

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
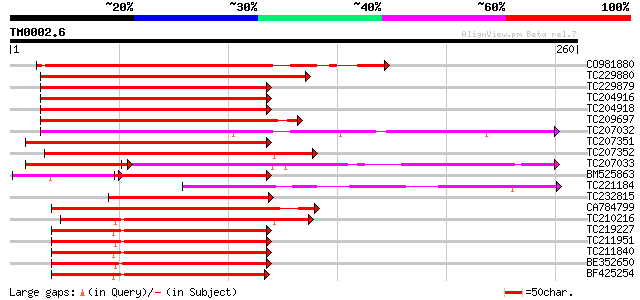
Query= TM0002.6
(260 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CO981880 220 6e-58
TC229880 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 182 2e-46
TC229879 homologue to UP|Q8H0H0 (Q8H0H0) Myb-like protein, parti... 181 3e-46
TC204916 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial... 179 1e-45
TC204918 similar to UP|O23160 (O23160) Myb-related protein (MYB ... 179 1e-45
TC209697 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial... 179 1e-45
TC207032 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial... 178 2e-45
TC207351 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucros... 171 2e-43
TC207352 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucros... 168 2e-42
TC207033 similar to UP|Q941B3 (Q941B3) AT5g67300/K8K14_2, partia... 122 3e-42
BM525863 133 3e-40
TC221184 similar to UP|O22179 (O22179) MYB family transcription ... 135 2e-32
TC232815 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucros... 130 8e-31
CA784799 124 4e-29
TC210216 homologue to UP|Q70KC5 (Q70KC5) MYB transcription facto... 112 2e-25
TC219227 homologue to UP|Q84U52 (Q84U52) MYB2, partial (36%) 111 4e-25
TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcri... 110 8e-25
TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4,... 108 2e-24
BE352650 108 3e-24
BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - gar... 106 9e-24
>CO981880
Length = 749
Score = 220 bits (560), Expect = 6e-58
Identities = 112/162 (69%), Positives = 122/162 (75%)
Frame = +1
Query: 13 GGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQH 72
GGD +RVKGSWSPQED TL+KLV +HG RNWSVISAGIPGRSGKSCRLRWCNQLSP+VQH
Sbjct: 328 GGD-NRVKGSWSPQEDATLLKLVNEHGARNWSVISAGIPGRSGKSCRLRWCNQLSPEVQH 504
Query: 73 RPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASP 132
RPFTP ED +II+AHA+HGNKWATISRLLPGRTDNAIKNHWNSTLRRRR A
Sbjct: 505 RPFTPAEDKMIIKAHAIHGNKWATISRLLPGRTDNAIKNHWNSTLRRRR-------TAXQ 663
Query: 133 SPSPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCV 174
S S S SPP AKRP L D +P K+ C+
Sbjct: 664 SSSSSASPP-----AKRP---------SSLFDTLHPLKKQCI 747
>TC229880 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (35%)
Length = 583
Score = 182 bits (461), Expect = 2e-46
Identities = 84/124 (67%), Positives = 97/124 (77%)
Frame = +1
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
D DR+KG W+P+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 97 DMDRIKGPWNPKEDEALQKLVERHGPRNWSLISRSIPGRSGKSCRLRWCNQLSPQVEHRA 276
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSP 134
FTP ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+ + + +
Sbjct: 277 FTPEEDETIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCASFMMAGDEAVAV 456
Query: 135 SPSP 138
SP P
Sbjct: 457 SPRP 468
>TC229879 homologue to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial (38%)
Length = 681
Score = 181 bits (459), Expect = 3e-46
Identities = 81/106 (76%), Positives = 91/106 (85%)
Frame = +1
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
+ DR+KG WSP+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 133 EMDRIKGPWSPEEDEALQKLVERHGPRNWSLISRSIPGRSGKSCRLRWCNQLSPQVEHRA 312
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
FTP ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 313 FTPEEDETIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK 450
>TC204916 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial (51%)
Length = 1505
Score = 179 bits (454), Expect = 1e-45
Identities = 81/106 (76%), Positives = 90/106 (84%)
Frame = +2
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
D DR+KG WSP+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 77 DMDRIKGPWSPEEDEALQKLVEKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQVEHRA 256
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
FT ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 257 FTAEEDDTIIRAHARFGNKWATIARLLHGRTDNAIKNHWNSTLKRK 394
>TC204918 similar to UP|O23160 (O23160) Myb-related protein (MYB
transcription factor), partial (41%)
Length = 565
Score = 179 bits (453), Expect = 1e-45
Identities = 81/106 (76%), Positives = 90/106 (84%)
Frame = +3
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
D DR+KG WSP+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 90 DMDRIKGPWSPEEDEALQKLVEKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPQVEHRA 269
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
FT ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 270 FTHEEDDTIIRAHARFGNKWATIARLLHGRTDNAIKNHWNSTLKRK 407
>TC209697 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial (51%)
Length = 882
Score = 179 bits (453), Expect = 1e-45
Identities = 83/120 (69%), Positives = 95/120 (79%)
Frame = +1
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
D DR+KG WSP+EDE L +LV +GPRNWSVIS IPGRSGKSCRLRWCNQLSP+V+ RP
Sbjct: 25 DMDRIKGPWSPEEDEALRRLVQTYGPRNWSVISKSIPGRSGKSCRLRWCNQLSPEVERRP 204
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSP 134
FT ED I++AHA GNKWATI+R L GRTDNAIKNHWNSTL+R+ E PL+ P P
Sbjct: 205 FTAEEDEAILKAHARFGNKWATIARFLNGRTDNAIKNHWNSTLKRKCSE----PLSEPRP 372
>TC207032 similar to UP|Q8H0H0 (Q8H0H0) Myb-like protein, partial (43%)
Length = 1001
Score = 178 bits (452), Expect = 2e-45
Identities = 109/253 (43%), Positives = 136/253 (53%), Gaps = 15/253 (5%)
Frame = +3
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
+ DRVKG WSP+EDE L +LV HGPRNWSVIS +PGRSGKSCRLRWCNQLSP V HRP
Sbjct: 87 EMDRVKGPWSPEEDEALRRLVQAHGPRNWSVISKSVPGRSGKSCRLRWCNQLSPQVAHRP 266
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLL-PGRTDNAIKNHWNSTLRRRRGESLPLPLASPS 133
F+P ED I++AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R++ ++ S
Sbjct: 267 FSPDEDEAIVRAHARFGNKWATIARLLNNGRTDNAVKNHWNSTLKRKK-------CSAVS 425
Query: 134 PSPSPSPPLPFSMAKRP-HGYELLDRYDGLMDQGYPFKRHCVEKDDEESRDGVEEAETMT 192
+ PPL S + P H L D P + + ET +
Sbjct: 426 DDVTDRPPLKRSASVGPAHLNPGSPSGSDLSDPSLP----ALSNPSPSAHYRPNLMETAS 593
Query: 193 EEEVLASSVPTSLSLFPMGEKKMVA-------------VAEKEEKKMDMQEGLMRTMMQR 239
A+S+ SL F G + A E++ M +MQ
Sbjct: 594 SACDPATSLSLSLPGFGSGSNPVCAPRPVSVSPPSPPPAEPVEQRGKQMFNAEFLRVMQE 773
Query: 240 MIAEEVRAYFDKM 252
MI +EVR+Y M
Sbjct: 774 MIRKEVRSYMSGM 812
>TC207351 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (52%)
Length = 974
Score = 171 bits (434), Expect = 2e-43
Identities = 79/113 (69%), Positives = 90/113 (78%)
Frame = +3
Query: 8 SNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLS 67
SN DR+KG WS QED L +LV Q+GPRNWS+IS I GRSGKSCRLRWCNQLS
Sbjct: 222 SNHRSPNKPDRIKGPWSAQEDRILTRLVEQYGPRNWSLISRYIKGRSGKSCRLRWCNQLS 401
Query: 68 PDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P V+HRPF+ ED II AHA +GN+WATI+RLLPGRTDNA+KNHWNSTL+RR
Sbjct: 402 PTVEHRPFSTQEDETIIAAHARYGNRWATIARLLPGRTDNAVKNHWNSTLKRR 560
>TC207352 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (58%)
Length = 811
Score = 168 bits (426), Expect = 2e-42
Identities = 80/126 (63%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WS +ED L LV ++GPRNWS+IS I GRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 124 DRIKGPWSAEEDRILTGLVERYGPRNWSLISRYIKGRSGKSCRLRWCNQLSPAVEHRPFS 303
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-RGESLPLPLASPSPS 135
ED II AHA +GN+WATI+RLLPGRTDNA+KNHWNSTL+RR +G ++ + A+ +
Sbjct: 304 AQEDDTIIAAHAQYGNRWATIARLLPGRTDNAVKNHWNSTLKRRAKGININVNHANNEVA 483
Query: 136 PSPSPP 141
S S P
Sbjct: 484 ASSSAP 501
>TC207033 similar to UP|Q941B3 (Q941B3) AT5g67300/K8K14_2, partial (42%)
Length = 1164
Score = 122 bits (307), Expect(2) = 3e-42
Identities = 87/221 (39%), Positives = 105/221 (47%), Gaps = 20/221 (9%)
Frame = +2
Query: 52 GRSGKSCRLRWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKN 111
G SGKSCRLRWCNQLSP V HRPF+ ED II AHA GNKWATI+RLL GRTDNA+KN
Sbjct: 227 GDSGKSCRLRWCNQLSPQVAHRPFSQEEDEAIIMAHAKFGNKWATIARLLNGRTDNAVKN 406
Query: 112 HWNSTLRR-----------------RRGESL---PLPLASPSPSPSPSPPLPFSMAKRPH 151
HWNSTL+R +R S+ L ASPS S P LP P
Sbjct: 407 HWNSTLKRKSSAVSDDDVVTHRQPLKRSNSVGPAHLNPASPSVSDLSDPGLPALSNPSPS 586
Query: 152 GYELLDRYDGLMDQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLASSVPTSLSLFPMG 211
+ + LM E+ + T + +VP +S P
Sbjct: 587 AHYM----PNLM----------------ETASSAPDPATSLSLSLPGFTVPQPVSPPPPP 706
Query: 212 EKKMVAVAEKEEKKMDMQEGLMRTMMQRMIAEEVRAYFDKM 252
E+ K+M E L +MQ MI +EVR+Y M
Sbjct: 707 LPLPAETVEERGKQMFNAEFL--RVMQEMIKKEVRSYMSGM 823
Score = 66.6 bits (161), Expect(2) = 3e-42
Identities = 31/49 (63%), Positives = 34/49 (69%)
Frame = +1
Query: 8 SNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGK 56
S+ G + DRVKG WSP+EDE L LV HGPRNWSVIS IPGR K
Sbjct: 94 SSSGKSREMDRVKGPWSPEEDEALRALVQAHGPRNWSVISKSIPGRFRK 240
>BM525863
Length = 439
Score = 133 bits (335), Expect(2) = 3e-40
Identities = 57/72 (79%), Positives = 65/72 (90%)
Frame = +2
Query: 49 GIPGRSGKSCRLRWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNA 108
G+PGRSGKSCRLRWCNQL P V+ +PFT ED+II+ AHA+HGNKWA I+RLLPGRTDNA
Sbjct: 206 GVPGRSGKSCRLRWCNQLDPCVKRKPFTEEEDSIIVSAHAIHGNKWAAIARLLPGRTDNA 385
Query: 109 IKNHWNSTLRRR 120
IKNHWNSTL+RR
Sbjct: 386 IKNHWNSTLKRR 421
Score = 49.3 bits (116), Expect(2) = 3e-40
Identities = 26/59 (44%), Positives = 33/59 (55%), Gaps = 8/59 (13%)
Frame = +1
Query: 2 IGGGDSSNGGGGGDRD--------RVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPG 52
+ G +S GG + D RVKG WSP+ED L +LV Q G RNWS+I+ G G
Sbjct: 40 VHNGVASTELGGEEMDIAEVTAAGRVKGPWSPEEDALLSRLVAQFGARNWSMIACGSAG 216
>TC221184 similar to UP|O22179 (O22179) MYB family transcription factor
(At2g23280) (MYB transcription factor), partial (15%)
Length = 705
Score = 135 bits (340), Expect = 2e-32
Identities = 83/191 (43%), Positives = 102/191 (52%), Gaps = 17/191 (8%)
Frame = +3
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPSPS 139
D +II+AHA+HGNKWATISRLLPGRTDNAIKNHWNSTLRRRR S S S S
Sbjct: 3 DKMIIKAHAIHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRA-------VESSSSSSAS 161
Query: 140 PPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCVEKDDEESRDGVEEAETMTEEEVLAS 199
PP R+ L D +PFK+ C+EK++EE R + S
Sbjct: 162 PP--------------AKRHSSLFDTLHPFKKQCIEKENEEER--------------VLS 257
Query: 200 SVPTSLSLFPMGEKKMVAVAEKEEKKMDMQ-----------------EGLMRTMMQRMIA 242
V TSLSLFP GEK E+E+++ + + + MMQRMIA
Sbjct: 258 PVTTSLSLFPPGEKSEEEEEEEEQEEEEKEKEELFQVNVNVNQKVTDQNYFMQMMQRMIA 437
Query: 243 EEVRAYFDKMR 253
EEVR Y + +R
Sbjct: 438 EEVRNYMETLR 470
>TC232815 similar to UP|Q9FZ13 (Q9FZ13) Tuber-specific and sucrose-responsive
element binding factor, partial (29%)
Length = 589
Score = 130 bits (326), Expect = 8e-31
Identities = 57/76 (75%), Positives = 66/76 (86%)
Frame = +1
Query: 46 ISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRT 105
IS I GRSGKSCRLRWCNQLSP V+HRPF+ ED +I+ AHA GNKWATI+R+LPGRT
Sbjct: 13 ISRHIKGRSGKSCRLRWCNQLSPTVEHRPFSTREDEVILHAHARFGNKWATIARMLPGRT 192
Query: 106 DNAIKNHWNSTLRRRR 121
DNA+KNHWN+TL+RRR
Sbjct: 193 DNAVKNHWNATLKRRR 240
>CA784799
Length = 387
Score = 124 bits (311), Expect = 4e-29
Identities = 58/123 (47%), Positives = 77/123 (62%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P EDE L +LV ++GP NW+ I+ + GRSGKSCRLRW NQL P + PFT E
Sbjct: 39 RGHWRPAEDEKLRELVERYGPHNWNAIAEKLRGRSGKSCRLRWFNQLDPRINRSPFTEEE 218
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPSPS 139
+ ++ +H +HG WA I+R PGR+DNA+KNHW+ + R R E S + SP
Sbjct: 219 EERLLASHRIHGFLWAVIARHFPGRSDNAVKNHWHVMMARIRRER--------SKNYSPK 374
Query: 140 PPL 142
PL
Sbjct: 375 QPL 383
>TC210216 homologue to UP|Q70KC5 (Q70KC5) MYB transcription factor, partial
(33%)
Length = 1252
Score = 112 bits (280), Expect = 2e-25
Identities = 58/119 (48%), Positives = 78/119 (64%), Gaps = 3/119 (2%)
Frame = +3
Query: 24 SPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTEDT 81
SP+ED+ LM + HG WS ++ AG+ R GKSCRLRW N L PD++ F+P E+
Sbjct: 9 SPEEDDKLMNYMLNHGQGCWSDVARNAGLQ-RCGKSCRLRWINYLRPDLKRGAFSPQEEE 185
Query: 82 IIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-RGESLPLPLASPSPSPSPS 139
+II H++ GN+W+ I+ LPGRTDN IKN WNST+++R R S SPSPS + S
Sbjct: 186 LIIHFHSLLGNRWSQIAARLPGRTDNEIKNFWNSTIKKRLRNMSSTTTTTSPSPSSNAS 362
>TC219227 homologue to UP|Q84U52 (Q84U52) MYB2, partial (36%)
Length = 1311
Score = 111 bits (277), Expect = 4e-25
Identities = 53/103 (51%), Positives = 72/103 (69%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WSP+EDE L++ + ++G WS + AG+ R GKSCRLRW N L PD+ F+
Sbjct: 43 KGLWSPEEDEKLLRHITKYGHGCWSSVPKQAGLQ-RCGKSCRLRWINYLRPDLNRGTFSQ 219
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+T+II+ HAV GN+W+ I+ LPGRTDN IKN WNS L+++
Sbjct: 220 EEETLIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKK 348
Score = 32.7 bits (73), Expect = 0.17
Identities = 18/61 (29%), Positives = 33/61 (53%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G++S +E+ +++L G R WS I+A +PGR+ + W + L ++ R
Sbjct: 193 DLNRGTFSQEEETLIIELHAVLGNR-WSQIAAQLPGRTDNEIKNLWNSCLKKKLRQRGID 369
Query: 77 P 77
P
Sbjct: 370 P 372
>TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcription factor
{Arabidopsis thaliana;} , partial (38%)
Length = 776
Score = 110 bits (274), Expect = 8e-25
Identities = 53/103 (51%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WSP+EDE L+ + +HG WS + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 131 KGLWSPEEDEKLLNYITKHGHGCWSSVPKLAGLQ-RCGKSCRLRWINYLRPDLKRGAFSQ 307
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ II+ HAV GN+W+ I+ LPGRTDN IKN WNS L+++
Sbjct: 308 QEENSIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKK 436
Score = 35.4 bits (80), Expect = 0.026
Identities = 19/61 (31%), Positives = 34/61 (55%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G++S QE+ ++++L G R WS I+A +PGR+ + W + L ++ R
Sbjct: 281 DLKRGAFSQQEENSIIELHAVLGNR-WSQIAAQLPGRTDNEIKNLWNSCLKKKLRQRGID 457
Query: 77 P 77
P
Sbjct: 458 P 460
>TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4, partial
(38%)
Length = 516
Score = 108 bits (271), Expect = 2e-24
Identities = 50/103 (48%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ +ED+ L+ + +HG NW + +AG+ R GKSCRLRW N L PD++ F+
Sbjct: 2 KGPWTTEEDQKLIDYIQKHGYGNWRTLPKNAGLQ-RCGKSCRLRWTNYLRPDIKRGRFSF 178
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ IIQ H++ GNKW+ I+ LPGRTDN IKN+WN+ +R+R
Sbjct: 179 EEEETIIQLHSILGNKWSAIASRLPGRTDNEIKNYWNTHIRKR 307
>BE352650
Length = 528
Score = 108 bits (269), Expect = 3e-24
Identities = 49/103 (47%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+P+EDE L+ + +HG +W + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 145 KGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGL-NRCGKSCRLRWTNYLRPDIKRGKFSE 321
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
++ III H+V GNKW+ I+ LPGRTDN IKN+W + +R++
Sbjct: 322 DDERIIINFHSVLGNKWSKIAAHLPGRTDNEIKNYWTTHIRKK 450
>BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - garden petunia,
partial (29%)
Length = 375
Score = 106 bits (265), Expect = 9e-24
Identities = 49/102 (48%), Positives = 69/102 (67%), Gaps = 2/102 (1%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+P+ED+ L+ + +HG +W + AG+ R GKSCRLRW N L PD++ F
Sbjct: 36 KGPWTPEEDQKLLAYIEEHGHGSWRALPAKAGLQ-RCGKSCRLRWTNYLRPDIKRGKFXM 212
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR 119
E+ IIQ HA+ GN+W++IS LP RTDN IKN+WN+ L++
Sbjct: 213 QEEQTIIQLHALLGNRWSSISTHLPKRTDNEIKNYWNTHLKK 338
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,928,766
Number of Sequences: 63676
Number of extensions: 225605
Number of successful extensions: 8011
Number of sequences better than 10.0: 425
Number of HSP's better than 10.0 without gapping: 4312
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6937
length of query: 260
length of database: 12,639,632
effective HSP length: 95
effective length of query: 165
effective length of database: 6,590,412
effective search space: 1087417980
effective search space used: 1087417980
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0002.6


