

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
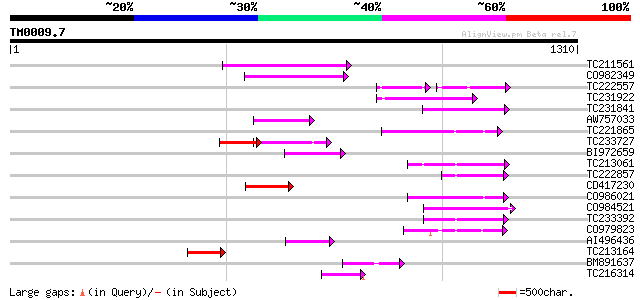
Query= TM0009.7
(1310 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 220 4e-57
CO982349 156 6e-38
TC222557 74 6e-22
TC231922 98 3e-20
TC231841 96 1e-19
AW757033 95 2e-19
TC221865 95 2e-19
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 93 9e-19
BI972659 85 2e-16
TC213061 84 5e-16
TC222857 80 6e-15
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 79 2e-14
CO986021 78 3e-14
CO984521 76 1e-13
TC233392 74 3e-13
CO979823 73 7e-13
AI496436 72 2e-12
TC213164 71 3e-12
BM891637 71 4e-12
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 69 1e-11
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 220 bits (560), Expect = 4e-57
Identities = 113/299 (37%), Positives = 177/299 (58%), Gaps = 1/299 (0%)
Frame = +1
Query: 491 LLANEVADFLSKKSEGGLMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMKSCISSA 550
L+ANEV D + ++ L+ KVD KAYD V W FL+ M+ M+F KWI W++ C+ SA
Sbjct: 13 LIANEVIDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEECVKSA 192
Query: 551 SLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRN- 609
S+++LVNGSPT F ++GLRQGDPL+P LFN+ GL+ L+ + + E Y + N
Sbjct: 193 SISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYLVGANG 372
Query: 610 LTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVER 669
+ I+ L +ADDT+ F E + +E + +L +F S L++N KS V +
Sbjct: 373 VPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTDQWKQE 552
Query: 670 LAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITK 729
A+ S+ P YLG P+G N RR W+ ++ K + K + + +S GR+ + K
Sbjct: 553 AANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTLIK 732
Query: 730 SVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQICLNRKAGGL 788
SVL SIP++ F+ P+ V+ ++ K+ F+WG +KKI+WI W+++CL ++ GG+
Sbjct: 733 SVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKERGGI 909
>CO982349
Length = 795
Score = 156 bits (394), Expect = 6e-38
Identities = 78/241 (32%), Positives = 138/241 (56%), Gaps = 1/241 (0%)
Frame = +3
Query: 542 WMKSCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENY 601
W++ C+ SAS++ILVN SP + FS ++GLRQGDPL+PLLFN+ GL+ L+ + + + +
Sbjct: 69 WIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRF 248
Query: 602 CGVRISRNLT-INHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGC 660
+ +N ++ L +ADDT+ F E ++ + +L F ASGLK+N +S
Sbjct: 249 NSFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGAI 428
Query: 661 NVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRIVFWNPMLEKLKCKTRSYNSKFMS 720
+ A S+ ++P SYLG P+ NPR +P++ K + K + + +S
Sbjct: 429 WKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHIS 608
Query: 721 LSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQIC 780
L GR+ + ++L ++P++ F+AP V+ ++ + F+WG +++I W+ W+ +C
Sbjct: 609 LGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETVC 788
Query: 781 L 781
L
Sbjct: 789 L 791
>TC222557
Length = 1002
Score = 73.6 bits (179), Expect(2) = 6e-22
Identities = 50/178 (28%), Positives = 80/178 (44%), Gaps = 6/178 (3%)
Frame = +1
Query: 986 MWQVPKSIAKLLPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHI 1045
+WQ+ +P K F W+ ++I T+ NL R + LN + C FCN +E +H+
Sbjct: 439 LWQLK------IPLKATTFAWRLIKERIPTKGNLWRRRVQLN-NLMCPFCNRQEEEASHL 597
Query: 1046 FLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWDFLPK--ISD----KSLWSLIPYAL 1099
F +C I LW E W + I +++ IS ++ W AL
Sbjct: 598 FFNCPRILPLWW-----ESLAWTKTVGVFSPIPRQNYMQHTLISKGKHLQARWKCWWVAL 762
Query: 1100 SWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFESIRF 1157
+WSIW RN I F +KA N S++ D + + W +L + + + S N + F
Sbjct: 763 TWSIWRHRNRIVFLNKAFNGSKLMDDAIFLVWSWFRQLEKGFAQPYNYWSSNLSTAFF 936
Score = 50.4 bits (119), Expect(2) = 6e-22
Identities = 36/127 (28%), Positives = 60/127 (46%), Gaps = 3/127 (2%)
Frame = +3
Query: 848 QDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDPWV-SNVALCHLFPRLYAMSENKE 906
QD+ V++E ++ F+ + +G G +++FW D W+ L +PRLY +S ++
Sbjct: 6 QDLRSVIHEQ-AVHGLFQSAIDWKVGCGDKIRFWEDCWLMEQEPLRAKYPRLYNISCQQQ 182
Query: 907 ARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASIN--HIIPAAVGGDQLIWCANQ 964
I +G +S W W L +R F E + FL I+ HI + D +W
Sbjct: 183 KLILVMGFHSANVWEWKLEWRRHLFDNEVQAAASFLDDISWGHIDRWTL--DCWVWKPEP 356
Query: 965 TGLFSVK 971
G FS +
Sbjct: 357 NGQFSTR 377
>TC231922
Length = 803
Score = 97.8 bits (242), Expect = 3e-20
Identities = 61/237 (25%), Positives = 112/237 (46%), Gaps = 4/237 (1%)
Frame = -3
Query: 848 QDIWRVLNED--HSLSAGFKENLLISIGDGSRVKFWRDPWVS-NVALCHLFPRLYAMSEN 904
+D+ R+ N+ HS+ +N++ +G G ++KFW+D W+S L + +LY +S
Sbjct: 765 KDLRRLYNQPDFHSIH----QNMVWRVGCGDKIKFWQDSWLSEGCNLQQKYNQLYTISRQ 598
Query: 905 KEARIAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQ 964
I+++G +S+ W +R + F +E + F+ I+ I D + W A+
Sbjct: 597 *NLTISKMGKFSQNARSWEFKWRRRLFDYEYAMAVDFMNEISGISIQNQAHDSMFWKADS 418
Query: 965 TGLFSVKELCYWLEDHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIATRANLLARG 1023
+G++S K L + ++ + + L +PP+ A+F W+ D++ TR NL R
Sbjct: 417 SGVYSTKSAYRLLMPSISPAPSRRIFQILWHLKIPPRAAVFSWRLFLDRLPTRGNLSRRS 238
Query: 1024 IDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEW 1080
I + C C E H+F HC LW + R + +PA A +++
Sbjct: 237 IPIQ-DIMCPLCGCQHEEAGHLFFHCKMTKGLWWESMRWIRVIGALPADPASHFIQF 70
>TC231841
Length = 791
Score = 95.5 bits (236), Expect = 1e-19
Identities = 60/202 (29%), Positives = 100/202 (48%), Gaps = 2/202 (0%)
Frame = +3
Query: 955 GDQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKI 1013
GDQ +W A+ +G ++ K L +F Q V + + KL LP K+ +F W+ D++
Sbjct: 60 GDQWVWKADPSGQYTAKSAYGVLWGEMFEEQQDGVFEELWKLKLPSKITIFAWRLIRDRL 239
Query: 1014 ATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASL 1073
TR+NL + I+++ +C FC + +ES H+F HC I +W ++ L V P
Sbjct: 240 PTRSNLRRKQIEVD-DPRCPFCRSAEESAAHLFFHCSRIAPVWWESLSWVNLLGVFPNHP 416
Query: 1074 ADLILEWDFLPKISDK-SLWSLIPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGW 1132
L+ + + S W AL+W+IW +RNN+ F++ N ++I D + I
Sbjct: 417 RQHFLQHIYGVTAGMRASRWKWWWLALTWTIWKQRNNMIFSNGTFNANKILDEAIFLIWT 596
Query: 1133 WTHELLRQNSYSIDQLSRNFES 1154
W L + S +Q S N +
Sbjct: 597 WLTHLEKDFSIHYNQWSSNIRA 662
>AW757033
Length = 441
Score = 95.1 bits (235), Expect = 2e-19
Identities = 53/142 (37%), Positives = 79/142 (55%), Gaps = 1/142 (0%)
Frame = -2
Query: 564 FSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISR-NLTINHLLFADDTL 622
FS +GLRQGDPL+PLLFN+ GL+ L+ + + +Y + + ++ L +ADDT+
Sbjct: 431 FSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTI 252
Query: 623 LFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLP 682
F E + + +L F ASGLK+N KS +V + A S+ +LP
Sbjct: 251 FFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSLLSLP 72
Query: 683 ISYLGAPLGGNPRRIVFWNPML 704
SYLG P+G NPRR W+P++
Sbjct: 71 FSYLGIPIGANPRRRETWDPII 6
>TC221865
Length = 1045
Score = 95.1 bits (235), Expect = 2e-19
Identities = 69/284 (24%), Positives = 126/284 (44%), Gaps = 6/284 (2%)
Frame = -3
Query: 860 LSAGFKENLLISIGDGSRVKFWRDPWVSN-VALCHLFPRLYAMSENKEARIAEVGTYSRG 918
L+ + ++ +G G + +FW D V L +PR+Y +S ++ I +VG+++
Sbjct: 959 LNRVLQNEIVWKVGCGDKFRFWEDKLVGEGETLMRKYPRMYQISCQQQQLIQQVGSHTET 780
Query: 919 KWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLE 978
W W +R F E + FL I+ D +W + G +S + + ++
Sbjct: 779 TWEWKFQWRRPLFDNEVDTTIGFLEDISRFPIHRHLTDCWVWKSEPNGHYSTRSAYHLMQ 600
Query: 979 DHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNA 1037
+ I KL +P K A+F + D++ T+ NL R + LNG T CLFC
Sbjct: 599 HEAAKANTDPAFEEIWKLKIPAKAAIFA*RLVKDRLPTKNNLRRRQVQLNG-TLCLFCRN 423
Query: 1038 FDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWDF----LPKISDKSLWS 1093
++E +H+F C +W ++ A+ L+ F L + + W
Sbjct: 422 YEEEASHLFFSCTKTQPVWWESLSWINTSGAFSANPRHHFLQHSFGLLGLKQYNR*KCWW 243
Query: 1094 LIPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHEL 1137
+ AL+++IW RN F+++ N S++ + + + W EL
Sbjct: 242 V---ALTFTIWQHRNRTIFSNEPFNGSKMMEDAMFLVWSWFREL 120
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 92.8 bits (229), Expect = 9e-19
Identities = 44/97 (45%), Positives = 67/97 (68%), Gaps = 1/97 (1%)
Frame = -3
Query: 486 IADCILLANEVADFLSKKSEGG-LMLKVDLAKAYDQVDWEFLINMLHEMNFGDKWISWMK 544
I DC + +EV + L KK GG L LK+D+ KA+D +D +FLI++L++ + + +W++
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIR 655
Query: 545 SCISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLF 581
+ SA L+I VNG P FFS ++G+RQGDPLSP L+
Sbjct: 654 VILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
Score = 59.3 bits (142), Expect = 1e-08
Identities = 42/182 (23%), Positives = 83/182 (45%), Gaps = 1/182 (0%)
Frame = -2
Query: 563 FFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVRISRNLTINHLLFADDTL 622
F + KG + PL + ++ L FL E Y +LT +H++ ADD +
Sbjct: 601 FLLLSKGCKTRRSPIPLFVSPKMS*AGNYLC*FLKESCYL*SVPMGSLTRSHVMHADDIM 422
Query: 623 LFCENNELQLETLCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLP 682
+FC+ + L N + + G ++ KS++ ++ + ++ + +P
Sbjct: 421 IFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISDLLQYFHSDIP 242
Query: 683 ISYLGAPL-GGNPRRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMV 741
+Y G P+ G P R + + +K+KCK S+ +S+ GR+ + SV+ + L+
Sbjct: 241 FNYFGVPIFKGKPTRRHLY-CVADKIKCKLASWKGSILSIMGRIQLVNSVIHGMLLYSFS 65
Query: 742 VF 743
V+
Sbjct: 64 VY 59
>BI972659
Length = 453
Score = 84.7 bits (208), Expect = 2e-16
Identities = 42/140 (30%), Positives = 72/140 (51%)
Frame = +1
Query: 635 LCNVLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNP 694
L ++L F SGL++N KS+ Q A +P YLG P+
Sbjct: 34 LKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPFHYLGMPIAVKA 213
Query: 695 RRIVFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVE 754
+ W P++ K K K +N K++S+ GR+ + KSVL ++P++L+ FK P+ ++ ++
Sbjct: 214 SSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFKIPQRIVDKLV 393
Query: 755 KLFHAFIWGQEQGRKKINWI 774
L F+WG Q +I+W+
Sbjct: 394 SLQRTFMWGGNQHHNRISWV 453
>TC213061
Length = 823
Score = 83.6 bits (205), Expect = 5e-16
Identities = 58/237 (24%), Positives = 105/237 (43%), Gaps = 3/237 (1%)
Frame = +2
Query: 920 WGWSLTFRGQFFAWEEELYQQFLASIN-HIIPAAVGGDQLIWCANQTGLFSVKELCYWLE 978
W W +R F E E+ F+ + H I + + DQ +W A +G + + +
Sbjct: 20 WEWQFQWRRPLFDSEIEMLVAFIQEVEGHKIRSDIE-DQWVWAAESSGSYLARSAYRVIR 196
Query: 979 DHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNA 1037
+ + + + K + KL +P KV +F W+ D + TR NL + ++L+ C C +
Sbjct: 197 EGIPEEEQDREFKELWKLKVPMKVTMFAWRLLKDILPTRDNLRRKRVELHEYV-CPLCRS 373
Query: 1038 FDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWDF-LPKISDKSLWSLIP 1096
DES +H+F HC I +W ++ + + P + + + + W
Sbjct: 374 MDESASHLFFHCSKILPIWWESLSWVKLVGAFPHHPRHHFHQHSHEVYQGLQGNRWKWWW 553
Query: 1097 YALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRNFE 1153
AL+W+IW RN+I F++ N ++ D + I W L + +Q S N +
Sbjct: 554 LALTWTIWKHRNDIIFSNATFNAHKVMDDAVFLIWTWFRHLENDFTSHYNQWSSNLK 724
>TC222857
Length = 657
Score = 80.1 bits (196), Expect = 6e-15
Identities = 44/156 (28%), Positives = 80/156 (51%), Gaps = 1/156 (0%)
Frame = -3
Query: 997 LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLW 1056
+P K +F W+ D++ T+ANL AR + ++ T C FC +E+ +H+F+HC +W
Sbjct: 598 IPSKFLMFAWRLLWDRLPTKANLRARQVQISDLT-CPFCRRVEETASHMFIHCIKTQPIW 422
Query: 1057 CVIVNRERFLWVMPASLADLILEWDFLPKISDKS-LWSLIPYALSWSIWIERNNICFNHK 1115
+N +P S+ D +++ L + +S W A++WSIW RN+I F++
Sbjct: 421 WETMNWINMQGPLPWSITDHFMQFSSLKEAGIRSRRWQWWWMAVTWSIWQLRNSIVFSNA 242
Query: 1116 ALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRN 1151
+ +++ + + W H L + S +Q S N
Sbjct: 241 TFDGNKLVEDASFLLWTWLHNLEKDFSLHFNQWSSN 134
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 78.6 bits (192), Expect = 2e-14
Identities = 44/112 (39%), Positives = 69/112 (61%), Gaps = 1/112 (0%)
Frame = -3
Query: 546 CISSASLAILVNGSPTDFFSIEKGLRQGDPLSPLLFNLCLNGLSCLLNQFLGEENYCGVR 605
C+S+ ++++L NG D FS G+RQ DP++P LF LCL LS L+N + ++ ++
Sbjct: 668 CMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQ 489
Query: 606 ISR-NLTINHLLFADDTLLFCENNELQLETLCNVLITFLFASGLKVNNQKSE 656
++R L I+HL F DD +LF E N Q++ + L F +SG KVN K++
Sbjct: 488 VNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTK 333
>CO986021
Length = 900
Score = 77.8 bits (190), Expect = 3e-14
Identities = 62/237 (26%), Positives = 96/237 (40%), Gaps = 5/237 (2%)
Frame = -1
Query: 920 WGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGLFSVKELCYWLED 979
W W +R F E E F+ I+ I D ++W A+ TG++S K L
Sbjct: 891 WNWDFKWRRNLFDHENEQAIAFMDDISAISIHQQLQDSMLWKADPTGIYSTKSAYRLLLP 712
Query: 980 HLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIATRANLLARGIDLNGSTQCLFCNAF 1038
+ Q K + KL +PP+ LF W+ D+++ RANLL R + L C C
Sbjct: 711 NNRPGQHSSNFKILWKLKIPPRAELFSWRLFRDRLSIRANLLRRHVALQ-DIMCPLCGNH 535
Query: 1039 DESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEWD----FLPKISDKSLWSL 1094
E H+F HC LW +N R L A +++ + + LW +
Sbjct: 534 QEEAGHLFFHCRMTIGLWWESMNWTRTLGAFLDDPAAHFIQFSDGFGAQRNHNRRCLWWI 355
Query: 1095 IPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHELLRQNSYSIDQLSRN 1151
AL+ +IW RN++ F ++ D L W + + +Q S N
Sbjct: 354 ---ALTSTIWQHRNSLLFKGTPFQPPKVMDDALFHAWTWLKATEKGFNIPFNQWSTN 193
>CO984521
Length = 716
Score = 75.9 bits (185), Expect = 1e-13
Identities = 57/215 (26%), Positives = 93/215 (42%), Gaps = 2/215 (0%)
Frame = -2
Query: 956 DQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIA 1014
DQ W A +G +S K L + K + KL +P KVA+F W+ DK+
Sbjct: 646 DQWKWAAEPSGCYSTKSAYKALHHVTVGEEQDGKFKELWKLRVPLKVAIFAWRLIQDKLP 467
Query: 1015 TRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLA 1074
T+ANL + ++L C C + +E+ +H+F HC + LW + + V P
Sbjct: 466 TKANLRKKRVELQ-EYLCPLCRSVEETASHLFFHCSKVSPLWWESQSWVNMMGVFPYQPD 290
Query: 1075 DLILEWDFLPKIS-DKSLWSLIPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWW 1133
+ F + W +AL++SIW RN+I F++ + ++ D + + W
Sbjct: 289 QHFSQHIFGASVGLQGKRWQWWWFALTYSIWKHRNSIIFSNANFDAHKLMDDAVFILWTW 110
Query: 1134 THELLRQNSYSIDQLSRNFESIRFASKPSLLRAAS 1168
+ S +Q S N K + LR AS
Sbjct: 109 LRCFEKDFSLHFNQWSSNL-------KDAFLRTAS 26
>TC233392
Length = 1145
Score = 74.3 bits (181), Expect = 3e-13
Identities = 53/198 (26%), Positives = 85/198 (42%), Gaps = 2/198 (1%)
Frame = -1
Query: 956 DQLIWCANQTGLFSVKELCYWLEDHLFTHQMWQVPKSIAKL-LPPKVALFFWQAQNDKIA 1014
D +W L+S + L+ HL + + KL +P K ++F W+ D++
Sbjct: 656 DTWVWKHEPNELYSTRSAYKLLQGHLEGEDQDGALQDLWKLKIPAKASIFAWRLIRDRLP 477
Query: 1015 TRANLLARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLA 1074
T++NL R + L S C FC +E +HIFL C+ I LW R L V P +
Sbjct: 476 TKSNLHRRQVVLEDSL-CPFCRIREEDASHIFLECNKIRPLWWESQTWVRSLGVSPINKR 300
Query: 1075 DLILEW-DFLPKISDKSLWSLIPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWW 1133
L+ P + W AL+WS+W RN + F + N ++ + + W
Sbjct: 299 QHFLQHVHGTPGSKRYNRWKTWWIALTWSLWKHRNQVIFQNAQFNGIKLLEDAVFLHWTW 120
Query: 1134 THELLRQNSYSIDQLSRN 1151
+ + S +Q S N
Sbjct: 119 IRAMEKDFSMHYNQWSSN 66
>CO979823
Length = 853
Score = 73.2 bits (178), Expect = 7e-13
Identities = 59/251 (23%), Positives = 104/251 (40%), Gaps = 10/251 (3%)
Frame = -2
Query: 909 IAEVGTYSRGKWGWSLTFRGQFFAWEEELYQQFLASINHIIPAAVGGDQLIWCANQTGLF 968
I ++GT + W L +R E + FL I D+ +W G +
Sbjct: 843 IQQLGTITNSGWE*QLNWRRPXXDSEIAMADSFLGEITQQQIHPQREDKWLWKPEPGGHY 664
Query: 969 SVK--------ELCYWLEDHLFTHQMWQVPKSIAKLLPPKVALFFWQAQNDKIATRANLL 1020
S K EL ++D F ++W++ +P K A+F W+ D++ T++NL
Sbjct: 663 STKSGYHVLWGELTEEIQDADFA-EIWKLK------IPTKAAVFAWRLVRDRLPTKSNLR 505
Query: 1021 ARGIDLNGSTQCLFCNAFDESCNHIFLHCHSIWSLWCVIVNRERFLWVMPASLADLILEW 1080
R + + C CN +E H+F +C LW ++ MP + L++
Sbjct: 504 RRQVMVQDMV-CPLCNNIEEGAAHLFFNCTKTLPLWWESMSWVNLKTAMPQTPRQHFLQY 328
Query: 1081 --DFLPKISDKSLWSLIPYALSWSIWIERNNICFNHKALNISEIWDFHLLRIGWWTHELL 1138
D + K W AL+W+IW RN + F + + +++ + LL + W L
Sbjct: 327 GTDIADGLKSKR-WKCWWIALTWTIWQHRNKVVFQNATFHGNKVLEDALLLLWSWFKALE 151
Query: 1139 RQNSYSIDQLS 1149
+ + +Q S
Sbjct: 150 KDFNLHFNQWS 118
>AI496436
Length = 414
Score = 72.0 bits (175), Expect = 2e-12
Identities = 37/113 (32%), Positives = 61/113 (53%)
Frame = +1
Query: 638 VLITFLFASGLKVNNQKSELIGCNVQTVEVERLAHSFGWSVGTLPISYLGAPLGGNPRRI 697
+L F +SGLK+N KS+ + + A SV ++P YLG P+G N R
Sbjct: 4 ILRCFELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRHS 183
Query: 698 VFWNPMLEKLKCKTRSYNSKFMSLSGRLVITKSVLCSIPLFLMVVFKAPKGVI 750
W P++ K + K S+ K++S GR+ + SVL ++P++L+ F+ P V+
Sbjct: 184 DVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVV 342
>TC213164
Length = 446
Score = 71.2 bits (173), Expect = 3e-12
Identities = 33/87 (37%), Positives = 57/87 (64%)
Frame = +3
Query: 412 EDFFSSGRISRGMNAAFITLIPKKENASSLNHYRPISLIRSSYKLIAKVLATRLQKVMPK 471
++F+++G + NA F+ LIPK + SLN Y+PISLI YK++AK+L+ R +KV+P
Sbjct: 162 DEFYANGVFLKRSNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPT 341
Query: 472 LLSQNQFAFTKERQIADCILLANEVAD 498
++ + Q F + R +++ANE+ +
Sbjct: 342 IIDERQTVFMEGRHTLHNVVIANEIME 422
>BM891637
Length = 427
Score = 70.9 bits (172), Expect = 4e-12
Identities = 43/149 (28%), Positives = 67/149 (44%), Gaps = 5/149 (3%)
Frame = -1
Query: 769 KKINWIGWDQICLNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLWRFFVIERY---- 824
KKI WI W Q C + GGLG+ I+ N LL+KW W + + LW +I +Y
Sbjct: 418 KKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKYKGWR 239
Query: 825 NLDPDSLIWHMASTTPAMGSRAIQDIWRVLNEDHSLSAGFKENLLISIGDGSRVKFWRDP 884
LD ++ + + R +N+ S+ A +G G ++ FW D
Sbjct: 238 GLDQGPQKYYFSPWWADL---------RAINQHQSMIAA-SNQFCWKVGRGDQILFWEDS 89
Query: 885 WVSN-VALCHLFPRLYAMSENKEARIAEV 912
WV + L FP LY +S + +A++
Sbjct: 88 WVDDGTPLKDQFPELYRLSSQRNFIMADM 2
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 69.3 bits (168), Expect = 1e-11
Identities = 35/113 (30%), Positives = 55/113 (47%), Gaps = 11/113 (9%)
Frame = +2
Query: 721 LSGRLVITKSVLCSIPLFLMVVFKAPKGVILEVEKLFHAFIWGQEQGRKKINWIGWDQIC 780
+ GR+ + KSVL ++P+ + FK P+ ++ ++ L F+WG Q +I W+ W IC
Sbjct: 2 MGGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADIC 181
Query: 781 LNRKAGGLGLGFIEWRNRGLLLKWAWRFGREINSLW-----------RFFVIE 822
+ GGLG+ + N L +W W N LW FFV+E
Sbjct: 182 NPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARLAEQ*S*RVAFFVVE 340
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,050,644
Number of Sequences: 63676
Number of extensions: 1179009
Number of successful extensions: 9641
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 9341
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9580
length of query: 1310
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1201
effective length of database: 5,698,948
effective search space: 6844436548
effective search space used: 6844436548
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0009.7


