjct: 449 MEIGCPTNVQHVTHIGWD 502
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,438,140
Number of Sequences: 63676
Number of extensions: 282302
Number of successful extensions: 1551
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1480
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1535
length of query: 493
length of database: 12,639,632
effective HSP length: 101
effective length of query: 392
effective length of database: 6,208,356
effective search space: 2433675552
effective search space used: 2433675552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0010.6


