

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.16
(405 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
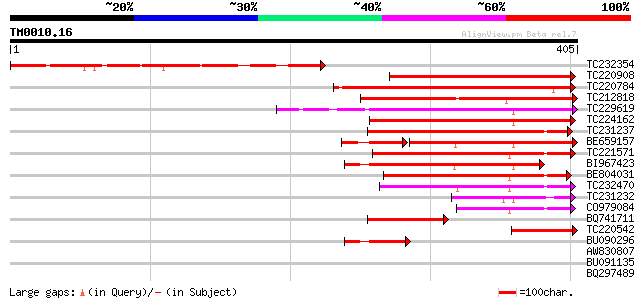
Score E
Sequences producing significant alignments: (bits) Value
TC232354 239 1e-63
TC220908 230 1e-60
TC220784 similar to GB|AAD03427.1|4115916|F3H7 F3H7.9 {Arabidops... 182 2e-46
TC212818 weakly similar to UP|Q9FNG2 (Q9FNG2) Emb|CAB81925.1, pa... 163 1e-40
TC229619 144 5e-35
TC224162 137 9e-33
TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644 At2g4... 133 2e-31
BE659157 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidops... 111 2e-31
TC221571 similar to GB|AAO63337.1|28950827|BT005273 At5g58530 {A... 121 6e-28
BI967423 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidops... 120 8e-28
BE804031 weakly similar to GP|15624060|dbj B1088C09.21 {Oryza sa... 115 4e-26
TC232470 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g5... 114 1e-25
TC231232 similar to UP|Q9LPI8 (Q9LPI8) F6N18.14, partial (10%) 78 6e-15
CO979084 74 1e-13
BQ741711 61 8e-10
TC220542 similar to GB|AAL69460.1|18491271|AY074644 At2g41330/F1... 61 8e-10
BU090296 homologue to GP|14532460|gb| AT5g13810/MAC12_24 {Arabid... 45 5e-05
AW830807 weakly similar to PIR|T48082|T480 glutaredoxin-like pro... 39 0.004
BU091135 homologue to GP|10241626|emb Contains similarity to F1N... 38 0.007
BQ297489 37 0.021
>TC232354
Length = 677
Score = 239 bits (611), Expect = 1e-63
Identities = 143/239 (59%), Positives = 167/239 (69%), Gaps = 14/239 (5%)
Frame = +1
Query: 1 MGCVSSKLIRKDIKQEHVIIDNCGGGGRYLSHVVSLTSSTYGALKLDKDNDQ----PVIA 56
MGCVSSK I+KD+KQE + GGG Y++HVVSLTSSTYGAL LDK+ +Q P +
Sbjct: 28 MGCVSSKHIKKDLKQEEEVTVTNGGG--YVNHVVSLTSSTYGALMLDKEKEQLQLQPTVE 201
Query: 57 AKP-------REEPEAITTINAWELMEGLEDGVPISNQPKKSPKSSSPFLRGFMNSDTR- 108
+ REEPE I NAWELMEGLE+GVPISN P KSPK S+PFLRGF+++D +
Sbjct: 202 EESKTSPPRIREEPEVI---NAWELMEGLEEGVPISNNPMKSPK-STPFLRGFISTDPKP 369
Query: 109 --SPLKFLNQLGSPKTVKKSTGKENKVQVNGVRTGGVRRLDYSNSPKGILKPSNLSPNAS 166
+P KFLNQ GSPK+++K TGKENKVQV GVRRLDY+ SPKGILKPSN
Sbjct: 370 KITPFKFLNQFGSPKSLRKFTGKENKVQVQ-PPNAGVRRLDYNFSPKGILKPSNFCS--- 537
Query: 167 KNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASYEKELSEEEEQV 225
P KGSPI ARR SFG + K R+S SPLFDPEL+ASYEKELSEEEEQ+
Sbjct: 538 -----PLKGSPIRARRNSFGTDIK--------RRSPSPLFDPELLASYEKELSEEEEQI 675
>TC220908
Length = 673
Score = 230 bits (586), Expect = 1e-60
Identities = 109/135 (80%), Positives = 125/135 (91%), Gaps = 2/135 (1%)
Frame = +1
Query: 272 GIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVG 331
GIRKTFE+CNKVR+I+ESYCV + ERDVSMDSGFKEELRKLMG +QVKVPVVFVKGRLVG
Sbjct: 1 GIRKTFEECNKVRSIVESYCVHVVERDVSMDSGFKEELRKLMGTKQVKVPVVFVKGRLVG 180
Query: 332 GVDEVVKLEDEEKLGVLLEGIP-KAVGGCEGCGGVRFVMCVECNGSCKVLD-EDKKKTVK 389
G +E+VKLE+E KLGVL EGIP KA+G CEGCGGVRFVMCVECNGSCKVLD E+ KKT++
Sbjct: 181 GAEEIVKLEEEGKLGVLFEGIPHKALGECEGCGGVRFVMCVECNGSCKVLDHENHKKTLR 360
Query: 390 CGKCNENGIMQCPIC 404
CG+CNENG++QCP+C
Sbjct: 361 CGQCNENGLIQCPMC 405
>TC220784 similar to GB|AAD03427.1|4115916|F3H7 F3H7.9 {Arabidopsis
thaliana;} , partial (43%)
Length = 777
Score = 182 bits (462), Expect = 2e-46
Identities = 98/178 (55%), Positives = 124/178 (69%), Gaps = 5/178 (2%)
Frame = +2
Query: 232 TPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYC 291
TP T K L + +L FEK CPP GE VVIYTT+LRG+R TFE CN VRA +E +
Sbjct: 107 TPPT--GSKLLLTPTLADRFEKICPPNGEKRVVIYTTSLRGVRTTFEACNAVRAALEGFG 280
Query: 292 VQMRERDVSMDSGFKEELRKLMGAEQVKVPV-VFVKGRLVGGVDEVVKLEDEEKLGVLLE 350
V + ERDVSM SGF+EELR L+ +QV VP VFVKG +GG DE++K+ +E LG LL+
Sbjct: 281 VVICERDVSMHSGFREELRTLLKGKQVMVPPRVFVKGLYIGGADEMLKVAEEGLLGDLLD 460
Query: 351 GIP-KAVGG-CEGCGGVRFVMCVECNGSCKVLDEDKKKT--VKCGKCNENGIMQCPIC 404
G+P K VG C GCG +RF+ C CNGSCK L +++ +T VKC CNENG++ CP+C
Sbjct: 461 GLPRKKVGAVCVGCGDLRFLPCFNCNGSCKTLVKEQGRTVVVKCTHCNENGLVLCPLC 634
>TC212818 weakly similar to UP|Q9FNG2 (Q9FNG2) Emb|CAB81925.1, partial (42%)
Length = 854
Score = 163 bits (413), Expect = 1e-40
Identities = 76/159 (47%), Positives = 110/159 (68%), Gaps = 4/159 (2%)
Frame = +1
Query: 251 FEKKCPPGGEN-SVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEEL 309
F++ CPPGG N S+++YTT+LRGIRKTF++CN +R ++ S+ + ERDVS+ F+EEL
Sbjct: 43 FKELCPPGGSNHSIILYTTSLRGIRKTFQECNTIRFLLRSFKIMYHERDVSLHLEFREEL 222
Query: 310 RKLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIP--KAVGGCEGCGGVRF 367
K++G + + P +F+KGR +GG DEVV L + LG LEG P + C GC +RF
Sbjct: 223 WKILGGKVIP-PKLFIKGRYIGGADEVVGLHEMGWLGKFLEGTPTHSSDSPCSGCANMRF 399
Query: 368 VMCVECNGSCKVL-DEDKKKTVKCGKCNENGIMQCPICC 405
+C C GSCKV D + + V+C +CNENG+++CP+CC
Sbjct: 400 AICSNCCGSCKVFTDNNDECFVRCSQCNENGLVKCPVCC 516
>TC229619
Length = 1334
Score = 144 bits (364), Expect = 5e-35
Identities = 84/218 (38%), Positives = 127/218 (57%), Gaps = 3/218 (1%)
Frame = +2
Query: 191 DTLQCSSRRKSQSPLFDPELVASYEKELSEEEEQVKRMVWATPKTRRARKSLDSLSLIQM 250
D+++ S R K L L S +K L E E + + + ++R DS S I +
Sbjct: 434 DSMKSSIRGKMVKKLCT--LFESSKKPLPEPESESESF-----SSSKSRSKTDSDSCITI 592
Query: 251 FEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELR 310
+ PG E+ +V+Y T+LRGIR+TFEDCN VR I++ + V + ERDVSMD ++EEL+
Sbjct: 593 LFRL--PGTEDRIVVYLTSLRGIRRTFEDCNAVRMILKGFRVWVDERDVSMDLSYREELQ 766
Query: 311 KLMGAEQVKVPVVFVKGRLVGGVDEVVKL-EDEEKLGVLLEGIPKAVGG--CEGCGGVRF 367
+ G V +P VF+ + +GG D + L E + ++LEG+PK G C+ CG RF
Sbjct: 767 HVPGDHHVALPQVFIPRKYIGGADVIKHLFESGDLPKMILEGLPKLKPGFVCDNCGDPRF 946
Query: 368 VMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
V C G+ KV DED+ + +C + +ENG+++CP CC
Sbjct: 947 VPCENSRGTRKVFDEDEGELKRCLESHENGLLRCPYCC 1060
>TC224162
Length = 831
Score = 137 bits (345), Expect = 9e-33
Identities = 67/150 (44%), Positives = 98/150 (64%), Gaps = 3/150 (2%)
Frame = +1
Query: 258 GGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE- 316
G E+ +V+Y T+LRGIRKT+EDC VR I+ + V + RD+SMDS +++EL+ L+G +
Sbjct: 121 GTEDRIVLYCTSLRGIRKTYEDCCSVRMILRGFRVAVVFRDISMDSSYRKELKDLLGGKA 300
Query: 317 QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGG--CEGCGGVRFVMCVECN 374
V +P VF++GR VG +E+ L + +L LLEG P G C+ CG RFV C C+
Sbjct: 301 AVTLPQVFIRGRYVGNAEEMKHLNESGELARLLEGFPTQDPGFVCDNCGDARFVPCPNCS 480
Query: 375 GSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
GS KV + + +C +CNENG+++CP C
Sbjct: 481 GSRKVFEHEDGGLRRCPECNENGLIRCPGC 570
>TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644
At2g41330/F13H10.12 {Arabidopsis thaliana;} , partial
(24%)
Length = 583
Score = 133 bits (334), Expect = 2e-31
Identities = 67/149 (44%), Positives = 98/149 (64%), Gaps = 2/149 (1%)
Frame = +1
Query: 256 PPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGA 315
PPG + VV+Y T+LR +R+TF+DC VR+I+ V + ERDVS+D F++EL ++G
Sbjct: 139 PPGLDQGVVVYFTSLRVVRRTFDDCRAVRSILRGLRVAVDERDVSIDDRFRDELHAVLGC 318
Query: 316 E-QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA-VGGCEGCGGVRFVMCVEC 373
+ +P VFV G VGG D+V +L + +L L+E +P++ + C+ CGG RFV+C EC
Sbjct: 319 RGNLALPRVFVGGVYVGGADDVRQLHESGELHRLIERLPRSNLNACDSCGGFRFVVCDEC 498
Query: 374 NGSCKVLDEDKKKTVKCGKCNENGIMQCP 402
NGS KV E K + C CN NG+++CP
Sbjct: 499 NGSHKVFAE-KNGFLCCSSCNANGLIRCP 582
>BE659157 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidopsis
thaliana}, partial (53%)
Length = 727
Score = 111 bits (277), Expect(2) = 2e-31
Identities = 55/131 (41%), Positives = 81/131 (60%), Gaps = 11/131 (8%)
Frame = -3
Query: 286 IIESYCVQMRERDVSMDSGFKEELRKLMGAEQ---------VKVPVVFVKGRLVGGVDEV 336
I + V + ERD+SMD+ +++EL + E V +P VF++GR VGG D +
Sbjct: 398 IFRGFRVWVDERDISMDANYRKELMSALFGENNNNNNKKGHVALPQVFIRGRHVGGADVI 219
Query: 337 VKLEDEEKLGVLLEGIPKAVGG--CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCN 394
+ + +L +LEG+P+ GG CE CG VRFV C C+GS KV DED++ +C +CN
Sbjct: 218 KHMWEVGELEKVLEGLPRTKGGFVCESCGDVRFVPCGNCSGSRKVFDEDEEVLKRCLECN 39
Query: 395 ENGIMQCPICC 405
ENG+++CP CC
Sbjct: 38 ENGLIRCPNCC 6
Score = 42.7 bits (99), Expect(2) = 2e-31
Identities = 23/47 (48%), Positives = 31/47 (65%)
Frame = -1
Query: 238 ARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVR 284
A KSLDS ++ PG E+ +V+Y T+LRGIR+T+EDC VR
Sbjct: 520 ASKSLDSPAIRL-------PGTEDRIVLYFTSLRGIRRTYEDCYAVR 401
>TC221571 similar to GB|AAO63337.1|28950827|BT005273 At5g58530 {Arabidopsis
thaliana;} , partial (30%)
Length = 965
Score = 121 bits (303), Expect = 6e-28
Identities = 67/149 (44%), Positives = 93/149 (61%), Gaps = 4/149 (2%)
Frame = +2
Query: 260 ENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQ-V 318
E VV+Y T+LR +R TFEDC VR+I+ + V + ERDVSMDSGF ELR++ G + +
Sbjct: 209 EQRVVVYFTSLRVVRATFEDCKTVRSILRGFRVALDERDVSMDSGFLSELRRVTGHKSGL 388
Query: 319 KVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA---VGGCEGCGGVRFVMCVECNG 375
+P VF+ GR VGG +E+ L + +L LLEG+P + C C RFV+C EC+G
Sbjct: 389 TLPRVFINGRYVGGAEELRWLHESGELKKLLEGLPAVDSHLRVCHVCDDHRFVLCGECSG 568
Query: 376 SCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
+ KV E K C CNE+G+++C C
Sbjct: 569 ARKVYAE-KGGFKTCTACNESGLIRCISC 652
>BI967423 similar to GP|14532460|gb AT5g13810/MAC12_24 {Arabidopsis
thaliana}, partial (45%)
Length = 683
Score = 120 bits (302), Expect = 8e-28
Identities = 66/155 (42%), Positives = 95/155 (60%), Gaps = 12/155 (7%)
Frame = -2
Query: 240 KSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDV 299
KS+DS I++ PG E+ +V+Y T+LRGIR+T+EDC VR I + V + ERD+
Sbjct: 490 KSIDSSPAIRL------PGTEDRIVLYFTSLRGIRRTYEDCYAVRMIFRGFRVWVDERDI 329
Query: 300 SMDSGFKEELRKLMGAE----------QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
SMD+ +++EL ++ E V +P VF++GR VGG D + + + +L +L
Sbjct: 328 SMDANYRKELMSVLFGENNNNNNKKKGHVALPQVFIRGRHVGGADVIKHMWEVGELEKVL 149
Query: 350 EGIPKAVGG--CEGCGGVRFVMCVECNGSCKVLDE 382
EG+P+ GG CE CG VRFV C C+GS KV DE
Sbjct: 148 EGLPRTKGGFVCESCGDVRFVPCGNCSGSRKVFDE 44
>BE804031 weakly similar to GP|15624060|dbj B1088C09.21 {Oryza sativa
(japonica cultivar-group)}, partial (37%)
Length = 413
Score = 115 bits (288), Expect = 4e-26
Identities = 60/137 (43%), Positives = 87/137 (62%), Gaps = 3/137 (2%)
Frame = +3
Query: 268 TTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGA-EQVKVPVVFVK 326
T+LR +R+TF+DC VR+I+ V + ERDVS+D F++EL ++G + +P VFV
Sbjct: 6 TSLRVVRRTFDDCRAVRSILRGLRVAVDERDVSIDDRFRDELHAVLGCRSNLALPRVFVG 185
Query: 327 GRLVGGVDEVVKLEDEEKLGVLLEGIPKA--VGGCEGCGGVRFVMCVECNGSCKVLDEDK 384
G VGG D+V +L + +L L+E +P++ C+ CGG RFV+C ECNGS KV E K
Sbjct: 186 GIYVGGADDVRQLHESGELHRLIERLPRSNQNNACDSCGGFRFVVCDECNGSHKVFTE-K 362
Query: 385 KKTVKCGKCNENGIMQC 401
C CN NG+++C
Sbjct: 363 NGFRSCSSCNANGLIRC 413
>TC232470 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g58530
{Arabidopsis thaliana;} , partial (48%)
Length = 625
Score = 114 bits (284), Expect = 1e-25
Identities = 63/146 (43%), Positives = 85/146 (58%), Gaps = 6/146 (4%)
Frame = +3
Query: 265 IYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQV----KV 320
+Y T+LR +R TFE V + VQ+ ERDVSMDSGF EL ++MG ++ +
Sbjct: 3 VYYTSLRVVRPTFEXXKSVXSXXRXXRVQIDERDVSMDSGFTAELNRIMGRPELGPSPSL 182
Query: 321 PVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA--VGGCEGCGGVRFVMCVECNGSCK 378
P VF+ GR VGG +E+ +L + +L +L +P C C G RFV+C ECNGS K
Sbjct: 183 PRVFIAGRYVGGAEELRQLNEVGELKKILLDLPAVDPTAECHVCAGHRFVLCDECNGSRK 362
Query: 379 VLDEDKKKTVKCGKCNENGIMQCPIC 404
V E K C CNENG+++CP C
Sbjct: 363 VYTE-KTGFKTCNACNENGLVKCPSC 437
>TC231232 similar to UP|Q9LPI8 (Q9LPI8) F6N18.14, partial (10%)
Length = 397
Score = 78.2 bits (191), Expect = 6e-15
Identities = 41/93 (44%), Positives = 54/93 (57%), Gaps = 4/93 (4%)
Frame = +2
Query: 316 EQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEG--IPKAVGG--CEGCGGVRFVMCV 371
E V +P VFVKGR VGG + L +LG +L + + +G C GCGG RFV C
Sbjct: 8 EGVALPRVFVKGRYVGGWRSLGDLXYTGRLGRILNATRVERGIGRQTCGGCGGARFVPCF 187
Query: 372 ECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
+C GSCK+L + +C CNENG++ CP C
Sbjct: 188 DCAGSCKLLHRE-----RCPNCNENGLVHCPAC 271
>CO979084
Length = 382
Score = 73.9 bits (180), Expect = 1e-13
Identities = 39/87 (44%), Positives = 51/87 (57%), Gaps = 2/87 (2%)
Frame = -2
Query: 320 VPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKA--VGGCEGCGGVRFVMCVECNGSC 377
+P VF+ GR VGG +EV +L + +L +L +P C C G RFV+C ECNGS
Sbjct: 375 LPRVFIAGRYVGGAEEVRQLNEVGELKKILMDLPAVDPTTECHVCAGHRFVLCDECNGSR 196
Query: 378 KVLDEDKKKTVKCGKCNENGIMQCPIC 404
KV E K C CNENG+++CP C
Sbjct: 195 KVYAE-KTGFKTCNACNENGLVKCPSC 118
>BQ741711
Length = 437
Score = 61.2 bits (147), Expect = 8e-10
Identities = 24/58 (41%), Positives = 43/58 (73%)
Frame = +1
Query: 256 PPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLM 313
P ++VV+Y T+LR +R+T++DC VR+I+ + + + ERDVS+D F+EEL++++
Sbjct: 250 PDSDRSAVVVYYTSLRVVRRTYDDCRAVRSILRGFAIAIDERDVSVDERFREELQRIL 423
>TC220542 similar to GB|AAL69460.1|18491271|AY074644 At2g41330/F13H10.12
{Arabidopsis thaliana;} , partial (12%)
Length = 560
Score = 61.2 bits (147), Expect = 8e-10
Identities = 23/47 (48%), Positives = 31/47 (65%)
Frame = +3
Query: 359 CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
C+ CG RFV C CNGS KV + ++ +C CNENG+++CP CC
Sbjct: 6 CDNCGDARFVPCPNCNGSRKVFEHEEGGLRRCPDCNENGLIRCPGCC 146
>BU090296 homologue to GP|14532460|gb| AT5g13810/MAC12_24 {Arabidopsis
thaliana}, partial (15%)
Length = 395
Score = 45.4 bits (106), Expect = 5e-05
Identities = 23/47 (48%), Positives = 32/47 (67%)
Frame = +2
Query: 240 KSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAI 286
KS+DS I++ PG E+ +V+Y T+LRGIR+T+EDC VR I
Sbjct: 269 KSIDSSPAIRL------PGTEDRIVLYFTSLRGIRRTYEDCYAVRMI 391
>AW830807 weakly similar to PIR|T48082|T480 glutaredoxin-like protein -
Arabidopsis thaliana, partial (80%)
Length = 378
Score = 38.9 bits (89), Expect = 0.004
Identities = 25/84 (29%), Positives = 37/84 (43%), Gaps = 13/84 (15%)
Frame = +2
Query: 301 MDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEG--------- 351
M SG + E + + VP VF+ + +GG DEV+KL + KL LL G
Sbjct: 83 MQSGHQVERALIQLGCKPSVPAVFIGQQFIGGADEVIKLNVQNKLAQLLLGAKAIFIWSR 262
Query: 352 ----IPKAVGGCEGCGGVRFVMCV 371
+P + C+ G F+ V
Sbjct: 263 *LCQVPNSSSSCDQIRGSYFLKAV 334
>BU091135 homologue to GP|10241626|emb Contains similarity to F1N 19.7 {Oryza
sativa (indica cultivar-group)}, partial (3%)
Length = 421
Score = 38.1 bits (87), Expect = 0.007
Identities = 32/86 (37%), Positives = 39/86 (45%), Gaps = 9/86 (10%)
Frame = +1
Query: 35 SLTSSTYGALKLDKDNDQPVIAA-------KPREEPEAITTINAWELMEGLEDGVPISNQ 87
SLTSSTYG + P+ + EEPE I NAWELMEGL+ P
Sbjct: 166 SLTSSTYGHSC*QGE*ATPLHSP*RFQHLHLDSEEPEVI---NAWELMEGLKRECPFQTT 336
Query: 88 PKKSPKSSSP--FLRGFMNSDTRSPL 111
P KS + P F+ N + PL
Sbjct: 337 P*KSQVHTFPSGFIXTTQNPNQNPPL 414
Score = 29.3 bits (64), Expect = 3.3
Identities = 22/87 (25%), Positives = 36/87 (41%), Gaps = 3/87 (3%)
Frame = +3
Query: 15 QEHVIIDNCGGGGRYLSHVVSLTSSTYGALKLDKDNDQPVIAAKPREEPEAITTINAWEL 74
QE VI N G Y++H + + + + ++ + K P +++
Sbjct: 117 QEEVIATN---DGSYVNHSIPHFLHLWTLMLTRRMSNTTSLPVKIPTSPPGFRRTRSYQR 287
Query: 75 ME---GLEDGVPISNQPKKSPKSSSPF 98
+ G E+GVPISN P K P F
Sbjct: 288 LGTHGGSEEGVPISNNPMKIPSPHLSF 368
>BQ297489
Length = 420
Score = 36.6 bits (83), Expect = 0.021
Identities = 21/61 (34%), Positives = 32/61 (52%), Gaps = 12/61 (19%)
Frame = +3
Query: 32 HVVSLTSSTYGAL--------KLDKDNDQPVIAAKPREEPE----AITTINAWELMEGLE 79
H+VSLTS+TYG+L + N + + +PE + IN WELM+GL+
Sbjct: 111 HLVSLTSTTYGSLLPIDQKDSNFTQKNQPHITKTSNQTDPEHSLSPDSVINTWELMDGLD 290
Query: 80 D 80
+
Sbjct: 291 E 293
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.134 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,095,455
Number of Sequences: 63676
Number of extensions: 234624
Number of successful extensions: 1175
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 1116
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1134
length of query: 405
length of database: 12,639,632
effective HSP length: 99
effective length of query: 306
effective length of database: 6,335,708
effective search space: 1938726648
effective search space used: 1938726648
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0010.16


