

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0006.8
(1142 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
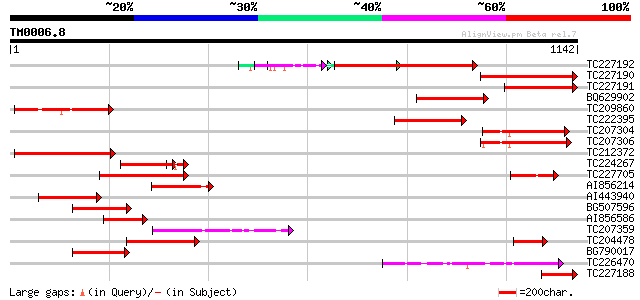
Score E
Sequences producing significant alignments: (bits) Value
TC227192 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, ... 447 e-126
TC227190 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, ... 364 e-100
TC227191 similar to GB|AAB84358.1|2618731|ATU49077 IAA21 {Arabid... 283 4e-76
BQ629902 similar to GP|1711205|gb|A IAA23 {Arabidopsis thaliana}... 245 9e-65
TC209860 similar to GB|AAB62404.1|2245390|ATU89296 auxin respons... 221 1e-57
TC222395 weakly similar to UP|Q6L8U3 (Q6L8U3) Auxin response fac... 216 6e-56
TC207304 similar to UP|Q6L8U2 (Q6L8U2) Auxin response factor 2, ... 213 4e-55
TC207306 similar to UP|Q6L8U2 (Q6L8U2) Auxin response factor 2, ... 212 7e-55
TC212372 similar to GB|AAN46837.1|24111405|BT001072 At5g62000/mt... 204 2e-52
TC224267 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, ... 203 3e-52
TC227705 similar to GB|AAM91657.1|22136676|AY133723 auxin respon... 194 2e-49
AI856214 193 4e-49
AI443940 homologue to GP|19352043|db auxin response factor 6b {O... 184 2e-46
BG507596 homologue to GP|9757747|dbj auxin response factor 4 {Ar... 177 3e-44
AI856586 168 1e-41
TC207359 similar to GB|BAB08972.1|9758525|AB024026 auxin respons... 161 2e-39
TC204478 similar to UP|ARFB_ARATH (Q94JM3) Auxin response factor... 150 4e-36
BG790017 similar to PIR|T08917|T08 auxin response factor 9 - Ara... 144 3e-34
TC226470 similar to UP|Q6L8U1 (Q6L8U1) Auxin response factor 3, ... 142 6e-34
TC227188 similar to GB|AAB84358.1|2618731|ATU49077 IAA21 {Arabid... 139 9e-33
>TC227192 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, partial
(14%)
Length = 875
Score = 447 bits (1151), Expect = e-126
Identities = 237/292 (81%), Positives = 253/292 (86%), Gaps = 4/292 (1%)
Frame = +3
Query: 654 ASLLQRQQLPQQTQLQQSSLHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKLQQ 713
ASLLQRQQ QQTQLQQS L LLQQN PQRAPQQPP M QQ PSEQQLHLQLLQKLQQ
Sbjct: 3 ASLLQRQQQQQQTQLQQSPLQLLQQNLPQRAPQQPPATQMLQQNPSEQQLHLQLLQKLQQ 182
Query: 714 HQQHQQ-LLSTSSPLLQPQLLQQQSTPQQNQQLTQL--PASQHQTQQLGNNAFSTEKVLN 770
QQHQQ LLSTS+PLL QLLQQQ+T QQNQQL Q P SQHQ QQLGNNAFSTEK+LN
Sbjct: 183 QQQHQQRLLSTSTPLLLSQLLQQQNTHQQNQQLPQPLPPISQHQPQQLGNNAFSTEKLLN 362
Query: 771 SNSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLL 830
SN+ SSSLMQ +L +NHPQNTQKSL ITRAPSTLT+GDAPSCSTSPSTNNCQ++P NLL
Sbjct: 363 SNNLSSSLMQSQKLSMNHPQNTQKSLTITRAPSTLTEGDAPSCSTSPSTNNCQVTPPNLL 542
Query: 831 KRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITDQMEAS 890
KRNQQ+PATL GSL+VE TSNLIQEL SK D QIK EF NVKG DQLKYKGTITDQ+EA
Sbjct: 543 KRNQQLPATLRGSLIVEPTSNLIQELHSKPDTQIKQEFLNVKGPDQLKYKGTITDQLEA- 719
Query: 891 SSGTSYCLD-PGNVQQSLPLSNFCMEGDVQSHSRSNLPFDSNLDGLTPDPVL 941
SSGTSYCLD PGNVQQ+LPLSNFCMEGDVQSH R++LPFDSNLDGLTPD +L
Sbjct: 720 SSGTSYCLDPPGNVQQNLPLSNFCMEGDVQSHPRNSLPFDSNLDGLTPDTML 875
Score = 76.3 bits (186), Expect = 7e-14
Identities = 60/158 (37%), Positives = 80/158 (49%), Gaps = 13/158 (8%)
Frame = +3
Query: 494 QQQQQQQQQQQQQQQLQLQHQQQQQQLQQQ-------QQQQQQQQLQQQQQQQLQQQQSQ 546
Q+QQQQQQ Q QQ LQL Q Q+ QQ QQ +QQL Q Q+LQQQQ
Sbjct: 15 QRQQQQQQTQLQQSPLQLLQQNLPQRAPQQPPATQMLQQNPSEQQLHLQLLQKLQQQQQH 194
Query: 547 QQQLQS------LLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQLGQQSQKAIMSNGVV 600
QQ+L S L Q+ Q + QQLP+P LP Q PQQLG A + ++
Sbjct: 195 QQRLLSTSTPLLLSQLLQQQNTHQQNQQLPQP--LPPISQHQPQQLG---NNAFSTEKLL 359
Query: 601 ASNQITNQLVLQQQQLLSGGISPQQSIHSASKTTFPLT 638
SN +++ L+ Q+ ++ PQ + S + T P T
Sbjct: 360 NSNNLSSSLMQSQKLSMN---HPQNTQKSLTITRAPST 464
Score = 51.6 bits (122), Expect = 2e-06
Identities = 77/278 (27%), Positives = 111/278 (39%), Gaps = 8/278 (2%)
Frame = +3
Query: 520 LQQQQQQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQL--QQPRQQQLPEPKNLPL 577
L Q+QQQQQQ QLQQ Q LQQ Q+ Q P Q+ Q P +QQL +L L
Sbjct: 9 LLQRQQQQQQTQLQQSPLQLLQQNLPQRAPQQP----PATQMLQQNPSEQQL----HLQL 164
Query: 578 PQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQQQLLSGGISP--QQSIHSASKTTF 635
Q+L QQ Q + S ++ S + Q QQ Q L + P Q F
Sbjct: 165 -LQKLQQQQQHQQRLLSTSTPLLLSQLLQQQNTHQQNQQLPQPLPPISQHQPQQLGNNAF 341
Query: 636 PLTSLPQESQFLQQIDQQASLLQRQQL----PQQTQLQQSSLHLLQQNQPQRAPQQPPVA 691
L L + +SL+Q Q+L PQ T Q SL + + P +
Sbjct: 342 STEKL------LNSNNLSSSLMQSQKLSMNHPQNT---QKSLTITRAPSTLTEGDAPSCS 494
Query: 692 TMSQQTPSEQQLHLQLLQKLQQHQQHQQLLSTSSPLLQPQLLQQQSTPQQNQQLTQLPAS 751
T +PS + L+++QQ L S + L Q+ + + Q+ Q +
Sbjct: 495 T----SPSTNNCQVTPPNLLKRNQQLPATLRGSLIVEPTSNLIQELHSKPDTQIKQEFLN 662
Query: 752 QHQTQQLGNNAFSTEKVLNSNSFSSSLMQPHQLPVNHP 789
QL T+++ S+ S L P + N P
Sbjct: 663 VKGPDQLKYKGTITDQLEASSGTSYCLDPPGNVQQNLP 776
Score = 43.9 bits (102), Expect = 4e-04
Identities = 54/209 (25%), Positives = 83/209 (38%), Gaps = 20/209 (9%)
Frame = +3
Query: 461 QAPALSAPSLQFNKPNLPNQINQ-------LHQSPVSWPQQQQQQQQQQQQQQQQLQLQH 513
Q L LQ + NLP + Q L Q+P Q Q+ QQQQQ Q +L
Sbjct: 33 QQTQLQQSPLQLLQQNLPQRAPQQPPATQMLQQNPSEQQLHLQLLQKLQQQQQHQQRLLS 212
Query: 514 QQQ----QQQLQQQQQQQQQQQL-------QQQQQQQLQQQQSQQQQLQSLLQIPMNQLQ 562
Q LQQQ QQ QQL Q Q QQL ++L + + + L
Sbjct: 213 TSTPLLLSQLLQQQNTHQQNQQLPQPLPPISQHQPQQLGNNAFSTEKLLNSNNL-SSSLM 389
Query: 563 QPRQQQLPEPKNL--PLPQQQLPQQLGQQSQKAIMSNGVVASNQITNQLVLQQQQLLSGG 620
Q ++ + P+N L + P L + + ++ + Q+T +L++ Q L
Sbjct: 390 QSQKLSMNHPQNTQKSLTITRAPSTLTEGDAPSCSTSPSTNNCQVTPPNLLKRNQQLPAT 569
Query: 621 ISPQQSIHSASKTTFPLTSLPQESQFLQQ 649
+ + S L S P ++Q Q+
Sbjct: 570 LRGSLIVEPTSNLIQELHSKP-DTQIKQE 653
>TC227190 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, partial (15%)
Length = 1255
Score = 364 bits (935), Expect = e-100
Identities = 181/198 (91%), Positives = 186/198 (93%), Gaps = 3/198 (1%)
Frame = +3
Query: 948 QKDLQNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNN--G 1005
QKDLQN+LSNYGGAPR+IETELSTADISSQSFGVPNMTF PGCS DVGINDTGVLNN G
Sbjct: 3 QKDLQNLLSNYGGAPREIETELSTADISSQSFGVPNMTFKPGCSSDVGINDTGVLNNNNG 182
Query: 1006 LRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKL 1065
LR NQT RMRTYTKVQKRGSVGRCIDVTRYKGYDELR+DLARMFGIEGQLEDP RTDWKL
Sbjct: 183 LRTNQTPRMRTYTKVQKRGSVGRCIDVTRYKGYDELRHDLARMFGIEGQLEDPLRTDWKL 362
Query: 1066 VYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDL-GNVPIPNQACSGTDS 1124
VYVDHENDILLVGDDPW+EFVSCVQSIKILS EVQQMSLDGDL GNVPIPNQA SGTDS
Sbjct: 363 VYVDHENDILLVGDDPWDEFVSCVQSIKILSSAEVQQMSLDGDLGGNVPIPNQAYSGTDS 542
Query: 1125 GNAWRGPYDDNSAASFNR 1142
GNAWRG Y+DNSAASFNR
Sbjct: 543 GNAWRGQYEDNSAASFNR 596
>TC227191 similar to GB|AAB84358.1|2618731|ATU49077 IAA21 {Arabidopsis
thaliana;} , partial (38%)
Length = 979
Score = 283 bits (723), Expect = 4e-76
Identities = 135/146 (92%), Positives = 141/146 (96%)
Frame = +3
Query: 997 NDTGVLNNGLRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLE 1056
ND GVLNNGL ANQTQRMRTYTKVQK GSVGRCIDVTRYKGYDELR+DLARMFGIEGQLE
Sbjct: 3 NDPGVLNNGLWANQTQRMRTYTKVQKCGSVGRCIDVTRYKGYDELRHDLARMFGIEGQLE 182
Query: 1057 DPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGNVPIPN 1116
DPQRT+WKLVYVDHENDILLVGDDPWEEFVSCVQSIKILS +EVQQMSLDGDLG+VP+PN
Sbjct: 183 DPQRTEWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSSSEVQQMSLDGDLGHVPVPN 362
Query: 1117 QACSGTDSGNAWRGPYDDNSAASFNR 1142
QACSGTD+GNAWRG YDDNSAASFNR
Sbjct: 363 QACSGTDNGNAWRGQYDDNSAASFNR 440
>BQ629902 similar to GP|1711205|gb|A IAA23 {Arabidopsis thaliana}, partial
(3%)
Length = 439
Score = 245 bits (625), Expect = 9e-65
Identities = 118/144 (81%), Positives = 128/144 (87%)
Frame = +3
Query: 820 NNCQISPSNLLKRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKY 879
NNCQ+SP NLLKRNQQ+PATLGG L+VE TSNLIQEL SK D QIK E NVKG DQLKY
Sbjct: 6 NNCQVSPPNLLKRNQQIPATLGGGLIVEPTSNLIQELHSKPDTQIKQELLNVKGPDQLKY 185
Query: 880 KGTITDQMEASSSGTSYCLDPGNVQQSLPLSNFCMEGDVQSHSRSNLPFDSNLDGLTPDP 939
KGTITD +EASSSGTSYCLDPGNVQQ+LPLSNFCME DVQSH R++LPFDSNLDGLTPD
Sbjct: 186 KGTITDSLEASSSGTSYCLDPGNVQQNLPLSNFCMERDVQSHPRNSLPFDSNLDGLTPDT 365
Query: 940 VLMRGYDSQKDLQNMLSNYGGAPR 963
+L+RGYDSQKDLQN+LSNY APR
Sbjct: 366 MLLRGYDSQKDLQNLLSNYASAPR 437
>TC209860 similar to GB|AAB62404.1|2245390|ATU89296 auxin response
transcription factor 3 {Arabidopsis thaliana;} , partial
(32%)
Length = 687
Score = 221 bits (563), Expect = 1e-57
Identities = 112/206 (54%), Positives = 138/206 (66%), Gaps = 7/206 (3%)
Frame = +3
Query: 11 NSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNL 70
+S T+ ELWHACAGP++SLP GS+VVYFPQGH EQ +P+ N+
Sbjct: 87 SSSSSSSSTVCLELWHACAGPMISLPKKGSVVVYFPQGHLEQHLHDFP-----LPASANI 251
Query: 71 PSKLICMLHNVALHADPETDEVYAQMTLQPVNK-------YDKDAILASDMGLKQNRQPT 123
PS + C + +V LHA+ +DEVY Q+ L P ++ +D D + ++ P
Sbjct: 252 PSHVFCRVLDVKLHAEEGSDEVYCQVVLVPESEQKLREGEFDADGEEEDAEAVMKSTTP- 428
Query: 124 EFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYR 183
FCKTLTASDTSTHGGFSVPRRAAE FPPLDYS Q P+QELVAKDLH W FRHIYR
Sbjct: 429 HMFCKTLTASDTSTHGGFSVPRRAAEDCFPPLDYSQQRPSQELVAKDLHGQEWRFRHIYR 608
Query: 184 GQPKRHLLTTGWSVFVSTKRLFAGDS 209
GQP+RHLLTTGWS FV+ K+L +GD+
Sbjct: 609 GQPRRHLLTTGWSAFVNKKKLVSGDA 686
>TC222395 weakly similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1,
partial (10%)
Length = 434
Score = 216 bits (549), Expect = 6e-56
Identities = 109/145 (75%), Positives = 123/145 (84%)
Frame = +3
Query: 775 SSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQISPSNLLKRNQ 834
SSSLMQ QL VN P NTQKSL TRAPSTLTDGDAPSCSTSPSTNNCQISP NL+KRNQ
Sbjct: 6 SSSLMQTQQLSVNQPHNTQKSLTNTRAPSTLTDGDAPSCSTSPSTNNCQISP-NLMKRNQ 182
Query: 835 QVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFSNVKGFDQLKYKGTITDQMEASSSGT 894
QV ATLGG VVE T++L+QEL SKS+MQIK+E +V+G DQLK+KGT+ DQMEA SSGT
Sbjct: 183 QVSATLGGPSVVEPTNHLMQELHSKSEMQIKHELPSVRGTDQLKFKGTVADQMEA-SSGT 359
Query: 895 SYCLDPGNVQQSLPLSNFCMEGDVQ 919
SYC+DP N+ Q+ PL NFCM+GDVQ
Sbjct: 360 SYCIDPNNIHQNFPLPNFCMDGDVQ 434
>TC207304 similar to UP|Q6L8U2 (Q6L8U2) Auxin response factor 2, partial (15%)
Length = 1006
Score = 213 bits (542), Expect = 4e-55
Identities = 115/185 (62%), Positives = 135/185 (72%), Gaps = 10/185 (5%)
Frame = +1
Query: 952 QNMLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVLNNG------ 1005
+ ML NY RD + ELS++ +S Q+FGVP+M FN S D I+D+ LN+G
Sbjct: 1 EGMLGNYX-XNRDAQQELSSSMVS-QTFGVPDMAFN---SIDSTIDDSNFLNSGPWAPPP 165
Query: 1006 ----LRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRT 1061
L Q QRMRTYTKV KRG+VGR ID+TRY GY+EL+ DLAR FGIEGQLED QR
Sbjct: 166 APPPLPPAQFQRMRTYTKVYKRGAVGRSIDITRYSGYEELKKDLARRFGIEGQLEDRQRI 345
Query: 1062 DWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGNVPIPNQACSG 1121
WKLVYVDHE+D+LLVGDDPWEEFV+CV+ IKILS EVQQMSLDGD GN + NQACS
Sbjct: 346 GWKLVYVDHESDVLLVGDDPWEEFVNCVRCIKILSPQEVQQMSLDGDFGNGGLQNQACSS 525
Query: 1122 TDSGN 1126
+D GN
Sbjct: 526 SDGGN 540
>TC207306 similar to UP|Q6L8U2 (Q6L8U2) Auxin response factor 2, partial (16%)
Length = 1125
Score = 212 bits (540), Expect = 7e-55
Identities = 119/200 (59%), Positives = 139/200 (69%), Gaps = 18/200 (9%)
Frame = +1
Query: 949 KDLQN------MLSNYGGAPRDIETELSTADISSQSFGVPNMTFNPGCSGDVGINDTGVL 1002
KDL N ML NY RD + E S++ +S Q+FGVP+M FN S D I+D+ L
Sbjct: 4 KDLSNNFSSEGMLGNYK-INRDAQQEPSSSMVS-QTFGVPDMAFN---SIDSTIDDSNFL 168
Query: 1003 NNG------------LRANQTQRMRTYTKVQKRGSVGRCIDVTRYKGYDELRNDLARMFG 1050
N+G L Q QRMRTYTKV KRG+VGR ID+TRY GY+EL+ DLAR FG
Sbjct: 169 NSGPWAPPPAPPLPPLPPAQFQRMRTYTKVYKRGAVGRSIDITRYSGYEELKQDLARRFG 348
Query: 1051 IEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLG 1110
IEGQLED QR WKLVYVDHE+D+LL+GDDPWEEFV+CV+ IKILS EVQQMSLDGD G
Sbjct: 349 IEGQLEDRQRIGWKLVYVDHESDVLLLGDDPWEEFVNCVRCIKILSPQEVQQMSLDGDFG 528
Query: 1111 NVPIPNQACSGTDSGNAWRG 1130
N +PNQACS +D GN G
Sbjct: 529 NGGLPNQACSSSDGGNT*NG 588
>TC212372 similar to GB|AAN46837.1|24111405|BT001072 At5g62000/mtg10_20
{Arabidopsis thaliana;} , partial (31%)
Length = 878
Score = 204 bits (519), Expect = 2e-52
Identities = 104/203 (51%), Positives = 132/203 (64%), Gaps = 1/203 (0%)
Frame = +1
Query: 11 NSGEGERKTINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETD-FIPSYPN 69
+S + ELWHACAGPLV++P V YFPQGH EQV AS + + +P Y +
Sbjct: 244 SSARDAEAALYRELWHACAGPLVTVPRERERVFYFPQGHIEQVEASTNQVAEQHMPVY-D 420
Query: 70 LPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKT 129
LP K++C + NV L A+P+TDEV+AQ+TL P D++A+ R FCKT
Sbjct: 421 LPPKILCRVINVMLKAEPDTDEVFAQVTLLPEPNQDENAVEKEGPPAAPPRFHVHSFCKT 600
Query: 130 LTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQPKRH 189
LTASDTSTHGGFSV RR A++ PPLD + QPP QELVAKDLH W FRHI+RGQP++H
Sbjct: 601 LTASDTSTHGGFSVLRRHADECLPPLDMTKQPPTQELVAKDLHGNXWRFRHIFRGQPRKH 780
Query: 190 LLTTGWSVFVSTKRLFAGDSVLF 212
LL +GW+VFV+ G + F
Sbjct: 781 LLQSGWNVFVTPXSXLLGMRLXF 849
>TC224267 similar to UP|Q6L8U3 (Q6L8U3) Auxin response factor 1, partial
(13%)
Length = 421
Score = 203 bits (517), Expect = 3e-52
Identities = 100/115 (86%), Positives = 108/115 (92%)
Frame = +3
Query: 223 GIRRANRQQPALSSSVISSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKF 282
GI+RANRQQPAL SSVISSDSMHIGILAAAAHAA+NNSPFTIFYNPRASPSEFVIPLAK+
Sbjct: 6 GIKRANRQQPALFSSVISSDSMHIGILAAAAHAASNNSPFTIFYNPRASPSEFVIPLAKY 185
Query: 283 NKAMYTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGW 337
NKA++ QVSLGMRFRMMFETEESGVRR+MGTITGI+DLD RWK+SQWRN GW
Sbjct: 186 NKALFNQVSLGMRFRMMFETEESGVRRYMGTITGITDLDPVRWKNSQWRNXS-GW 347
Score = 57.8 bits (138), Expect = 3e-08
Identities = 30/48 (62%), Positives = 32/48 (66%), Gaps = 5/48 (10%)
Frame = +1
Query: 317 ISDLDSARWKSSQWR-----NLQVGWDESTAGERPSRVSIWEIEPVVT 359
+ L S W S R LQVGWDEST GERPSRVSIW+IEPVVT
Sbjct: 274 VQSLVSLTWILSDGRIHSGAXLQVGWDESTGGERPSRVSIWDIEPVVT 417
>TC227705 similar to GB|AAM91657.1|22136676|AY133723 auxin response factor 1
{Arabidopsis thaliana;} , partial (54%)
Length = 1772
Score = 194 bits (493), Expect = 2e-49
Identities = 94/179 (52%), Positives = 130/179 (72%), Gaps = 1/179 (0%)
Frame = +3
Query: 182 YRGQPKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISS 241
+ GQPKRHLLTTGWSVFVS+K+L AGD+ +F+R E +L +G+RR RQQ + SSVISS
Sbjct: 159 FPGQPKRHLLTTGWSVFVSSKKLAAGDAFIFLRGENGELRVGVRRVMRQQSNVPSSVISS 338
Query: 242 DSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMRFRMMFE 301
SMH+G+LA A+HA A + F++FY PR S SEF++ + K+ + ++S+GMRF+M FE
Sbjct: 339 HSMHLGVLATASHAIATGTLFSVFYKPRTSRSEFIVSVNKYLEVQSHKLSVGMRFKMRFE 518
Query: 302 TEESGVRRHMGTITGISD-LDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIEPVVT 359
+E RR GTI G+ D S+ W S+WR+L+V WDE ++ RP RVS WE+EP+V+
Sbjct: 519 GDEIPERRFSGTIVGVGDNKSSSVWPDSEWRSLKVQWDEPSSIMRPDRVSSWELEPLVS 695
Score = 84.3 bits (207), Expect = 3e-16
Identities = 41/97 (42%), Positives = 70/97 (71%), Gaps = 1/97 (1%)
Frame = +3
Query: 1009 NQTQRMRTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVY 1067
+Q++++R+ TKV +G +VGR +D+TR+ GY++L L MF I+ +L + W++VY
Sbjct: 1251 SQSKQIRSCTKVHMQGMAVGRAVDLTRFDGYEDLLRKLEDMFNIKTELCGSLKK-WQVVY 1427
Query: 1068 VDHENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMS 1104
D+E+D+++VGDDPW+EF S V+ I I + EV+++S
Sbjct: 1428 TDNEDDMMMVGDDPWDEFCSVVRKIFIYTAEEVKKLS 1538
>AI856214
Length = 464
Score = 193 bits (490), Expect = 4e-49
Identities = 94/124 (75%), Positives = 100/124 (79%)
Frame = -1
Query: 287 YTQVSLGMRFRMMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERP 346
YTQVSLGMRFRMMFETEESGVR +MGTITGISDLD RWKSSQWRN+QVGWDESTAGERP
Sbjct: 464 YTQVSLGMRFRMMFETEESGVRGYMGTITGISDLDPVRWKSSQWRNIQVGWDESTAGERP 285
Query: 347 SRVSIWEIEPVVTPFYICPPPFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMKDA 406
RVSIWEIEPVVTPFYICPPPFFRPKFPRQPGMP ++ P F +K +
Sbjct: 284 RRVSIWEIEPVVTPFYICPPPFFRPKFPRQPGMPGSKTS---------PLASSAFILKIS 132
Query: 407 SGSI 410
S SI
Sbjct: 131 SSSI 120
>AI443940 homologue to GP|19352043|db auxin response factor 6b {Oryza
sativa}, partial (14%)
Length = 391
Score = 184 bits (467), Expect = 2e-46
Identities = 87/128 (67%), Positives = 107/128 (82%), Gaps = 2/128 (1%)
Frame = +3
Query: 59 KETD-FIPSYPNLPSKLICMLHNVALHADPETDEVYAQMTLQPVNKYDK-DAILASDMGL 116
KE D IP+YP+LP +LIC LHN+ +HAD ETDEVYAQMTLQP+N ++ +A L +++G
Sbjct: 6 KEVDAHIPNYPSLPPQLICQLHNMTMHADVETDEVYAQMTLQPLNPQEQNEAYLPAELGT 185
Query: 117 KQNRQPTEFFCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTW 176
++QPT +FCKTLTASDTSTHGGFSVPRRAAEK+FPPLD+S QPPAQEL+A+DLH W
Sbjct: 186 A-SKQPTNYFCKTLTASDTSTHGGFSVPRRAAEKVFPPLDFSQQPPAQELIARDLHGNEW 362
Query: 177 SFRHIYRG 184
FRHI+RG
Sbjct: 363 KFRHIFRG 386
>BG507596 homologue to GP|9757747|dbj auxin response factor 4 {Arabidopsis
thaliana}, partial (15%)
Length = 433
Score = 177 bits (448), Expect = 3e-44
Identities = 84/120 (70%), Positives = 95/120 (79%)
Frame = +3
Query: 126 FCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQ 185
FCKTLTASDTSTHGGFSVPRRAAE FPPLDY Q P+QELVAKDLH W FRHIYRGQ
Sbjct: 72 FCKTLTASDTSTHGGFSVPRRAAEDCFPPLDYKKQRPSQELVAKDLHGVEWKFRHIYRGQ 251
Query: 186 PKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMH 245
P+RHLLTTGWS+FVS K L +GD+VLF+R E +L LGIRRA R + L S++ S S +
Sbjct: 252 PRRHLLTTGWSIFVSQKNLVSGDAVLFLRGENGELRLGIRRAARPRNGLPESIVGSQSYY 431
>AI856586
Length = 321
Score = 168 bits (426), Expect = 1e-41
Identities = 86/87 (98%), Positives = 86/87 (98%)
Frame = +1
Query: 190 LLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISSDSMHIGIL 249
LLTTGWSVFVSTKRLFAGDSVLFIRDEKQ LLLGIRRANRQQPALSSSVISSDSMHIGIL
Sbjct: 4 LLTTGWSVFVSTKRLFAGDSVLFIRDEKQHLLLGIRRANRQQPALSSSVISSDSMHIGIL 183
Query: 250 AAAAHAAANNSPFTIFYNPRASPSEFV 276
AAAAHAAANNSPFTIFYNPRASPSEFV
Sbjct: 184 AAAAHAAANNSPFTIFYNPRASPSEFV 264
>TC207359 similar to GB|BAB08972.1|9758525|AB024026 auxin responsive
transcription factor {Arabidopsis thaliana;} , partial
(25%)
Length = 1263
Score = 161 bits (407), Expect = 2e-39
Identities = 109/290 (37%), Positives = 145/290 (49%), Gaps = 5/290 (1%)
Frame = +2
Query: 288 TQVSLGMRFRMMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPS 347
T+VS+GMRFRM+FETEES VRR+MGTITGISDLD RW +S WR+++VGWDESTAGER
Sbjct: 2 TRVSVGMRFRMLFETEESSVRRYMGTITGISDLDPVRWPNSHWRSVKVGWDESTAGERQP 181
Query: 348 RVSIWEIEPVVT-PFY--ICPPPFFRPKFPRQPGMPDDESDIENAFKRAMPWLGDDFGMK 404
RVS+WEIEP+ T P Y + P RP P D + N + WL G +
Sbjct: 182 RVSLWEIEPLTTFPMYPSLFPLRLKRPWHPGTSSFHDGRDEATN----GLMWLRGGPGDQ 349
Query: 405 DASGSIFPGLSLVQWMSMQQNNQFSAAQSGCFPPMLSPNTLHSNLGTDD--PSKLLSFQA 462
+ F G L+ WM + + A + + L NLG+ D ++++FQ
Sbjct: 350 ALNSLNFQGSGLLPWMQQRMDPTLLANDHNQQYQAMFASGL-QNLGSGDLMRQQIMNFQQ 526
Query: 463 PALSAPSLQFNKPNLPNQINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQ 522
P Q PN P Q+ Q PQ QQ Q Q Q+ + Q L Q
Sbjct: 527 PFNYLQ--QSGNPNPPLQLQQ--------PQAIQQSVSSNNILQPQAQVMAENLSQHLLQ 676
Query: 523 QQQQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQLQQPRQQQLPEP 572
+ ++ Q QQQQQ Q+ Q + + +QL Q + LP P
Sbjct: 677 KSHNNREDQTQQQQQ--------QRHTYQDTVLLQSDQLHQRQHSGLPSP 802
Score = 33.9 bits (76), Expect = 0.42
Identities = 57/213 (26%), Positives = 78/213 (35%), Gaps = 25/213 (11%)
Frame = +2
Query: 479 NQINQLHQSPVSWPQQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQ 538
N +N + W QQ+ QQ Q Q L +QQ QQ
Sbjct: 356 NSLNFQGSGLLPWMQQRMDPTLLANDHNQQYQAMFASGLQNLGSGDLMRQQIMNFQQPFN 535
Query: 539 QLQQQQSQQQQLQSLLQIPMNQLQQPRQ-QQLPEPKNLPLPQQQ-LPQQLGQQSQKAIMS 596
LQQ + LQ LQQP+ QQ N+ PQ Q + + L Q + +
Sbjct: 536 YLQQSGNPNPPLQ---------LQQPQAIQQSVSSNNILQPQAQVMAENLSQHLLQKSHN 688
Query: 597 NGVVASNQITNQ-------LVLQQQQL----LSGGISPQQSIHS--ASKTTFP------- 636
N + Q Q ++LQ QL SG SP S S FP
Sbjct: 689 NREDQTQQQQQQRHTYQDTVLLQSDQLHQRQHSGLPSPSYSKPDFLDSSMKFPASVSPGQ 868
Query: 637 --LTSL-PQESQFLQQIDQQASLLQRQQLPQQT 666
L SL P+ S L + + + + +QLPQQ+
Sbjct: 869 NMLGSLCPEGSGNLLNLSRSSQSMLTEQLPQQS 967
Score = 32.7 bits (73), Expect = 0.93
Identities = 52/217 (23%), Positives = 85/217 (38%), Gaps = 3/217 (1%)
Frame = +2
Query: 612 QQQQLLSGGISPQQSIHSASKTTFPLTSLPQESQFLQQIDQQASLLQRQQLPQQTQLQQS 671
Q Q + + G+ Q++ S + + Q +LQQ LQ QQ PQ Q S
Sbjct: 443 QYQAMFASGL---QNLGSGDLMRQQIMNFQQPFNYLQQSGNPNPPLQLQQ-PQAIQQSVS 610
Query: 672 SLHLLQQNQPQRAPQQPPVATMSQQTPSEQQLHLQLLQKLQQHQQHQQLLSTSSPLLQPQ 731
S ++LQ PQ +A + L LLQK +++ Q Q
Sbjct: 611 SNNILQ-------PQAQVMA---------ENLSQHLLQKSHNNREDQT-----------Q 709
Query: 732 LLQQQSTPQQNQQLTQL-PASQHQTQQLGNNAFSTEKVLNSNSFSSSLMQPHQLPVNH-- 788
QQQ Q+ L Q Q Q L + ++S L+S+ + + P Q +
Sbjct: 710 QQQQQRHTYQDTVLLQSDQLHQRQHSGLPSPSYSKPDFLDSSMKFPASVSPGQNMLGSLC 889
Query: 789 PQNTQKSLAITRAPSTLTDGDAPSCSTSPSTNNCQIS 825
P+ + L ++R+ ++ P S +P QI+
Sbjct: 890 PEGSGNLLNLSRSSQSMLTEQLPQQSWAPKFTPLQIN 1000
>TC204478 similar to UP|ARFB_ARATH (Q94JM3) Auxin response factor 2
(ARF1-binding protein) (ARF1-BP), partial (37%)
Length = 1975
Score = 150 bits (378), Expect = 4e-36
Identities = 72/149 (48%), Positives = 99/149 (66%), Gaps = 3/149 (2%)
Frame = +3
Query: 236 SSVISSDSMHIGILAAAAHAAANNSPFTIFYNPRASPSEFVIPLAKFNKAMYTQVSLGMR 295
SSVISS SMH+G+LA A HA + FT++Y PR SP+EF++P ++ +++ ++GMR
Sbjct: 15 SSVISSHSMHLGVLATAWHAILTGTMFTVYYKPRTSPAEFIVPYDQYMESLKNNYTIGMR 194
Query: 296 FRMMFETEESGVRRHMGTITGISDLDSARWKSSQWRNLQVGWDESTAGERPSRVSIWEIE 355
F+M FE EE+ +R GTI GI D D+ RW S+WR+L+V WDE++ RP RVS W+IE
Sbjct: 195 FKMRFEGEEAPEQRFTGTIVGIEDADTKRWPKSKWRSLKVRWDETSNIPRPERVSQWKIE 374
Query: 356 PVVTPFYICPPPFFRPKFPRQ---PGMPD 381
P + P + P P RPK PR P PD
Sbjct: 375 PALAPPALNPLPMPRPKRPRSNVVPSSPD 461
Score = 73.6 bits (179), Expect = 5e-13
Identities = 33/70 (47%), Positives = 51/70 (72%), Gaps = 1/70 (1%)
Frame = +3
Query: 1015 RTYTKVQKRG-SVGRCIDVTRYKGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHEND 1073
R+ TKV K+G ++GR +D+T++ Y EL +L ++F G+L PQ+ DW +VY D+E D
Sbjct: 1407 RSCTKVHKKGIALGRSVDLTKFSDYGELITELDQLFEFGGELTSPQK-DWLIVYTDNEGD 1583
Query: 1074 ILLVGDDPWE 1083
++LVGDDPW+
Sbjct: 1584 MMLVGDDPWQ 1613
>BG790017 similar to PIR|T08917|T08 auxin response factor 9 - Arabidopsis
thaliana, partial (21%)
Length = 440
Score = 144 bits (362), Expect = 3e-34
Identities = 70/116 (60%), Positives = 85/116 (72%)
Frame = +1
Query: 126 FCKTLTASDTSTHGGFSVPRRAAEKIFPPLDYSMQPPAQELVAKDLHDTTWSFRHIYRGQ 185
FCK LTASDTSTHGGFSV R+ A + P LD S P QELVAKDL W F+HI+RGQ
Sbjct: 88 FCKVLTASDTSTHGGFSVLRKHATECLPALDMSKSTPTQELVAKDLQGFEWRFKHIFRGQ 267
Query: 186 PKRHLLTTGWSVFVSTKRLFAGDSVLFIRDEKQQLLLGIRRANRQQPALSSSVISS 241
P+RHLLTTGWS FV++KRL AGD+ +F+R +L +G+RR Q ++ SSVISS
Sbjct: 268 PRRHLLTTGWSTFVTSKRLVAGDTFVFLRGNNGELRVGVRRIAPLQSSMPSSVISS 435
>TC226470 similar to UP|Q6L8U1 (Q6L8U1) Auxin response factor 3, partial (30%)
Length = 1359
Score = 142 bits (359), Expect = 6e-34
Identities = 120/381 (31%), Positives = 188/381 (48%), Gaps = 17/381 (4%)
Frame = +1
Query: 751 SQHQT-QQLGNNAFSTEKVLNSNSFSSSLMQPHQLPVNHPQNTQKSLAITRAPSTLTDGD 809
+QH QQ + ++V++ SS++ Q V+ PQ+ + +A S+L
Sbjct: 4 NQHNLLQQQQQSQLQQQQVVDHQQISSAVSTMSQF-VSAPQSQSPPM---QAISSLGHQQ 171
Query: 810 APSCSTSPSTNNCQISPSNLLKRNQQVPATLGGSLVVESTSNLIQELQSKSDMQIKNEFS 869
+ S S +SP + ++ GS + TS+L+ +S S + +++ +
Sbjct: 172 SFSDSNGNPVTTAVVSPLH----------SILGSFPQDDTSHLLNLPRSTSWVPVQHSTA 321
Query: 870 NVKGFDQLKYKGTITDQMEASSSGTSYCLDPGNVQQSLPLSNFCMEGDVQ--------SH 921
K D + SSG S C+ P Q P S G +
Sbjct: 322 WPSS------KRVAVDPL--FSSGASQCVLPQVEQLGQPQSTMAQNGIALPPFPGRECTI 477
Query: 922 SRSNLPFDSNLDGLTPDPVLMRGYDSQKDLQNMLSNYGGAPRDIETELSTADISSQSF-- 979
SN P + L G+ +P + ++ L+ + SN ++ T S ++
Sbjct: 478 EGSNDPQNHLLFGVNIEPSSLLMHNGMSSLKGVSSN---------SDSPTIPFQSSNYLN 630
Query: 980 -GVPNMTFNPGCSGDVGINDTGVLN---NGLRANQTQRMRTYTKVQKRGSVGRCIDVTRY 1035
VP+ + NPG + ++G ++G L NG + N T + T+ KV K GS GR +D+T++
Sbjct: 631 TTVPDSSLNPGMTHNIG--ESGFLQTPENGGQGNPTNK--TFVKVYKSGSFGRSLDITKF 798
Query: 1036 KGYDELRNDLARMFGIEGQLEDPQRTDWKLVYVDHENDILLVGDDPWEEFVSCVQSIKIL 1095
Y ELR++LARMFG+EG+LEDP R+ W+LV+VD END+LL+GD PW EFV+ V IKIL
Sbjct: 799 TSYPELRSELARMFGLEGELEDPVRSGWQLVFVDQENDVLLLGDGPWPEFVNSVGCIKIL 978
Query: 1096 SYTEVQQMSLDG--DLGNVPI 1114
S EVQQM +G L +VPI
Sbjct: 979 SPQEVQQMGNNGLELLNSVPI 1041
Score = 37.4 bits (85), Expect = 0.038
Identities = 41/122 (33%), Positives = 51/122 (41%)
Frame = +1
Query: 526 QQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQLQQPRQQQLPEPKNLPLPQQQLPQQ 585
Q Q LQQQQQ QLQQQQ Q S M+Q Q Q P P Q +
Sbjct: 1 QNQHNLLQQQQQSQLQQQQVVDHQQISSAVSTMSQFVSAPQSQSP-------PMQAI-SS 156
Query: 586 LGQQSQKAIMSNGVVASNQITNQLVLQQQQLLSGGISPQQSIHSASKTTFPLTSLPQESQ 645
LG Q Q SNG N +T +V +L G PQ T L +LP+ +
Sbjct: 157 LGHQ-QSFSDSNG----NPVTTAVVSPLHSIL--GSFPQDD-------TSHLLNLPRSTS 294
Query: 646 FL 647
++
Sbjct: 295 WV 300
Score = 32.3 bits (72), Expect = 1.2
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 2/57 (3%)
Frame = +1
Query: 506 QQQLQLQHQQQQQQLQQQQ--QQQQQQQLQQQQQQQLQQQQSQQQQLQSLLQIPMNQ 560
Q Q L QQQQ QLQQQQ QQ Q + QSQ +Q++ + Q
Sbjct: 1 QNQHNLLQQQQQSQLQQQQVVDHQQISSAVSTMSQFVSAPQSQSPPMQAISSLGHQQ 171
Score = 31.6 bits (70), Expect = 2.1
Identities = 19/55 (34%), Positives = 22/55 (39%)
Frame = +1
Query: 493 QQQQQQQQQQQQQQQQLQLQHQQQQQQLQQQQQQQQQQQLQQQQQQQLQQQQSQQ 547
Q QQQQQ Q QQQ + HQQ + Q Q Q Q + QQ
Sbjct: 7 QHNLLQQQQQSQLQQQQVVDHQQISSAVSTMSQFVSAPQSQSPPMQAISSLGHQQ 171
>TC227188 similar to GB|AAB84358.1|2618731|ATU49077 IAA21 {Arabidopsis
thaliana;} , partial (19%)
Length = 784
Score = 139 bits (349), Expect = 9e-33
Identities = 64/72 (88%), Positives = 69/72 (94%)
Frame = +1
Query: 1071 ENDILLVGDDPWEEFVSCVQSIKILSYTEVQQMSLDGDLGNVPIPNQACSGTDSGNAWRG 1130
ENDILLVGDDPWEEFVSCVQSIKILS EVQ+MSLDGDLG+VP+PNQACSGTD+GNAWRG
Sbjct: 1 ENDILLVGDDPWEEFVSCVQSIKILSSAEVQKMSLDGDLGHVPVPNQACSGTDNGNAWRG 180
Query: 1131 PYDDNSAASFNR 1142
Y+DNSAASFNR
Sbjct: 181 QYEDNSAASFNR 216
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.128 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,152,268
Number of Sequences: 63676
Number of extensions: 750227
Number of successful extensions: 15355
Number of sequences better than 10.0: 447
Number of HSP's better than 10.0 without gapping: 5135
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8554
length of query: 1142
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1034
effective length of database: 5,762,624
effective search space: 5958553216
effective search space used: 5958553216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0006.8


