

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
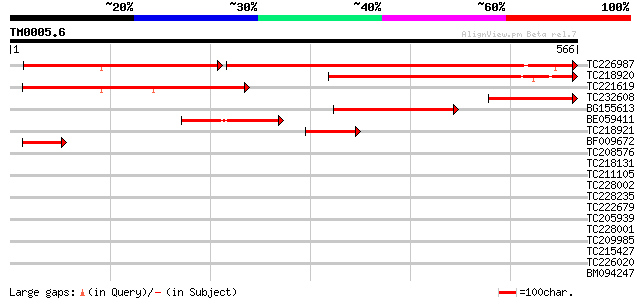
Query= TM0005.6
(566 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC226987 similar to UP|Q9FEV9 (Q9FEV9) Microtubule-associated pr... 342 e-151
TC218920 similar to GB|AAP37732.1|30725420|BT008373 At1g14690 {A... 188 6e-48
TC221619 similar to GB|AAP37732.1|30725420|BT008373 At1g14690 {A... 173 2e-43
TC232608 147 9e-36
BG155613 134 1e-31
BE059411 similar to GP|20805034|db cytokinesis regulating protei... 86 3e-17
TC218921 weakly similar to GB|AAP37732.1|30725420|BT008373 At1g1... 52 7e-07
BF009672 51 1e-06
TC208576 weakly similar to UP|Q9UN50 (Q9UN50) ELL complex EAP30 ... 32 0.99
TC218131 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (8%) 31 1.3
TC211105 similar to UP|Q6K7P2 (Q6K7P2) Transcription factor PCF3... 31 1.7
TC228002 30 2.2
TC228235 homologue to UP|Q9M607 (Q9M607) Transcription factor, p... 30 2.2
TC222679 similar to UP|Q9LH58 (Q9LH58) GTP-binding protein-like,... 30 3.8
TC205939 weakly similar to UP|Q761Z8 (Q761Z8) BRI1-KD interactin... 30 3.8
TC228001 30 3.8
TC209985 similar to PIR|T52285|T52285 serine/threonine-specific ... 29 6.4
TC215427 similar to GB|AAN46773.1|24111299|BT001019 At3g52990/F8... 29 6.4
TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, ... 29 6.4
BM094247 similar to PIR|B84668|B846 ethylene-insensitive3-like1 ... 28 8.4
>TC226987 similar to UP|Q9FEV9 (Q9FEV9) Microtubule-associated protein
MAP65-1a, partial (97%)
Length = 2204
Score = 342 bits (878), Expect(2) = e-151
Identities = 180/356 (50%), Positives = 250/356 (69%), Gaps = 6/356 (1%)
Frame = +1
Query: 217 QSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVTRLI 276
+SISNDTLA L + +LK++K+QRL K+QEL L+DLW+L++T +E++ F+HVT +
Sbjct: 898 KSISNDTLARLAKTVLTLKEDKKQRLHKLQELASQLIDLWNLMDTHPEERRLFDHVTCNM 1077
Query: 277 SASVDEVSTHGCLSSDVIEQVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDS 336
SASVDEV+ G L+ D+IEQ E EV+RL+ LKAS+MK++ FK+Q ELEEI+ H++VD
Sbjct: 1078 SASVDEVTVPGALALDLIEQAEVEVERLDQLKASRMKEIAFKKQAELEEIFARAHIEVDP 1257
Query: 337 EAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVEKWRFAAEEEKWLD 396
+AAR+ + +LI+SGNI+ ++LL MD+QI +AKE+A SR++ILD+VEKW A EEE WL+
Sbjct: 1258 DAAREKIMALIDSGNIEPTELLADMDNQIAKAKEEALSRKDILDKVEKWMSACEEESWLE 1437
Query: 397 EYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWEMEKGIPFLYEKAP 456
+Y RDENRY+A RGAH NLKRAEKARI+V+KIP++V+ L K +AWE + G+ F Y+ P
Sbjct: 1438 DYNRDENRYNASRGAHINLKRAEKARILVNKIPALVDTLVAKTRAWEEDHGMSFTYDGVP 1617
Query: 457 LLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIV 516
LL L+EY + R REEEKR+ R+QK+ EQ EQE +FGSR + +P+ S +
Sbjct: 1618 LLAMLDEYAMLRHEREEEKRRMRDQKKHHEQRNTEQETIFGSRPSPARPVSSSKSGGP-- 1791
Query: 517 GTPNGRRMLTPSSRYGTSGGKERRESV-----RGNNI-IPVNYVALPKDDSVSRGS 566
NG TP+ R + K S R N + PVNYVA+ K+D+ S S
Sbjct: 1792 -RANGGANATPNRRLSLNAHKNGNRSTSKDGKRENRLSAPVNYVAISKEDAASHVS 1956
Score = 213 bits (541), Expect(2) = e-151
Identities = 112/202 (55%), Positives = 142/202 (69%), Gaps = 3/202 (1%)
Frame = +3
Query: 14 TCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAE 73
TC SLL +LQ IWDE+GESD RD MLLQLEQECLD+Y+RKV+ K +A+L Q L+DA+
Sbjct: 279 TCGSLLKKLQEIWDEVGESDEQRDKMLLQLEQECLDVYKRKVEQAAKSRAQLLQALSDAK 458
Query: 74 AELINLVSSLGESSSFS---RGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQ 130
EL L+S+LGE S GT+K+QLA I PVLE L +K+ER+KEFS ++SQI Q
Sbjct: 459 LELSTLLSALGEKSFAGIPENTSGTIKEQLAAIAPVLEQLWQQKEERIKEFSDVQSQIQQ 638
Query: 131 ICVEIAGYEQPKSSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELS 190
IC EIAG S VD+SDL+ KKL E +S LQ+LQ EK R KV +ST+ +L
Sbjct: 639 ICGEIAGNLNLNDVSPAVDESDLSLKKLDEYQSELQELQKEKSERLHKVLEFVSTVHDLC 818
Query: 191 AVMSIDFWKTLSEIHPSLGDSS 212
AV+ +DF+ T +E+HPSL DSS
Sbjct: 819 AVLGMDFFSTATEVHPSLNDSS 884
>TC218920 similar to GB|AAP37732.1|30725420|BT008373 At1g14690 {Arabidopsis
thaliana;} , partial (39%)
Length = 804
Score = 188 bits (477), Expect = 6e-48
Identities = 105/264 (39%), Positives = 163/264 (60%), Gaps = 16/264 (6%)
Frame = +2
Query: 319 RQNELEEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREI 378
+++ELEEI + H + + ++LI+SG +D S+LL +++ QI +AK++A SR+E+
Sbjct: 2 KRSELEEICKLTHTEPYTSTTADKASALIDSGLVDPSELLANIEAQIIKAKDEALSRKEV 181
Query: 379 LDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTK 438
DR++KW A EEE WLDEY +D+NRY A RGAH NLKRAE+ARI +SKIP++V+NL K
Sbjct: 182 TDRIDKWFAACEEENWLDEYNQDDNRYCAGRGAHINLKRAERARITISKIPAMVDNLINK 361
Query: 439 VKAWEMEKGIPFLYEKAPLLQSLEEYHVQRQLREEEKRKSREQKRLQEQHAVEQEALFGS 498
AWE EK FLY+ L+ L++Y + RQ +EE+KR+ R+ K++Q+ ++EA++GS
Sbjct: 362 TLAWEDEKKTHFLYDGVRLVSILDDYKLARQQKEEDKRRHRDLKKMQDLLLNQKEAMYGS 541
Query: 499 RSATKKPLGQSTTANTIVGTPNG----------------RRMLTPSSRYGTSGGKERRES 542
+ + +K T N+ NG +LTP S G G +
Sbjct: 542 KPSPRKNNSFRKT-NSYRANGNGSMPPTPRRNSLSGGTTSELLTPRSYSGRQNGYFK--E 712
Query: 543 VRGNNIIPVNYVALPKDDSVSRGS 566
+R + P+N+VA+ K+D++S S
Sbjct: 713 MRRLSTAPLNFVAISKEDTMSYAS 784
>TC221619 similar to GB|AAP37732.1|30725420|BT008373 At1g14690 {Arabidopsis
thaliana;} , partial (33%)
Length = 1044
Score = 173 bits (438), Expect = 2e-43
Identities = 93/235 (39%), Positives = 146/235 (61%), Gaps = 8/235 (3%)
Frame = +2
Query: 13 TTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADA 72
+TC +LL +L+ IW++IGE++ ++D ML++LE+ECL++YRRKVD KA Q +A
Sbjct: 137 STCTALLRELEQIWNDIGETEVEKDRMLMELERECLEVYRRKVDEAANTKARFHQTVAAK 316
Query: 73 EAELINLVSSLGESSSFS-----RGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQ 127
EAEL L+++LGE S + +LKQ+LA+I P +E+L+ KKDER+K+F +K+Q
Sbjct: 317 EAELATLMAALGEHDIHSPIKTEKRSASLKQKLASITPWVEELKKKKDERLKQFEDVKAQ 496
Query: 128 ISQICVEIAGYEQPK---SSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHIS 184
I +I EI G+ SS+ D DL+ ++L E ++HL+ LQ EK R QKV ++
Sbjct: 497 IEKISGEIFGFHSVNNALSSTTVEDDQDLSLRRLNEYQTHLRTLQKEKSDRLQKVLQCVN 676
Query: 185 TISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQ 239
+ L +V+ +DF +T+ ++HPSL + +ISN TL +K K+
Sbjct: 677 EVHSLCSVLGLDFGQTVGDVHPSLHGTQVEQSTNISNSTLGRSRAGHSQVKNRKE 841
>TC232608
Length = 435
Score = 147 bits (372), Expect = 9e-36
Identities = 76/89 (85%), Positives = 82/89 (91%), Gaps = 1/89 (1%)
Frame = +3
Query: 479 REQKRLQEQHAVEQEALFGSRSATKKPLGQSTTANTIVGTPNGRRML-TPSSRYGTSGGK 537
+EQKRLQEQHAVEQEALFGSRSATKKPLGQST ANTI+GTP GRRML TPS R+G SGGK
Sbjct: 18 QEQKRLQEQHAVEQEALFGSRSATKKPLGQSTHANTILGTPTGRRMLSTPSGRHGNSGGK 197
Query: 538 ERRESVRGNNIIPVNYVALPKDDSVSRGS 566
+RRES R N+IIPVNYVALPKDDSVSRG+
Sbjct: 198 DRRESGRVNSIIPVNYVALPKDDSVSRGN 284
>BG155613
Length = 381
Score = 134 bits (337), Expect = 1e-31
Identities = 66/125 (52%), Positives = 91/125 (72%)
Frame = +2
Query: 324 EEIYRGVHMDVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRREILDRVE 383
EEI R H+ + + A + IESG++D + +L+ ++ QI Q KE+A R+EIL++VE
Sbjct: 2 EEICRKTHLIPEIDNAVESAVDAIESGSVDPACILEQIELQISQVKEEAFGRKEILEKVE 181
Query: 384 KWRFAAEEEKWLDEYERDENRYSAVRGAHKNLKRAEKARIVVSKIPSIVENLTTKVKAWE 443
KW A +EE WL+EY RD+NRY+A RGAH LKRAEKAR +V+KIP++V+ LT+K AWE
Sbjct: 182 KWLAACDEESWLEEYNRDDNRYNAGRGAHLTLKRAEKARALVNKIPAMVDGLTSKTIAWE 361
Query: 444 MEKGI 448
EKGI
Sbjct: 362 KEKGI 376
>BE059411 similar to GP|20805034|db cytokinesis regulating protein - like
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 314
Score = 86.3 bits (212), Expect = 3e-17
Identities = 44/102 (43%), Positives = 72/102 (70%)
Frame = +1
Query: 172 KILRQQKVKSHISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYI 231
K R ++V+ H+ T++ L V+ DF +T++ IHPSL DS KG+ +S+SNDT+ L I
Sbjct: 1 KSSRLKQVQEHLCTLNSLCLVLGFDFKQTINGIHPSLVDS-KGS-KSVSNDTIQQLAVAI 174
Query: 232 HSLKQEKQQRLLKVQELTKFLVDLWDLIETPIDEQKAFNHVT 273
L++ K QR+ K+Q+L +++LW+L++TPI+EQ+ F +VT
Sbjct: 175 QELREVKLQRMQKLQDLATTMLELWNLMDTPIEEQQMFQNVT 300
>TC218921 weakly similar to GB|AAP37732.1|30725420|BT008373 At1g14690
{Arabidopsis thaliana;} , partial (9%)
Length = 276
Score = 52.0 bits (123), Expect = 7e-07
Identities = 26/55 (47%), Positives = 40/55 (72%)
Frame = +1
Query: 296 QVEAEVQRLNALKASKMKDLVFKRQNELEEIYRGVHMDVDSEAARQILTSLIESG 350
Q AEV RL LKAS+MK+LVFK++++LEEI + H++ D+ A + ++LI+ G
Sbjct: 97 QAFAEVDRLAKLKASRMKELVFKKRSKLEEICKLTHIEPDTSTAAEKASALIDFG 261
>BF009672
Length = 211
Score = 51.2 bits (121), Expect = 1e-06
Identities = 21/44 (47%), Positives = 35/44 (78%)
Frame = +2
Query: 13 TTCASLLSQLQMIWDEIGESDSDRDNMLLQLEQECLDIYRRKVD 56
+T +LL +L+ IW+++ E+ D+D M+++LE+ECLD+YRR VD
Sbjct: 41 STRTALLRELEQIWNDLVETVLDKDLMVMELERECLDVYRRTVD 172
>TC208576 weakly similar to UP|Q9UN50 (Q9UN50) ELL complex EAP30 subunit,
partial (33%)
Length = 475
Score = 31.6 bits (70), Expect = 0.99
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +2
Query: 96 LKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEI 135
+K+QLAT R LED K +++ +SQ ++C ++
Sbjct: 179 MKEQLATFRSQLEDFARKHKNDIRKNPAFRSQFHEMCAKV 298
>TC218131 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (8%)
Length = 1077
Score = 31.2 bits (69), Expect = 1.3
Identities = 29/124 (23%), Positives = 50/124 (39%), Gaps = 5/124 (4%)
Frame = +3
Query: 193 MSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTKFL 252
+ ++FW+T + I+PS+ P + S + E+ L +Q L +
Sbjct: 240 LRLEFWRTQNNIYPSVAMEHNNNPAAAS------------EILSERDHILRCMQRLERLE 383
Query: 253 VDLWDLIETPI-----DEQKAFNHVTRLISASVDEVSTHGCLSSDVIEQVEAEVQRLNAL 307
+L P E N + R+ S D T L + V++Q+E + L L
Sbjct: 384 KTFGELSHKPAGIPLEKEHMLTNSLDRIKSVEFDLEKTKRVLHATVMKQLEI-AELLENL 560
Query: 308 KASK 311
+ASK
Sbjct: 561 QASK 572
>TC211105 similar to UP|Q6K7P2 (Q6K7P2) Transcription factor PCF3-like,
partial (8%)
Length = 430
Score = 30.8 bits (68), Expect = 1.7
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Frame = +1
Query: 469 QLREEEKRKSREQKRLQEQHAVEQEALFGSRS---ATKKPLGQSTTANTIVGTPNGRRML 525
QL+EEEK KRL+EQ+ + ++ + ATKK G+ +G P R L
Sbjct: 25 QLKEEEKCLLEVNKRLREQYRIVRQRCLSDQDVEFATKKE-GEEVETELFIGRPERRMPL 201
>TC228002
Length = 805
Score = 30.4 bits (67), Expect = 2.2
Identities = 27/111 (24%), Positives = 54/111 (48%), Gaps = 3/111 (2%)
Frame = +1
Query: 143 SSSEEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLS 202
S EE+++S + LG ++S QDLQ+ + +VK + +SEL ++++ + +
Sbjct: 244 SKIEELNES--RSELLGRVQSLKQDLQSWRSKWDTQVKVYRDELSELKKTLNVEVDQLRA 417
Query: 203 E---IHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQEKQQRLLKVQELTK 250
E + +L + S+ N L ++ H+ QE K++E+ K
Sbjct: 418 EFQDLRTTLQQQQEDVTASLRNLGLQDVSDVKHAPSQE-----TKIEEIVK 555
>TC228235 homologue to UP|Q9M607 (Q9M607) Transcription factor, partial (29%)
Length = 855
Score = 30.4 bits (67), Expect = 2.2
Identities = 23/104 (22%), Positives = 43/104 (41%), Gaps = 21/104 (20%)
Frame = +2
Query: 456 PLLQSLEEYHVQRQLREEEKRKSREQKRLQEQ----------HAVEQEALFGSRSATKKP 505
P+ + ++YH + R+ + E+K ++ + HA+ + G R KK
Sbjct: 14 PVYVNAKQYHGILRRRQSRAKAELEKKVIKNRKPYLHESRHLHAMRRARGNGGRFLNKKK 193
Query: 506 L-----------GQSTTANTIVGTPNGRRMLTPSSRYGTSGGKE 538
L GQ+T AN +PN + + T + G+S +
Sbjct: 194 LENYNSDATSDIGQNTGANPSTNSPNTQHLFTNNENLGSSNASQ 325
>TC222679 similar to UP|Q9LH58 (Q9LH58) GTP-binding protein-like, partial
(18%)
Length = 653
Score = 29.6 bits (65), Expect = 3.8
Identities = 16/45 (35%), Positives = 24/45 (52%)
Frame = +3
Query: 333 DVDSEAARQILTSLIESGNIDLSDLLQSMDDQIRQAKEQAQSRRE 377
D+D A Q +E+ + LS +L D++ KE AQSRR+
Sbjct: 207 DLDLVALEQEAKDAVEAYSSSLSQILSIEDEEKSDRKESAQSRRK 341
>TC205939 weakly similar to UP|Q761Z8 (Q761Z8) BRI1-KD interacting protein
116 (Fragment), partial (46%)
Length = 1434
Score = 29.6 bits (65), Expect = 3.8
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +3
Query: 367 QAKEQAQSRREILDRVEKWRFAAEEEKWLDEYERDENRYSAVRGAHKNL 415
+ ++A+ RRE ++E+ A EEEK E + E +A++ KNL
Sbjct: 672 RCSDRAEKRREFYLKLEEKHRALEEEKNQYEARKKEEEDAAIKQLRKNL 818
>TC228001
Length = 843
Score = 29.6 bits (65), Expect = 3.8
Identities = 34/167 (20%), Positives = 73/167 (43%), Gaps = 4/167 (2%)
Frame = +1
Query: 86 SSSFSRGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICVEIAGYEQPKSSS 145
S+S S T+++ L + + D + K K ++ I +IA + +S
Sbjct: 157 SNSESNPSTTIQRNLLLLNSPIPSSMENPDSSLPLSIK-KENVTPITSKIAELNESRS-- 327
Query: 146 EEVDQSDLTFKKLGELKSHLQDLQNEKILRQQKVKSHISTISELSAVMSIDFWKTLSE-- 203
+ LG ++S QDLQ+ + +VK + +SEL ++++ + +E
Sbjct: 328 ----------ELLGRVQSLKQDLQSWRSKLDTQVKVYRDELSELKKTLNVEVDQLRAEFQ 477
Query: 204 -IHPSLGDSSKGTPQSISNDTLASLTGYIHSLKQE-KQQRLLKVQEL 248
+ +L + S+ N L + H+ QE K ++++K +++
Sbjct: 478 DLRTTLQQQQEDVTASLRNLGLQDASDVKHAPSQETKIEKIVKEEQV 618
>TC209985 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (38%)
Length = 1034
Score = 28.9 bits (63), Expect = 6.4
Identities = 15/49 (30%), Positives = 26/49 (52%), Gaps = 6/49 (12%)
Frame = +1
Query: 164 HLQDLQNEKILRQQ------KVKSHISTISELSAVMSIDFWKTLSEIHP 206
HL+ +N ++++ + K K + +ISEL ++S FW IHP
Sbjct: 541 HLRGHRNMRVIQPKYLGVLHKSKMEMQSISELLGLVSFTFWVNTFYIHP 687
>TC215427 similar to GB|AAN46773.1|24111299|BT001019 At3g52990/F8J2_160
{Arabidopsis thaliana;} , complete
Length = 2020
Score = 28.9 bits (63), Expect = 6.4
Identities = 27/108 (25%), Positives = 47/108 (43%), Gaps = 4/108 (3%)
Frame = +3
Query: 126 SQISQICVEIAGYEQPKSSSEEVDQSDLTFKK----LGELKSHLQDLQNEKILRQQKVKS 181
S + +IC E +E+V DL FKK +GE SHL+ + + + KVK+
Sbjct: 1194 SIVGKICAE----------AEKVHNQDLYFKKAVKYVGEPMSHLESIASSAVRAAIKVKA 1343
Query: 182 HISTISELSAVMSIDFWKTLSEIHPSLGDSSKGTPQSISNDTLASLTG 229
+ S + + +++ P++ S PQ +N + TG
Sbjct: 1344 SVIICFTSSGRAA----RLIAKYRPTMPVISVVIPQLKTNQLRWTFTG 1475
>TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial (93%)
Length = 1441
Score = 28.9 bits (63), Expect = 6.4
Identities = 18/38 (47%), Positives = 24/38 (62%), Gaps = 2/38 (5%)
Frame = +3
Query: 156 KKLGELKSHLQDLQNEKILRQ--QKVKSHISTISELSA 191
KKL + K HLQDL+NEK L++ QK K + E+ A
Sbjct: 1077 KKLAKEK-HLQDLENEKHLQELDQKAKLEAKPLKEILA 1187
>BM094247 similar to PIR|B84668|B846 ethylene-insensitive3-like1 (EIL1)
[imported] - Arabidopsis thaliana, partial (3%)
Length = 428
Score = 28.5 bits (62), Expect = 8.4
Identities = 27/114 (23%), Positives = 48/114 (41%), Gaps = 4/114 (3%)
Frame = +3
Query: 31 ESDSDRDNMLLQLEQECLDIYRRKVDATRKHKAELCQWLADAEAELINLVSSLGESSSFS 90
++D + L LEQE +D+Y K AT + ELI +S
Sbjct: 150 DADKSLTSKKLVLEQEEMDLYTAKNSATPLSSVD----------ELIEEISE-------- 275
Query: 91 RGKGTLKQQLATIRPVLEDLRSKKDERVKEFSKIKSQISQICV----EIAGYEQ 140
+++++ + +LE LR K+ E + +K + ++C E A YE+
Sbjct: 276 -----IQKKIKDEKVLLEGLRQKECEAAGKADDLKVKFDKLCESANGEFASYEK 422
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,181,899
Number of Sequences: 63676
Number of extensions: 225889
Number of successful extensions: 1111
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 1096
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1104
length of query: 566
length of database: 12,639,632
effective HSP length: 102
effective length of query: 464
effective length of database: 6,144,680
effective search space: 2851131520
effective search space used: 2851131520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0005.6


