

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.12
(526 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
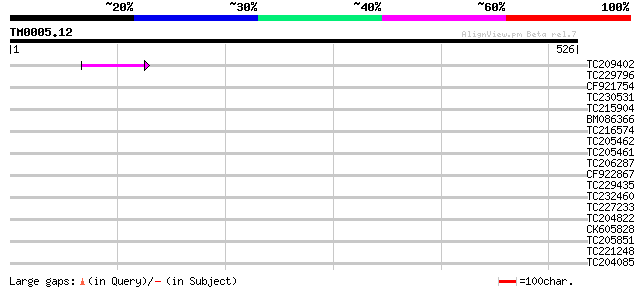
Score E
Sequences producing significant alignments: (bits) Value
TC209402 47 3e-05
TC229796 38 0.010
CF921754 38 0.013
TC230531 weakly similar to UP|Q941H9 (Q941H9) RNA-binding protei... 36 0.048
TC215904 similar to UP|Q9M7N4 (Q9M7N4) MFP1 attachment factor 1,... 35 0.082
BM086366 35 0.11
TC216574 similar to UP|RK4_SPIOL (O49937) 50S ribosomal protein ... 35 0.11
TC205462 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MS... 35 0.11
TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MS... 35 0.11
TC206287 UP|HMGA_SOYBN (Q00423) HMG-Y related protein A (SB16A p... 34 0.18
CF922867 33 0.24
TC229435 weakly similar to UP|Q14883 (Q14883) Intestinal mucin (... 33 0.31
TC232460 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, par... 33 0.41
TC227233 similar to GB|AAO42915.1|28416769|BT004669 At4g10262 {A... 32 0.53
TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 32 0.53
CK605828 32 0.53
TC205851 similar to GB|AAG53636.1|12407751|AF291712 initiation f... 32 0.53
TC221248 weakly similar to GB|CAD40580.1|21741678|OSJN00141 oj00... 32 0.53
TC204085 similar to UP|Q8VX74 (Q8VX74) Glycine-rich RNA-binding ... 32 0.69
>TC209402
Length = 1163
Score = 46.6 bits (109), Expect = 3e-05
Identities = 20/63 (31%), Positives = 34/63 (53%)
Frame = +1
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRK 126
++V + +++ +++LF HGK+T V L ++ K RFGFV +A R MK
Sbjct: 223 VYVKNLPENITQDRLKELFEHHGKITKVVLPSAKSGQEKSRFGFVHFAERSSAMKALKNT 402
Query: 127 QSY 129
+ Y
Sbjct: 403 EKY 411
>TC229796
Length = 413
Score = 38.1 bits (87), Expect = 0.010
Identities = 18/46 (39%), Positives = 25/46 (54%)
Frame = +2
Query: 84 LFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQSY 129
LF +HGK+T V L ++ K R GFV +A R + MK + Y
Sbjct: 2 LFERHGKITKVVLPPAKSGQEKNRIGFVHFAERSNAMKALKNTERY 139
>CF921754
Length = 529
Score = 37.7 bits (86), Expect = 0.013
Identities = 23/56 (41%), Positives = 31/56 (55%)
Frame = +1
Query: 438 SPPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPPPQLHP 493
+PP P G+PK+KK K L+G + P P K+G A P P SPPP ++P
Sbjct: 100 NPPPKNPLGKPKKKK--KAPLKGLKTP---PPKAG----GAPPPPLRGSPPPGVNP 240
>TC230531 weakly similar to UP|Q941H9 (Q941H9) RNA-binding protein precursor,
partial (29%)
Length = 851
Score = 35.8 bits (81), Expect = 0.048
Identities = 25/91 (27%), Positives = 38/91 (41%)
Frame = +3
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRK 126
LFV G++ N +RD F QHG++ V + G+ +GFVR+ + R
Sbjct: 420 LFVTGLSYDTNEPILRDAFGQHGEIIEVKVICDHVTGKSRGYGFVRFVSETTAAAARKEM 599
Query: 127 QSYVSMAPPTTVKDSRAKERLHLSGGFASSS 157
+ V + ERL L +SS
Sbjct: 600 NGQILDGRRIRVSYAHKGERLSLPNNS*NSS 692
>TC215904 similar to UP|Q9M7N4 (Q9M7N4) MFP1 attachment factor 1, partial
(64%)
Length = 761
Score = 35.0 bits (79), Expect = 0.082
Identities = 25/72 (34%), Positives = 32/72 (43%)
Frame = +1
Query: 439 PPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPPPQLHPAHITR 498
P R R R P + GSR + PR+S + +PS P+PPP L P IT
Sbjct: 292 PARSRTRPSPSPPAPPPPTMTGSRSSRSTPRRSA----NGCSTPSRPNPPP-LPPLSITG 456
Query: 499 GLQRLGFWQNPQ 510
L R+ PQ
Sbjct: 457 LLPRMLLLPPPQ 492
>BM086366
Length = 427
Score = 34.7 bits (78), Expect = 0.11
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 3/58 (5%)
Frame = +2
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANR---EDVMK 121
+FV G++ V + F ++GK+ + +R+ GR FGF+ +A+R ED +K
Sbjct: 95 IFVGGLSWEVTERQLEHAFARYGKILECQIMMERDTGRPRGFGFITFADRRGMEDAIK 268
>TC216574 similar to UP|RK4_SPIOL (O49937) 50S ribosomal protein L4,
chloroplast precursor (R-protein L4), partial (74%)
Length = 1364
Score = 34.7 bits (78), Expect = 0.11
Identities = 18/55 (32%), Positives = 25/55 (44%)
Frame = +2
Query: 439 PPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPPPQLHP 493
P R RP RP R++ R HPR+ +A P+ + S PP+L P
Sbjct: 302 PSRHRPSRRPPRRRHRPPEQAPRHRLHPHPRRGPRRRTEALPAEKNRSRPPRLQP 466
>TC205462 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;} , partial (54%)
Length = 1403
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/79 (27%), Positives = 37/79 (45%), Gaps = 3/79 (3%)
Frame = +3
Query: 84 LFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQSYVSMAPPTTVKDSR- 142
LF ++GK+ ++F+ + R G F FVRY ++ K R + TV+ ++
Sbjct: 192 LFDKYGKVVDIFIPKDRRTGESRGFAFVRYKYADEAQKAVERLDGRMVDGREITVQFAKY 371
Query: 143 --AKERLHLSGGFASSSRS 159
ER+H +S RS
Sbjct: 372 GPNAERIHKGRIIETSPRS 428
>TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;} , partial (69%)
Length = 1148
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/79 (27%), Positives = 37/79 (45%), Gaps = 3/79 (3%)
Frame = +3
Query: 84 LFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQSYVSMAPPTTVKDSR- 142
LF ++GK+ ++F+ + R G F FVRY ++ K R + TV+ ++
Sbjct: 195 LFDKYGKVVDIFIPKDRRTGESRGFAFVRYKYADEAQKAVERLDGRMVDGREITVQFAKY 374
Query: 143 --AKERLHLSGGFASSSRS 159
ER+H +S RS
Sbjct: 375 GPNAERIHKGRIIETSPRS 431
>TC206287 UP|HMGA_SOYBN (Q00423) HMG-Y related protein A (SB16A protein),
complete
Length = 1058
Score = 33.9 bits (76), Expect = 0.18
Identities = 23/62 (37%), Positives = 29/62 (46%), Gaps = 10/62 (16%)
Frame = +3
Query: 442 PRPRGRPKR------KKEVKRGLEGSRRPQGHPRKSGTMDLDAAP---SPSHP-SPPPQL 491
PRPRGRP + K + GS RP+G P+K AAP S P PP++
Sbjct: 642 PRPRGRPPKDPNAPPKSPKAKATPGSGRPRGRPKKVPRSPAVAAPTAVSSGRPRGRPPKV 821
Query: 492 HP 493
P
Sbjct: 822 KP 827
>CF922867
Length = 450
Score = 33.5 bits (75), Expect = 0.24
Identities = 19/63 (30%), Positives = 29/63 (45%), Gaps = 2/63 (3%)
Frame = +1
Query: 439 PPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAP--SPSHPSPPPQLHPAHI 496
PPR +P+ PK K + KR + + P P+K G + P + P + P+H
Sbjct: 37 PPREKPKPNPK*KSQRKRAPQKTPPPPKLPKKGGPQTPNRRPWEKKTPPGGKKEKKPSHP 216
Query: 497 TRG 499
RG
Sbjct: 217 PRG 225
>TC229435 weakly similar to UP|Q14883 (Q14883) Intestinal mucin (Fragment),
partial (26%)
Length = 1075
Score = 33.1 bits (74), Expect = 0.31
Identities = 20/58 (34%), Positives = 28/58 (47%)
Frame = +1
Query: 434 VLDFSPPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPPPQL 491
+L+ +PP P+PR RP+R++ +R RRPQ P P PPP L
Sbjct: 334 LLEEAPPTPQPRRRPRRRRRRRR---RRRRPQ----------------PLPPQPPPLL 450
>TC232460 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, partial (22%)
Length = 637
Score = 32.7 bits (73), Expect = 0.41
Identities = 32/100 (32%), Positives = 36/100 (36%), Gaps = 3/100 (3%)
Frame = +2
Query: 430 ADDPVLDFSPPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDA--APSPSHPSP 487
A P S PRP P P R+ G HP SG A P+P HP P
Sbjct: 263 AGPPPPSSSGPRPSPSPPPSRRLPRLAARAGPAARWAHPAASGGDHRGAPGRPAPGHPGP 442
Query: 488 P-PQLHPAHITRGLQRLGFWQNPQACLFREVMKRPFEGWH 526
Q P RG Q F+ P R V+ R G H
Sbjct: 443 AGRQPAPGRHPRGPQAGAFFL-PARAPARGVLPRLTPGRH 559
>TC227233 similar to GB|AAO42915.1|28416769|BT004669 At4g10262 {Arabidopsis
thaliana;} , partial (19%)
Length = 571
Score = 32.3 bits (72), Expect = 0.53
Identities = 24/59 (40%), Positives = 28/59 (46%), Gaps = 8/59 (13%)
Frame = -3
Query: 438 SPPRPRPRGRPKRKKEVKRGLEGSRRPQGHPRKSG-------TMDLD-AAPSPSHPSPP 488
SPP RPR P+R+KE L G RR +G G T D P+P PSPP
Sbjct: 230 SPPPSRPRQIPRRRKEA-GDLTGLRRRRGGSGCGGWR*APSCTCGRDRCGPAPPPPSPP 57
>TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (67%)
Length = 1232
Score = 32.3 bits (72), Expect = 0.53
Identities = 19/72 (26%), Positives = 33/72 (45%), Gaps = 4/72 (5%)
Frame = +3
Query: 52 RVARVVAPPARTCFP----LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFR 107
R R PP R F ++V + V+ ++ +F +HG + N + R GR
Sbjct: 567 RGTRPERPPPRRSFESSLSIYVGNLPWDVDNTRLKQIFSKHGNVVNARVVYDRESGRSRG 746
Query: 108 FGFVRYANREDV 119
FGFV ++ ++
Sbjct: 747 FGFVTMSDETEM 782
>CK605828
Length = 553
Score = 32.3 bits (72), Expect = 0.53
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 3/59 (5%)
Frame = +2
Query: 438 SPPRPRP---RGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPPPQLHP 493
+PP+ P RG PK+ + K+G EG+ RP R+ G + HP PP+ +P
Sbjct: 224 NPPKRGPAQKRGPPKKTQ--KKGGEGTPRPPFGGRRGGPTRGNPPQGGYHPQTPPRENP 394
>TC205851 similar to GB|AAG53636.1|12407751|AF291712 initiation factor 3g
{Arabidopsis thaliana;} , partial (45%)
Length = 641
Score = 32.3 bits (72), Expect = 0.53
Identities = 14/47 (29%), Positives = 24/47 (50%)
Frame = +1
Query: 83 DLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQSY 129
+LF G ++ V++ + G FGFV + NRED + ++ Y
Sbjct: 262 ELFRPFGPVSRVYVAIDQKTGMSRGFGFVNFVNREDAQRAINKLNGY 402
>TC221248 weakly similar to GB|CAD40580.1|21741678|OSJN00141 oj000126_13.2
{Oryza sativa (japonica cultivar-group);} , partial
(40%)
Length = 861
Score = 32.3 bits (72), Expect = 0.53
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Frame = +1
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKL--TNVFLQRKRNRGRKFRFGFVRYANREDVMKGR 123
+FV G+A S + F Q+G + N+ L + +NR + F G+V +A E+ K +
Sbjct: 388 VFVKGLAFSTTEEELAKAFSQYGSVLKANIILNKAKNRSKGF--GYVTFAKEEEACKAQ 558
>TC204085 similar to UP|Q8VX74 (Q8VX74) Glycine-rich RNA-binding protein,
partial (87%)
Length = 628
Score = 32.0 bits (71), Expect = 0.69
Identities = 18/54 (33%), Positives = 29/54 (53%)
Frame = +3
Query: 68 FVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
FV G+A + + + + F Q+G + + R GR FGFV +A+ ED M+
Sbjct: 36 FVGGLAWATDNYDLEKAFSQYGDVVESKIINDRETGRSRGFGFVTFAS-EDSMR 194
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,033,393
Number of Sequences: 63676
Number of extensions: 419948
Number of successful extensions: 4825
Number of sequences better than 10.0: 148
Number of HSP's better than 10.0 without gapping: 4106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4683
length of query: 526
length of database: 12,639,632
effective HSP length: 102
effective length of query: 424
effective length of database: 6,144,680
effective search space: 2605344320
effective search space used: 2605344320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0005.12


