

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0004a.3
(148 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
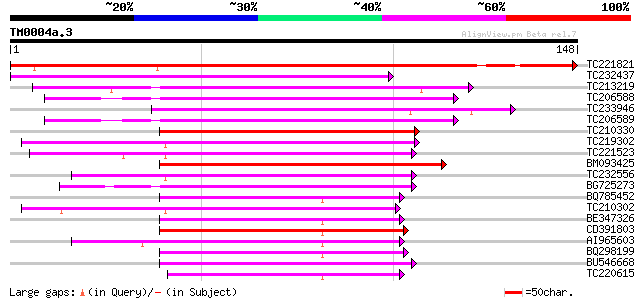
TC221821 weakly similar to UP|WSC2_YEAST (P53832) Cell wall inte... 193 2e-50
TC232437 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial ... 74 3e-14
TC213219 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial ... 74 3e-14
TC206588 weakly similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, p... 71 2e-13
TC233946 UP|Q7R3F4 (Q7R3F4) GLP_158_77883_78482, partial (5%) 71 2e-13
TC206589 similar to UP|Q6NMZ3 (Q6NMZ3) At4g34750, partial (39%) 70 3e-13
TC210330 67 3e-12
TC219302 similar to UP|Q9LR00 (Q9LR00) F10A5.21, partial (95%) 66 7e-12
TC221523 similar to UP|O49581 (O49581) Auxin induced like-protei... 63 5e-11
BM093425 homologue to GP|24943206|gb auxin-regulated protein {Ph... 62 1e-10
TC232556 62 1e-10
BG725273 similar to PIR|F86331|F863 F6F9.11 protein - Arabidopsi... 60 3e-10
BQ785452 60 4e-10
TC210302 60 4e-10
BE347326 similar to SP|P32295|ARG7_P Indole-3-acetic acid induce... 59 9e-10
CD391803 similar to SP|P33079|AXA5_S Auxin-induced protein 10A5.... 59 9e-10
AI965603 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [... 59 1e-09
BQ298199 similar to SP|P33079|AXA5_S Auxin-induced protein 10A5.... 59 1e-09
BU546668 weakly similar to GP|20161745|dbj B1033B05.12 {Oryza sa... 58 1e-09
TC220615 homologue to UP|AX6B_SOYBN (P33083) Auxin-induced prote... 58 2e-09
>TC221821 weakly similar to UP|WSC2_YEAST (P53832) Cell wall integrity and
stress response component 2 precursor, partial (6%)
Length = 735
Score = 193 bits (491), Expect = 2e-50
Identities = 104/153 (67%), Positives = 122/153 (78%), Gaps = 5/153 (3%)
Frame = +2
Query: 1 MAGAM-KKVDKIRQIVRLKQVMMRWKQISLRRHDSSDP----RPPSGCLYVYVGHERQRF 55
MAGAM KVDKIRQIVRLKQ+M RWK ISLRR S +P RPPSG ++VYVG ER RF
Sbjct: 71 MAGAMGMKVDKIRQIVRLKQLMTRWKHISLRRRSSDEPSAVRRPPSGFIFVYVGPERTRF 250
Query: 56 AIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKYGKLTLE 115
AIPARFLNL +F GLL +TEEEFGLRG+GGL+LPC V FF+++VK LHK+E+KYG L+L+
Sbjct: 251 AIPARFLNLALFEGLLKQTEEEFGLRGNGGLVLPCQVPFFSNVVKYLHKDEHKYGSLSLQ 430
Query: 116 QFVITFSDAAANFDSCKDQNVVVFTPLLQKAEV 148
FV S A++ DSCK +NV VF PLLQKAEV
Sbjct: 431 DFVNMLS--ASSSDSCK-ENVAVFAPLLQKAEV 520
>TC232437 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial (71%)
Length = 587
Score = 73.9 bits (180), Expect = 3e-14
Identities = 37/100 (37%), Positives = 56/100 (56%)
Frame = +1
Query: 1 MAGAMKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPAR 60
M+ A+ K +IR IVRL+Q++ RW+ + P+G + V VG+ +RF +
Sbjct: 13 MSAALGKCSRIRHIVRLRQMLRRWRSKARTSAHRIPSDVPAGHVAVCVGNNSKRFVVRTT 192
Query: 61 FLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVK 100
+LN PVF LL E EEE+G G L +PCD F +++
Sbjct: 193 YLNHPVFKRLLVEAEEEYGFSNHGPLAIPCDEAIFEQLLR 312
>TC213219 similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial (67%)
Length = 476
Score = 73.6 bits (179), Expect = 3e-14
Identities = 44/122 (36%), Positives = 69/122 (56%), Gaps = 7/122 (5%)
Frame = +2
Query: 7 KVDKIRQIVRLKQVMMRWK---QISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLN 63
K +IR IVRL+Q++ RW+ ++S R SD P+G + V VG RF + A +LN
Sbjct: 89 KCSQIRHIVRLRQMLRRWRNKARMSANRAPPSDV--PAGHVAVCVGSNLTRFVVRATYLN 262
Query: 64 LPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNE----NKYGKLTLEQFVI 119
PVF LL + EEE+G G L +PCD F +++ + +++ N++ L L+ F
Sbjct: 263 HPVFKKLLLQAEEEYGFTNHGPLAIPCDETLFRDVLRFISRSDPAKSNRFLNLELDNFNP 442
Query: 120 TF 121
+F
Sbjct: 443 SF 448
>TC206588 weakly similar to UP|Q9FXI2 (Q9FXI2) F6F9.11 protein, partial (44%)
Length = 633
Score = 70.9 bits (172), Expect = 2e-13
Identities = 42/108 (38%), Positives = 63/108 (57%)
Frame = +1
Query: 10 KIRQIVRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAG 69
KIR+IVR++Q+++RW RR ++D P+G + V VG R+RF + A LN P+F
Sbjct: 127 KIRRIVRVRQMLLRW-----RRKAAADV--PAGHVAVCVGPSRRRFIVRATHLNHPIFKM 285
Query: 70 LLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKYGKLTLEQF 117
LL + EEE+G G L +PCD F +++ + + G TLE F
Sbjct: 286 LLVKAEEEYGFCNHGPLAIPCDESLFEELLRVVSRPVPVPGFSTLEDF 429
>TC233946 UP|Q7R3F4 (Q7R3F4) GLP_158_77883_78482, partial (5%)
Length = 677
Score = 70.9 bits (172), Expect = 2e-13
Identities = 39/104 (37%), Positives = 56/104 (53%), Gaps = 9/104 (8%)
Frame = +2
Query: 38 RPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTH 97
+ P GC+ VYVG ER+RF I + N P+F LLD E E+G R +G L LPCDV F+
Sbjct: 188 KAPQGCICVYVGAERERFVIKVKIANHPLFKALLDAAEREYGYRNNGPLWLPCDVDLFSE 367
Query: 98 IVKRLH---KNENKYGKLTLEQFVI------TFSDAAANFDSCK 132
+K + + E + G + F + S A+ N+ SC+
Sbjct: 368 ALKDMENSIQEEEETGSVGCAAFAAGHSFRRSPSSASNNYISCR 499
>TC206589 similar to UP|Q6NMZ3 (Q6NMZ3) At4g34750, partial (39%)
Length = 686
Score = 70.5 bits (171), Expect = 3e-13
Identities = 42/108 (38%), Positives = 63/108 (57%)
Frame = +3
Query: 10 KIRQIVRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAG 69
KIR+IVR++Q+++RW RR + D P+G + V VG R+RF + A LN P+F
Sbjct: 93 KIRRIVRVRQMLLRW-----RRKVAVDV--PAGHVAVCVGPSRRRFIVRATHLNHPIFKM 251
Query: 70 LLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNENKYGKLTLEQF 117
LL + EEE+G G L +PCD F H+++ + + G +LE F
Sbjct: 252 LLVKAEEEYGFCNHGPLAIPCDESLFEHLLRVVARPVPLPGFSSLEDF 395
>TC210330
Length = 706
Score = 67.0 bits (162), Expect = 3e-12
Identities = 29/68 (42%), Positives = 43/68 (62%)
Frame = +1
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P GC V+VG ERQRF + +++N P+F LL+ETE+E+G G + LPC+V F ++
Sbjct: 319 PHGCFSVHVGPERQRFVVKTKYVNHPLFQMLLEETEQEYGFESDGPIWLPCNVDLFYKVL 498
Query: 100 KRLHKNEN 107
+ EN
Sbjct: 499 AEMDGEEN 522
>TC219302 similar to UP|Q9LR00 (Q9LR00) F10A5.21, partial (95%)
Length = 731
Score = 65.9 bits (159), Expect = 7e-12
Identities = 38/107 (35%), Positives = 52/107 (48%), Gaps = 3/107 (2%)
Frame = +1
Query: 4 AMKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPAR 60
A++K +K+ Q LKQ++ R + + D P P G VYVG R R+ +P
Sbjct: 10 AIRKSNKLPQHAVLKQILKRCSSLGKKNGYDDDGHPVDVPKGHFAVYVGENRTRYIVPIS 189
Query: 61 FLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNEN 107
FL P F LL + EEEFG GL +PCD F + L +N
Sbjct: 190 FLAHPQFQSLLRQAEEEFGYDHEMGLTIPCDEDVFRSLTSSLR*TKN 330
>TC221523 similar to UP|O49581 (O49581) Auxin induced like-protein, partial
(48%)
Length = 564
Score = 63.2 bits (152), Expect = 5e-11
Identities = 42/132 (31%), Positives = 62/132 (46%), Gaps = 31/132 (23%)
Frame = +1
Query: 6 KKVDKIRQIVRLKQVMMRWKQIS----------------------LRRHDSSDPRP---- 39
KK +KIR IVRL+Q++ +W++++ L+R S R
Sbjct: 22 KKPNKIRDIVRLQQILKKWRRVANSSKTSRSNNSNNNTSSRSINFLKRTLSISEREGGGS 201
Query: 40 -----PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCF 94
P G + V VG + RF IP +L F LL E EEEFG +G L +PC+V
Sbjct: 202 SSNVVPKGYVAVCVGVDLNRFVIPTEYLGHQAFLMLLREAEEEFGFEQTGVLRIPCEVSV 381
Query: 95 FTHIVKRLHKNE 106
F I+K + + +
Sbjct: 382 FESILKIVERKD 417
>BM093425 homologue to GP|24943206|gb auxin-regulated protein {Phaseolus
vulgaris}, partial (73%)
Length = 420
Score = 61.6 bits (148), Expect = 1e-10
Identities = 28/75 (37%), Positives = 46/75 (61%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P G L VYVG + +RF IP +L+ +F LL++ EEFG SGGL +PC++ F +++
Sbjct: 161 PKGYLAVYVGPQLRRFIIPTSYLSHSLFKALLEKAAEEFGFDQSGGLTIPCEIETFKYLL 340
Query: 100 KRLHKNENKYGKLTL 114
+ +++ KL +
Sbjct: 341 NCIENHDDSSSKLIM 385
>TC232556
Length = 666
Score = 61.6 bits (148), Expect = 1e-10
Identities = 34/93 (36%), Positives = 45/93 (47%), Gaps = 3/93 (3%)
Frame = +3
Query: 17 LKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDE 73
LKQ++ R + + D P P G VYVG R+R+ +P FL P F LL +
Sbjct: 6 LKQILKRCSSLGKKNGYDDDGHPVDVPKGHFAVYVGENRRRYIVPISFLAHPEFQSLLRQ 185
Query: 74 TEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNE 106
EEEFG GL +PCD F + L N+
Sbjct: 186 AEEEFGYDHEMGLTIPCDEVVFRSLTSSLR*NK 284
>BG725273 similar to PIR|F86331|F863 F6F9.11 protein - Arabidopsis thaliana,
partial (56%)
Length = 395
Score = 60.5 bits (145), Expect = 3e-10
Identities = 35/93 (37%), Positives = 53/93 (56%)
Frame = +2
Query: 14 IVRLKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDE 73
IVRL+Q++ RW+ S SD P+ + V VG +RF + A +LN PVF LL E
Sbjct: 56 IVRLRQMLRRWR--SKAHRIPSDV--PAVHVAVCVGTNSRRFVVRATYLNHPVFKKLLVE 223
Query: 74 TEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNE 106
EEE+G G L +PCD F +++ + +++
Sbjct: 224 AEEEYGFSNHGLLAIPCDEALFEQLLRFISRSD 322
>BQ785452
Length = 409
Score = 60.1 bits (144), Expect = 4e-10
Identities = 30/65 (46%), Positives = 39/65 (59%), Gaps = 1/65 (1%)
Frame = +3
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG +++RF IP +LN P F LL + E+E+G GGL +PC F HI
Sbjct: 213 PKGYLAVYVGDKQKRFVIPISYLNQPSFQDLLSQAEKEYGYDHPMGGLTIPCSEDVFQHI 392
Query: 99 VKRLH 103
RL+
Sbjct: 393 TSRLN 407
>TC210302
Length = 694
Score = 60.1 bits (144), Expect = 4e-10
Identities = 36/103 (34%), Positives = 49/103 (46%), Gaps = 4/103 (3%)
Frame = +2
Query: 4 AMKKVDKIR-QIVRLKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPA 59
A++K +K Q LKQ++ R + + D P P G VYVG R R+ +P
Sbjct: 20 AIRKSNKTTSQTTVLKQILKRCSSLGKKNGYDDDGLPLDVPKGHFAVYVGQNRSRYIVPI 199
Query: 60 RFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRL 102
FL P F LL + EEEFG GL +PC+ F + L
Sbjct: 200 SFLTHPEFQSLLRQAEEEFGFDHEMGLTIPCEEVVFRSLTSML 328
>BE347326 similar to SP|P32295|ARG7_P Indole-3-acetic acid induced protein
ARG7. [Mung bean Vigna radiata] {Phaseolus aureus},
partial (96%)
Length = 470
Score = 58.9 bits (141), Expect = 9e-10
Identities = 31/65 (47%), Positives = 36/65 (54%), Gaps = 1/65 (1%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN P F LL EEEFG GGL +PC F HI
Sbjct: 170 PKGYLAVYVGEKMKRFVIPVSYLNQPSFQDLLSRAEEEFGYDHPMGGLTIPCSEDVFQHI 349
Query: 99 VKRLH 103
L+
Sbjct: 350 TSCLN 364
>CD391803 similar to SP|P33079|AXA5_S Auxin-induced protein 10A5. [Soybean]
{Glycine max}, complete
Length = 514
Score = 58.9 bits (141), Expect = 9e-10
Identities = 30/66 (45%), Positives = 41/66 (61%), Gaps = 1/66 (1%)
Frame = -3
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN P F LL++ EEEFG +GGL +PC F ++
Sbjct: 350 PKGYLAVYVGDKMRRFMIPVSYLNPPSFQELLNQAEEEFGYDHPTGGLTIPCQEDEFLNV 171
Query: 99 VKRLHK 104
RL++
Sbjct: 170 TSRLNE 153
>AI965603 similar to SP|P33083|AX6B_S Auxin-induced protein 6B. [Soybean]
{Glycine max}, complete
Length = 421
Score = 58.5 bits (140), Expect = 1e-09
Identities = 36/95 (37%), Positives = 49/95 (50%), Gaps = 8/95 (8%)
Frame = +2
Query: 17 LKQVMMRWKQISLRRHD-------SSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAG 69
+KQ M ++ ++RR S + P G L VYVG ++ RF +P +LN P F
Sbjct: 74 IKQNRMGFRLPAVRRASFTASQAASKSVQVPKGYLAVYVGEKQ*RFVVPVSYLNQPSFED 253
Query: 70 LLDETEEEFGL-RGSGGLILPCDVCFFTHIVKRLH 103
LL + EEEFG GGL +PC F HI L+
Sbjct: 254 LLCQAEEEFGYDHPLGGLTIPCSEDVFQHITSHLN 358
>BQ298199 similar to SP|P33079|AXA5_S Auxin-induced protein 10A5. [Soybean]
{Glycine max}, complete
Length = 427
Score = 58.5 bits (140), Expect = 1e-09
Identities = 29/66 (43%), Positives = 40/66 (59%), Gaps = 1/66 (1%)
Frame = +2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHI 98
P G L VYVG + +RF IP +LN P F LL + +EEFG +GGL +PC F ++
Sbjct: 158 PKGYLAVYVGDKMRRFVIPVSYLNQPSFQELLSQAKEEFGYDHPTGGLTIPCQEDVFLNV 337
Query: 99 VKRLHK 104
RL++
Sbjct: 338 TSRLNE 355
>BU546668 weakly similar to GP|20161745|dbj B1033B05.12 {Oryza sativa
(japonica cultivar-group)}, partial (27%)
Length = 574
Score = 58.2 bits (139), Expect = 1e-09
Identities = 27/67 (40%), Positives = 36/67 (53%)
Frame = -2
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P GC VYVG + QR I + N P+F LL+E E E+G G L LPC V F ++
Sbjct: 405 PEGCFSVYVGPQMQRXXIKTEYANHPLFKMLLEEAESEYGYNSQGPLALPCHVDVFYKVL 226
Query: 100 KRLHKNE 106
+ +E
Sbjct: 225 MEMDSDE 205
>TC220615 homologue to UP|AX6B_SOYBN (P33083) Auxin-induced protein 6B,
complete
Length = 523
Score = 57.8 bits (138), Expect = 2e-09
Identities = 30/63 (47%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Frame = +1
Query: 42 GCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGLILPCDVCFFTHIVK 100
G L VYVG + +RF IP +LN P F LL + EEEFG +GGL +PC F HI
Sbjct: 142 GYLAVYVGEKMRRFVIPVSYLNKPSFQDLLSQAEEEFGYHHPNGGLTIPCSEDVFQHITS 321
Query: 101 RLH 103
L+
Sbjct: 322 FLN 330
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,746,345
Number of Sequences: 63676
Number of extensions: 88125
Number of successful extensions: 710
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 688
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 689
length of query: 148
length of database: 12,639,632
effective HSP length: 88
effective length of query: 60
effective length of database: 7,036,144
effective search space: 422168640
effective search space used: 422168640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0004a.3


