

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
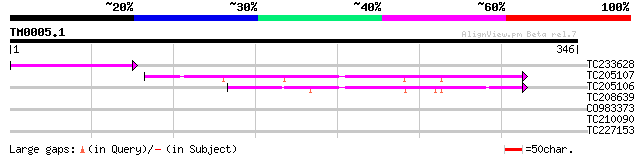
TC233628 weakly similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (6%) 55 4e-08
TC205107 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (34%) 50 2e-06
TC205106 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (33%) 42 5e-04
TC208639 similar to UP|Q9LMB8 (Q9LMB8) F14D16.25, partial (14%) 30 1.6
CO983373 29 2.7
TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, par... 28 8.0
TC227153 similar to UP|Q9FMI3 (Q9FMI3) Emb|CAB77570.1, partial (... 28 8.0
>TC233628 weakly similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (6%)
Length = 626
Score = 55.5 bits (132), Expect = 4e-08
Identities = 26/78 (33%), Positives = 45/78 (57%)
Frame = +1
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+ + LN +YQ+ SKL++H S P +PK++ +EK + ++ E A ++ KS ++
Sbjct: 22 MKENYLPELNEMYQKSASKLQRHTSLPQQPKSYKLEKLKKFMKMLECAIAFLQVSKSNMS 201
Query: 61 PEISERVDRVETLINSII 78
P ++D E I II
Sbjct: 202PNYRLKLDSCEKQIIKII 255
>TC205107 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (34%)
Length = 1762
Score = 49.7 bits (117), Expect = 2e-06
Identities = 70/266 (26%), Positives = 110/266 (41%), Gaps = 32/266 (12%)
Frame = +2
Query: 83 LSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQV-----------AK 131
+SS Q +A Q +Q PQV Q +HL +T + +
Sbjct: 203 VSSPQLLQAASPQIQQHSSPQV--DQQNHLPSKTKVTTPLQSSNSPFVGPTPSPPLAPSP 376
Query: 132 QYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEAS------GVSTNDMGFSSSALIKGSG 185
G SE+ + +S +QT A A S G+S + + SA G
Sbjct: 377 MPGESEKPIPCVSSISNAANIGHQQTGGAIAPAQSLAIGTPGISASPLLAEFSAPDGAHG 556
Query: 186 NLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPA------ 239
N + + ++E+P ++RLI SMSS+AL A++ I VV +ND I
Sbjct: 557 NALAATSGKSTVTEQP---LERLINAVKSMSSKALSAAVMDIGSVVSMNDRIAGSAPGNG 727
Query: 240 --AKFIDEPLEMAGQQNQPSR-IPQG-----RKMARNINAMAFDTSRSCARTHESFNQLT 291
A ++ + M + Q I Q ++M R +A+ + S ++S QLT
Sbjct: 728 SRAAVGEDLVSMTNCRLQARNFITQDGSNGIKRMKRYTSAIPLNVVPSAGSMNDSIKQLT 907
Query: 292 DVEEPDLDLVA-SKVKRSRIEVIYLL 316
E DL+ A S VK+ +IEV + L
Sbjct: 908 ASETSDLESTATSSVKKPKIEVNHAL 985
>TC205106 similar to UP|Q8H6Q7 (Q8H6Q7) CTV.22, partial (33%)
Length = 1796
Score = 41.6 bits (96), Expect = 5e-04
Identities = 57/205 (27%), Positives = 88/205 (42%), Gaps = 22/205 (10%)
Frame = +3
Query: 134 GNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIK-------GSGN 186
G+SE+ ++ +S +QT A A S ++ G S+S L+ GN
Sbjct: 324 GDSEKLISGVSSISNAANIGYQQTGGAAAPGQS-LAIGTPGISASPLLAEFTGPDGAHGN 500
Query: 187 LNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAA------ 240
+ + ++E+P ++RLI SMS +AL +++ I VV +ND I +
Sbjct: 501 SLAPTSGKSTVTEQP---IERLIKAVKSMSPKALSSAVSDIGSVVSMNDRIAGSAPGNGS 671
Query: 241 -KFIDEPLEMAGQQNQPSR--IPQG-----RKMARNINAMAFDTSRSCARTHESFNQLTD 292
+ E L +R I Q R+M R NA + S ++S QLT
Sbjct: 672 RAAVGEDLVAMTNCRLQARNFITQDGANGTRRMKRYTNATPLNVVTSAGSMNDSIKQLT- 848
Query: 293 VEEPDLDLVA-SKVKRSRIEVIYLL 316
E DLD A S+ K RIE + L
Sbjct: 849 AEASDLDSTATSRFKMPRIEANHSL 923
>TC208639 similar to UP|Q9LMB8 (Q9LMB8) F14D16.25, partial (14%)
Length = 1214
Score = 30.0 bits (66), Expect = 1.6
Identities = 17/63 (26%), Positives = 29/63 (45%)
Frame = +3
Query: 95 QSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQ 154
+ K+++ P+V Q ++ N C +YGNSEQ +L I+ G +
Sbjct: 570 KDKKKLPPRVTSMQKPLAIEKPVMLNRSCSPVVNTKIEYGNSEQCSTKLQIRSEQGSHNK 749
Query: 155 RQT 157
R+T
Sbjct: 750 RKT 758
>CO983373
Length = 713
Score = 29.3 bits (64), Expect = 2.7
Identities = 21/56 (37%), Positives = 30/56 (53%), Gaps = 4/56 (7%)
Frame = -1
Query: 63 ISERVDRVETLINSIIQHQNLSSHQGKHS----ADVQSKQQVLPQVLGTQNDHLAK 114
I E + ETL N I L +HQ K A++++K+++L Q L T NDH K
Sbjct: 350 IKEGEKQRETLKNDSI----LWAHQTKDLVFDLAEIEAKEKILGQQLETDNDHQTK 195
>TC210090 similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, partial (12%)
Length = 1189
Score = 27.7 bits (60), Expect = 8.0
Identities = 14/55 (25%), Positives = 27/55 (48%), Gaps = 4/55 (7%)
Frame = -3
Query: 41 HKRLYEDIFAMFRLRKSQITPEISERVDRVETLINSIIQ----HQNLSSHQGKHS 91
H R Y+D F++ L ++ TP S+ + + S Q ++NL+ + H+
Sbjct: 1136 HNRSYKDQFSIANLENNKFTPTTSQEISHYLNNLQSSYQFSRNYKNLNHYHRAHA 972
>TC227153 similar to UP|Q9FMI3 (Q9FMI3) Emb|CAB77570.1, partial (10%)
Length = 1583
Score = 27.7 bits (60), Expect = 8.0
Identities = 20/78 (25%), Positives = 38/78 (48%), Gaps = 6/78 (7%)
Frame = +1
Query: 68 DRVETLINSIIQHQNLSSHQGKH------SADVQSKQQVLPQVLGTQNDHLAKQTSQRNN 121
D++E+ I++ IQ+++ ++ K D++S QQ+ PQVL Q Q +N+
Sbjct: 874 DKLESPISAKIQYESKEANLEKFPMGCLPDEDMKSNQQISPQVLEAQ-----PQDQFKNS 1038
Query: 122 MCLQEQQVAKQYGNSEQG 139
C + A ++ G
Sbjct: 1039HCRDSSENASNNDLADAG 1092
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,784,551
Number of Sequences: 63676
Number of extensions: 151094
Number of successful extensions: 973
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 971
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 972
length of query: 346
length of database: 12,639,632
effective HSP length: 98
effective length of query: 248
effective length of database: 6,399,384
effective search space: 1587047232
effective search space used: 1587047232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0005.1


