

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
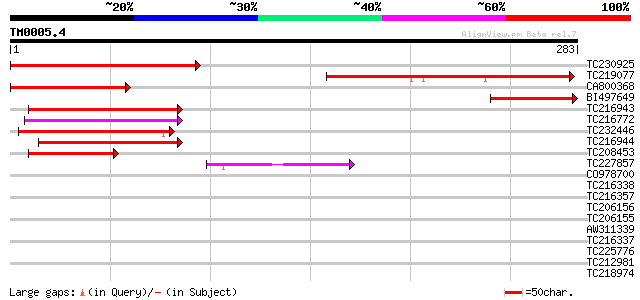
Score E
Sequences producing significant alignments: (bits) Value
TC230925 homologue to UP|Q9LHG0 (Q9LHG0) Arabidopsis thaliana ge... 189 2e-48
TC219077 homologue to UP|Q9W7B8 (Q9W7B8) Transcription factor Tc... 144 3e-35
CA800368 108 3e-24
BI497649 79 2e-15
TC216943 similar to UP|Q76FX1 (Q76FX1) CCAAT-box binding factor ... 60 1e-09
TC216772 homologue to UP|Q9SMP0 (Q9SMP0) Transcription factor Ha... 59 2e-09
TC232446 57 7e-09
TC216944 similar to UP|Q76FX1 (Q76FX1) CCAAT-box binding factor ... 52 2e-07
TC208453 similar to UP|O04033 (O04033) F7G19.16 protein, partial... 45 3e-05
TC227857 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarit... 41 5e-04
CO978700 40 0.001
TC216338 similar to UP|O19031 (O19031) Bactinecin 11, partial (7%) 39 0.002
TC216357 homologue to GB|AAM70547.1|21700847|AY124838 AT3g07590/... 39 0.003
TC206156 homologue to UP|Q9SY09 (Q9SY09) Small nuclear riboprote... 37 0.013
TC206155 homologue to GB|AAM70547.1|21700847|AY124838 AT3g07590/... 37 0.013
AW311339 similar to GP|18146720|dbj ribosomal protein S2 {Arabid... 36 0.017
TC216337 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-... 36 0.017
TC225776 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, par... 36 0.017
TC212981 similar to UP|BCT7_BOVIN (P19661) Bactenecin 7 precurso... 36 0.023
TC218974 weakly similar to UP|PCD7_MOUSE (Q9WTY1) Programmed cel... 35 0.029
>TC230925 homologue to UP|Q9LHG0 (Q9LHG0) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MQC3 (At3g12480/MQC3.32),
partial (31%)
Length = 491
Score = 189 bits (479), Expect = 2e-48
Identities = 92/95 (96%), Positives = 94/95 (98%)
Frame = +2
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK
Sbjct: 206 MRKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 385
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHG 95
TMNSLHLKHCVQSYNVFDFLRDVVS+VPDY HGHG
Sbjct: 386 TMNSLHLKHCVQSYNVFDFLRDVVSRVPDYGHGHG 490
>TC219077 homologue to UP|Q9W7B8 (Q9W7B8) Transcription factor Tcf3b
(Fragment), partial (6%)
Length = 718
Score = 144 bits (364), Expect = 3e-35
Identities = 79/138 (57%), Positives = 92/138 (66%), Gaps = 14/138 (10%)
Frame = +1
Query: 159 ETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSSETKELPKE-NAAVPA----ESAEFHNL 213
ET HQ E EPC SVQ +TNT + + + SE+KE+PKE N A P + N+
Sbjct: 1 ETYHQDAESEPCTSVQPSNPQNTNTSIAMDNGSESKEIPKEENIAAPVVPVVSTDSLRNI 180
Query: 214 DLNANTNENEDKKASTTAKQEIS---------EPPTESQHEEIPGWSLSDVDKMAIDSVQ 264
DLNA TNEN+DKKAS A S EPPTES+HEEIPGWSLSDVDKMAIDS+Q
Sbjct: 181 DLNAITNENDDKKASAEADASASVPEPDASLPEPPTESKHEEIPGWSLSDVDKMAIDSLQ 360
Query: 265 LANLGTHIEEDEEDYDEE 282
LA LG +EE+EEDYDEE
Sbjct: 361 LAQLGRPLEEEEEDYDEE 414
>CA800368
Length = 345
Score = 108 bits (270), Expect = 3e-24
Identities = 55/60 (91%), Positives = 57/60 (94%)
Frame = +3
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAV L SKALELFLQDLCDRTYE+TLQRGAK
Sbjct: 165 MRKKLDTRFPAARIKKIMQADEDVGKIALAVFFLXSKALELFLQDLCDRTYEVTLQRGAK 344
>BI497649
Length = 421
Score = 79.0 bits (193), Expect = 2e-15
Identities = 35/43 (81%), Positives = 40/43 (92%)
Frame = +1
Query: 241 ESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
E +HEEIPGWSLSDVDKM ID++QLANLG+ +EEDEEDYDEEG
Sbjct: 25 EIKHEEIPGWSLSDVDKMGIDTMQLANLGSRLEEDEEDYDEEG 153
>TC216943 similar to UP|Q76FX1 (Q76FX1) CCAAT-box binding factor HAP5
homolog, partial (60%)
Length = 1267
Score = 59.7 bits (143), Expect = 1e-09
Identities = 30/77 (38%), Positives = 48/77 (61%)
Frame = +2
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ +KA E+F+ +L R++ T + +T+ +
Sbjct: 566 PLARIKKIMKADEDVRMISAEAPVIFAKACEMFILELTLRSWIHTEENKRRTLQKNDIAA 745
Query: 70 CVQSYNVFDFLRDVVSK 86
+ +VFDFL D++ +
Sbjct: 746 AISRNDVFDFLVDIIPR 796
>TC216772 homologue to UP|Q9SMP0 (Q9SMP0) Transcription factor Hap5a, partial
(68%)
Length = 1040
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/79 (37%), Positives = 48/79 (59%)
Frame = +2
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+ P ARIKKIM+ADEDV I+ P+L +KA ELF+ +L R++ + +T+ +
Sbjct: 278 QLPLARIKKIMKADEDVRMISAEAPILFAKACELFILELTIRSWLHAEENKRRTLQKNDI 457
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 458 AAAITRTDIFDFLVDIVPR 514
>TC232446
Length = 622
Score = 57.4 bits (137), Expect = 7e-09
Identities = 28/79 (35%), Positives = 49/79 (61%), Gaps = 1/79 (1%)
Frame = +3
Query: 5 LDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNS 64
+ FP R+KKIM D+DV +++ LVS++ ELFLQ L +++ ++ +++ KT+N
Sbjct: 21 IQLEFPKGRVKKIMALDKDVKRVSSEALFLVSRSTELFLQFLAEKSAQVAIEKKRKTVNL 200
Query: 65 LHLKHCVQSYN-VFDFLRD 82
H++ V+ + DFL D
Sbjct: 201EHIREAVKRHQPTRDFLLD 257
>TC216944 similar to UP|Q76FX1 (Q76FX1) CCAAT-box binding factor HAP5
homolog, partial (42%)
Length = 864
Score = 52.4 bits (124), Expect = 2e-07
Identities = 26/72 (36%), Positives = 44/72 (61%)
Frame = +2
Query: 15 KKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCVQSY 74
KKIM+ADEDV I+ PV+ +KA E+F+ +L R++ T + +T+ + +
Sbjct: 2 KKIMKADEDVRMISAEAPVIFAKACEMFILELTLRSWIHTEENKRRTLQKNDIAAAISRN 181
Query: 75 NVFDFLRDVVSK 86
+VFDFL D++ +
Sbjct: 182DVFDFLVDIIPR 217
>TC208453 similar to UP|O04033 (O04033) F7G19.16 protein, partial (62%)
Length = 1093
Score = 45.4 bits (106), Expect = 3e-05
Identities = 22/45 (48%), Positives = 32/45 (70%)
Frame = +2
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEIT 54
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T
Sbjct: 401 PLARIKKIMKADEDVRMISAEAPVIFARACEMFILELTLRSWNHT 535
>TC227857 weakly similar to UP|Q8LA02 (Q8LA02) Contains similarity to
RNA-binding protein from Arabidopsis thaliana gi|2129727
and contains RNA recognition PF|00076 domain, partial
(37%)
Length = 995
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/79 (35%), Positives = 36/79 (45%), Gaps = 5/79 (6%)
Frame = +2
Query: 99 PSADDRT-----IPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG 153
PS +D T I K G D E RG + GRGRGRGRGRG GRG
Sbjct: 17 PSQEDATRNALKILSHGKDDGSDTGRGREYGGRGGLDR-----GRGRGRGRGRGRGMGRG 181
Query: 154 RTAVRETPHQAVEPEPCAS 172
R R+ + ++ + A+
Sbjct: 182 RFVERDVDEKVMDTDDYAT 238
>CO978700
Length = 728
Score = 40.0 bits (92), Expect = 0.001
Identities = 20/36 (55%), Positives = 23/36 (63%)
Frame = -1
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGR 154
DSD + G+ G GRGRGRG+GRGRGRGR
Sbjct: 269 DSDSSHQMGQ-------GGRGRGRGRGKGRGRGRGR 183
>TC216338 similar to UP|O19031 (O19031) Bactinecin 11, partial (7%)
Length = 420
Score = 39.3 bits (90), Expect = 0.002
Identities = 17/19 (89%), Positives = 17/19 (89%)
Frame = +1
Query: 139 GRGRGRGRGRGRGRGRTAV 157
GRGRGRGRGRGRGRGR V
Sbjct: 76 GRGRGRGRGRGRGRGRDQV 132
Score = 36.2 bits (82), Expect = 0.017
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +1
Query: 140 RGRGRGRGRGRGRGR 154
RGRGRGRGRGRGRGR
Sbjct: 73 RGRGRGRGRGRGRGR 117
Score = 32.3 bits (72), Expect = 0.25
Identities = 14/18 (77%), Positives = 15/18 (82%)
Frame = +1
Query: 142 RGRGRGRGRGRGRTAVRE 159
RGRGRGRGRGRGR R+
Sbjct: 73 RGRGRGRGRGRGRGRGRD 126
>TC216357 homologue to GB|AAM70547.1|21700847|AY124838 AT3g07590/MLP3_4
{Arabidopsis thaliana;} , complete
Length = 809
Score = 38.5 bits (88), Expect = 0.003
Identities = 16/16 (100%), Positives = 16/16 (100%)
Frame = +2
Query: 139 GRGRGRGRGRGRGRGR 154
GRGRGRGRGRGRGRGR
Sbjct: 437 GRGRGRGRGRGRGRGR 484
Score = 35.0 bits (79), Expect = 0.038
Identities = 16/25 (64%), Positives = 16/25 (64%)
Frame = +2
Query: 128 KMPELSHTGSTGRGRGRGRGRGRGR 152
K P GRGRGRGRGRGRGR
Sbjct: 410 KKPTAGKPLGRGRGRGRGRGRGRGR 484
Score = 33.5 bits (75), Expect = 0.11
Identities = 15/29 (51%), Positives = 18/29 (61%)
Frame = +2
Query: 137 STGRGRGRGRGRGRGRGRTAVRETPHQAV 165
+ G+ GRGRGRGRGRGR R H +
Sbjct: 419 TAGKPLGRGRGRGRGRGRGRGR*VAHHKI 505
>TC206156 homologue to UP|Q9SY09 (Q9SY09) Small nuclear riboprotein Sm-D1,
partial (98%)
Length = 842
Score = 36.6 bits (83), Expect = 0.013
Identities = 17/27 (62%), Positives = 17/27 (62%)
Frame = +1
Query: 128 KMPELSHTGSTGRGRGRGRGRGRGRGR 154
K P GRGRGRGRGRGRG GR
Sbjct: 421 KKPTAGKPLGRGRGRGRGRGRGRGGGR 501
>TC206155 homologue to GB|AAM70547.1|21700847|AY124838 AT3g07590/MLP3_4
{Arabidopsis thaliana;} , complete
Length = 985
Score = 36.6 bits (83), Expect = 0.013
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +1
Query: 139 GRGRGRGRGRGRGRG 153
GRGRGRGRGRGRGRG
Sbjct: 406 GRGRGRGRGRGRGRG 450
Score = 34.3 bits (77), Expect = 0.066
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = +1
Query: 141 GRGRGRGRGRGRGR 154
GRGRGRGRGRGRGR
Sbjct: 406 GRGRGRGRGRGRGR 447
Score = 33.1 bits (74), Expect = 0.15
Identities = 15/24 (62%), Positives = 15/24 (62%)
Frame = +1
Query: 128 KMPELSHTGSTGRGRGRGRGRGRG 151
K P GRGRGRGRGRGRG
Sbjct: 379 KKPTAGKPLGRGRGRGRGRGRGRG 450
Score = 32.0 bits (71), Expect = 0.33
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = +1
Query: 137 STGRGRGRGRGRGRGRGR 154
+ G+ GRGRGRGRGRGR
Sbjct: 388 TAGKPLGRGRGRGRGRGR 441
>AW311339 similar to GP|18146720|dbj ribosomal protein S2 {Arabidopsis
thaliana}, partial (6%)
Length = 358
Score = 36.2 bits (82), Expect = 0.017
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
RGRGRGRGRGRGRGR
Sbjct: 60 RGRGRGRGRGRGRGR 104
Score = 35.8 bits (81), Expect = 0.023
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
GRGRGRGRGRGRGR R
Sbjct: 63 GRGRGRGRGRGRGRDR 110
Score = 34.3 bits (77), Expect = 0.066
Identities = 15/18 (83%), Positives = 15/18 (83%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTA 156
GRGRGRGRGRGR RG A
Sbjct: 69 GRGRGRGRGRGRDRGPPA 122
Score = 33.1 bits (74), Expect = 0.15
Identities = 15/20 (75%), Positives = 15/20 (75%)
Frame = +3
Query: 142 RGRGRGRGRGRGRTAVRETP 161
RGRGRGRGRGRGR R P
Sbjct: 60 RGRGRGRGRGRGRGRDRGPP 119
>TC216337 similar to UP|Q7XJ13 (Q7XJ13) Chloroplast nucleoid DNA-binding
protein-like protein, partial (21%)
Length = 1139
Score = 36.2 bits (82), Expect = 0.017
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
RGRGRGRGRGRGRGR
Sbjct: 801 RGRGRGRGRGRGRGR 845
Score = 35.8 bits (81), Expect = 0.023
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
GRGRGRGRGRGRGR R
Sbjct: 804 GRGRGRGRGRGRGRDR 851
Score = 34.3 bits (77), Expect = 0.066
Identities = 15/18 (83%), Positives = 15/18 (83%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTA 156
GRGRGRGRGRGR RG A
Sbjct: 810 GRGRGRGRGRGRDRGPPA 863
Score = 34.3 bits (77), Expect = 0.066
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = +3
Query: 140 RGRGRGRGRGRGRG 153
RGRGRGRGRGRGRG
Sbjct: 588 RGRGRGRGRGRGRG 629
Score = 33.1 bits (74), Expect = 0.15
Identities = 15/20 (75%), Positives = 15/20 (75%)
Frame = +3
Query: 142 RGRGRGRGRGRGRTAVRETP 161
RGRGRGRGRGRGR R P
Sbjct: 801 RGRGRGRGRGRGRGRDRGPP 860
Score = 32.7 bits (73), Expect = 0.19
Identities = 14/15 (93%), Positives = 14/15 (93%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
R RGRGRGRGRGRGR
Sbjct: 582 RMRGRGRGRGRGRGR 626
Score = 32.3 bits (72), Expect = 0.25
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = +3
Query: 139 GRGRGRGRGRGRG 151
GRGRGRGRGRGRG
Sbjct: 591 GRGRGRGRGRGRG 629
Score = 30.0 bits (66), Expect = 1.2
Identities = 15/20 (75%), Positives = 15/20 (75%), Gaps = 1/20 (5%)
Frame = +3
Query: 136 GSTGRGRGRGRGRG-RGRGR 154
G GRGRGR RGR RGRGR
Sbjct: 678 GYGGRGRGRARGRAFRGRGR 737
Score = 29.6 bits (65), Expect = 1.6
Identities = 13/16 (81%), Positives = 13/16 (81%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
G R RGRGRGRGRGR
Sbjct: 573 GSPRMRGRGRGRGRGR 620
Score = 27.7 bits (60), Expect = 6.1
Identities = 12/16 (75%), Positives = 12/16 (75%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGR 154
G G R RGRGRGRGR
Sbjct: 567 GEGSPRMRGRGRGRGR 614
>TC225776 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, partial (26%)
Length = 1663
Score = 36.2 bits (82), Expect = 0.017
Identities = 38/129 (29%), Positives = 57/129 (43%), Gaps = 20/129 (15%)
Frame = +1
Query: 137 STGRGRGRGRGR----GRGRGR------TAVRETPHQAV-EPEPCASVQQGIQHDTNTDM 185
+ GRGRGRGRGR GRGRGR +R++ + V P+ ++ Q+ N D+
Sbjct: 718 AAGRGRGRGRGRGGGGGRGRGRGRGQLPAQLRKSGARPVGRPKKGSTSASTTQNAANEDL 897
Query: 186 ---TIHDSSETKE-----LPKENAAVPAES-AEFHNLDLNANTNENEDKKASTTAKQEIS 236
H S+ KE P N P + A L++ + N + T QE
Sbjct: 898 RRKLEHFQSKVKESLAVLKPHFNHESPVTAIAAIQELEVLGTMDLNAPLRDETLPPQE-E 1074
Query: 237 EPPTESQHE 245
+PP + Q +
Sbjct: 1075QPPLQQQQQ 1101
>TC212981 similar to UP|BCT7_BOVIN (P19661) Bactenecin 7 precursor (BAC7)
(PR-59), partial (9%)
Length = 574
Score = 35.8 bits (81), Expect = 0.023
Identities = 16/19 (84%), Positives = 16/19 (84%)
Frame = +3
Query: 139 GRGRGRGRGRGRGRGRTAV 157
GR RGRGRGRGRGRGR V
Sbjct: 75 GRVRGRGRGRGRGRGRDQV 131
Score = 32.7 bits (73), Expect = 0.19
Identities = 14/15 (93%), Positives = 14/15 (93%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGR 154
RGR RGRGRGRGRGR
Sbjct: 72 RGRVRGRGRGRGRGR 116
Score = 32.3 bits (72), Expect = 0.25
Identities = 13/17 (76%), Positives = 15/17 (87%)
Frame = +3
Query: 140 RGRGRGRGRGRGRGRTA 156
RGRGRGRGRGRGR + +
Sbjct: 84 RGRGRGRGRGRGRDQVS 134
Score = 29.3 bits (64), Expect = 2.1
Identities = 14/24 (58%), Positives = 16/24 (66%)
Frame = +3
Query: 136 GSTGRGRGRGRGRGRGRGRTAVRE 159
G+ RGR RGRGRGRGR R+
Sbjct: 54 GAPPPXRGRVRGRGRGRGRGRGRD 125
>TC218974 weakly similar to UP|PCD7_MOUSE (Q9WTY1) Programmed cell death
protein 7 (ES18), partial (7%)
Length = 890
Score = 35.4 bits (80), Expect = 0.029
Identities = 30/86 (34%), Positives = 41/86 (46%), Gaps = 8/86 (9%)
Frame = -3
Query: 82 DVVSKVPDYSHGHGHSE---PSADDRTIPKRRKAAGD-DCNDSDEEVKRGKM---PELSH 134
+V + +P + H H+E P A RRK G+ + ++ EEV+R P +
Sbjct: 414 EVAAVLPRQARDHAHAEVGEPVAQVHGA*YRRKGQGEPEEGET*EEVRREVAVVGPAVEE 235
Query: 135 TGSTGRGRGRG-RGRGRGRGRTAVRE 159
G G GRG G RG G GRG RE
Sbjct: 234 YGLAGGGRGGGGRGGGGGRGHV*PRE 157
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,711,353
Number of Sequences: 63676
Number of extensions: 140127
Number of successful extensions: 2225
Number of sequences better than 10.0: 138
Number of HSP's better than 10.0 without gapping: 1326
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1714
length of query: 283
length of database: 12,639,632
effective HSP length: 96
effective length of query: 187
effective length of database: 6,526,736
effective search space: 1220499632
effective search space used: 1220499632
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0005.4


