

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0001.6
(381 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
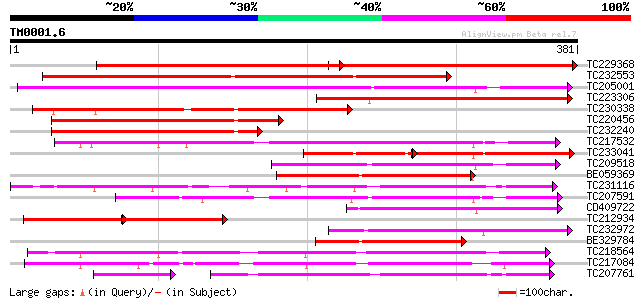
Score E
Sequences producing significant alignments: (bits) Value
TC229368 similar to PIR|T48618|T48618 early nodule-specific prot... 307 6e-84
TC232553 similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (59%) 290 7e-79
TC205001 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetyles... 276 8e-75
TC223306 weakly similar to PIR|T48618|T48618 early nodule-specif... 226 1e-59
TC230338 similar to UP|Q8W0Y6 (Q8W0Y6) Enod8.2, partial (49%) 204 7e-53
TC220456 weakly similar to UP|O81262 (O81262) Early nodule-speci... 172 3e-43
TC232240 weakly similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial ... 160 7e-40
TC217532 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E608... 156 2e-38
TC233041 weakly similar to UP|Q8W0Y7 (Q8W0Y7) Enod8.3 (Fragment)... 109 2e-38
TC209518 weakly similar to UP|O82076 (O82076) Lipase homolog (Fr... 148 5e-36
BE059369 weakly similar to PIR|T09416|T094 coil protein PO22 mi... 144 7e-35
TC231116 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrola... 131 4e-31
TC207591 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E608... 130 1e-30
CD409722 126 1e-29
TC212934 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetyles... 81 2e-29
TC232972 weakly similar to UP|O82681 (O82681) Lanatoside 15'-O-a... 115 3e-26
BE329784 112 2e-25
TC218564 110 1e-24
TC217084 109 2e-24
TC207761 weakly similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, parti... 82 5e-24
>TC229368 similar to PIR|T48618|T48618 early nodule-specific protein-like -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(61%)
Length = 1238
Score = 307 bits (786), Expect = 6e-84
Identities = 138/167 (82%), Positives = 147/167 (87%)
Frame = +3
Query: 215 GRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTE 274
G T AP GC+PV LFYKHN+P GYLD YGCVKDQNVMA EFNKQLKDRV+KLRTE
Sbjct: 474 GDTSGYTTLAPFGCMPVQLFYKHNIPEGYLDQYGCVKDQNVMATEFNKQLKDRVIKLRTE 653
Query: 275 LPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGN 334
LPEAAITYVD+YAAKY LISNTK EGFVDP+KICCGYHVNDTHIWCG +GT NGKDVFG+
Sbjct: 654 LPEAAITYVDVYAAKYALISNTKKEGFVDPMKICCGYHVNDTHIWCGNLGTDNGKDVFGS 833
Query: 335 ACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYRH 381
ACE PS Y+SWD VHYAEAANHWVANRILNGS+TDPPT ITQACYRH
Sbjct: 834 ACENPSQYISWDSVHYAEAANHWVANRILNGSYTDPPTPITQACYRH 974
Score = 301 bits (770), Expect = 4e-82
Identities = 146/167 (87%), Positives = 155/167 (92%)
Frame = +2
Query: 59 GESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNE 118
GE F KPS RDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIR+QNE
Sbjct: 5 GEGFFHKPSGRDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRKQNE 184
Query: 119 TIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQN 178
TIFQYGISPFSLD+Q VQF QFKARTKQLY+EAK E+SKLPVPEEFSKALYTFDIGQN
Sbjct: 185 TIFQYGISPFSLDIQIVQFNQFKARTKQLYEEAKAPHEKSKLPVPEEFSKALYTFDIGQN 364
Query: 179 DLSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAP 225
DLSVGFR MNFDQ+RESMPDI+NQLA+AVKNIY+ GGR FWIHNT+P
Sbjct: 365 DLSVGFRKMNFDQIRESMPDILNQLANAVKNIYQQGGRYFWIHNTSP 505
>TC232553 similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (59%)
Length = 832
Score = 290 bits (742), Expect = 7e-79
Identities = 150/277 (54%), Positives = 194/277 (69%), Gaps = 2/277 (0%)
Frame = +2
Query: 23 LKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAE 82
L S C FPAI+NFGDSNSDTGG+SAAF PP+GES+ P+ R CDGRLIVDF+A+
Sbjct: 11 LAASKQCHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGESYFHHPAGRYCDGRLIVDFLAK 190
Query: 83 KLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQY-GISPFSLDMQFVQFKQFK 141
KL LPYLSA+L+S+G+NY HGANFAT GSTIR QN T+ Q G SPFSLD+QF QF F+
Sbjct: 191 KLGLPYLSAFLDSVGSNYSHGANFATAGSTIRPQNTTLHQTGGFSPFSLDVQFNQFSDFQ 370
Query: 142 ARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIV 200
RT+ + K + ++ LP E+FS+ALYTFDIGQNDL+ G F M+ DQ++ +PD++
Sbjct: 371 RRTQFFHN--KGGVYKTLLPKAEDFSQALYTFDIGQNDLASGYFHNMSTDQVKAYVPDVL 544
Query: 201 NQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEF 260
Q + +K +Y GGR+FW+HNT P+GCLP + H + +D C N +A F
Sbjct: 545 AQFKNVIKYVYNHGGRSFWVHNTGPVGCLPY-IMDLHPVKPSLVDKAXCATPYNEVAKFF 721
Query: 261 NKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTK 297
N +LK+ VV+L ELP AAITYVD+Y+ K+ LI K
Sbjct: 722 NSKLKEVVVQLTKELPLAAITYVDVYSVKHSLIRQPK 832
>TC205001 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetylesterase
precursor, partial (62%)
Length = 1384
Score = 276 bits (707), Expect = 8e-75
Identities = 150/378 (39%), Positives = 215/378 (56%), Gaps = 5/378 (1%)
Frame = +1
Query: 6 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQK 65
+F F + C + + S+ + C F AI+NFGDSNSDTGG +F P PYG ++ +K
Sbjct: 82 IFSKFLVICMVMMISLVDSSYSLCDFEAIFNFGDSNSDTGGFHTSFPAQPAPYGMTYFKK 261
Query: 66 PSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGI 125
P R DGRLIVDF+A+ L LPYLS YL S+G++Y HGANFA+ ST+ + G+
Sbjct: 262 PVGRASDGRLIVDFLAQGLGLPYLSPYLQSIGSDYTHGANFASSASTVIPPTTSFSVSGL 441
Query: 126 SPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFR 185
SPFSL +Q Q +QFKA+ + +Q +K+P P+ F KALYTF IGQND +
Sbjct: 442 SPFSLSVQLRQMEQFKAKVDEFHQTGTRISSGTKIPSPDIFGKALYTFYIGQNDFTSKIA 621
Query: 186 MM-NFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYL 244
D +R S+P IV+Q+ +A+K +Y GGR F + N P+GC P L + + Y
Sbjct: 622 ATGGIDGVRGSLPHIVSQINAAIKELYAQGGRAFMVFNLGPVGCYPGYLVELPHATSDY- 798
Query: 245 DPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDP 304
D +GC+ N ++NK L+D + + L +A++ Y D ++A L + G
Sbjct: 799 DEFGCIVSHNNAVNDYNKLLRDTLTQTGESLVDASLIYADTHSALLELFHHPTFYGLKYN 978
Query: 305 LKICCGY----HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVAN 360
+ CCGY + + I CG + +AC++P YVSWDG+H+ EAAN VA+
Sbjct: 979 TRTCCGYGGGVYNFNPKILCG--------HMLASACDEPQNYVSWDGIHFTEAANKIVAH 1134
Query: 361 RILNGSFTDPPTLITQAC 378
ILNGS PP + + C
Sbjct: 1135AILNGSLFYPPFPLHKHC 1188
>TC223306 weakly similar to PIR|T48618|T48618 early nodule-specific
protein-like - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (25%)
Length = 669
Score = 226 bits (576), Expect = 1e-59
Identities = 102/176 (57%), Positives = 128/176 (71%), Gaps = 4/176 (2%)
Frame = +1
Query: 207 VKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLP----AGYLDPYGCVKDQNVMAVEFNK 262
V+ + LG RTFWIHNT PIGCLPV + + + AGYLD GC+ QN MA EFNK
Sbjct: 16 VQTLLGLGARTFWIHNTGPIGCLPVAMPVHNAMNTTPGAGYLDQNGCINYQNDMAREFNK 195
Query: 263 QLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGT 322
+LK+ VVKLR + P+A++ YVD+++AKY LISN EGFVDP ICCGYH + H++CG
Sbjct: 196 KLKNTVVKLRVQFPDASLIYVDMFSAKYELISNANKEGFVDPSGICCGYHQDGYHLYCGN 375
Query: 323 IGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
NGK++F + C+ PS Y+SWDGVHY EAANHW+ANRILNGSF+DPP I +C
Sbjct: 376 KAIINGKEIFADTCDDPSKYISWDGVHYTEAANHWIANRILNGSFSDPPLSIAHSC 543
>TC230338 similar to UP|Q8W0Y6 (Q8W0Y6) Enod8.2, partial (49%)
Length = 737
Score = 204 bits (518), Expect = 7e-53
Identities = 108/224 (48%), Positives = 145/224 (64%), Gaps = 9/224 (4%)
Frame = +2
Query: 16 LCVNSVELKNSPP------CAFPAIYNFGDSNSDTGGISAAFEPIPP--PYGESFPQKPS 67
LC+ + L N C FPAI+NFG SN+DTGG++A+F P P GE++ +P+
Sbjct: 83 LCIATTILNNPAMATKQYYCDFPAIFNFGASNADTGGLAASFFVAAPKSPNGETYFHRPA 262
Query: 68 ARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISP 127
R DGRLI+DF+A+ LPYLS YL+SLGTN+ GA+FAT GSTI Q + SP
Sbjct: 263 GRFSDGRLIIDFLAQSFGLPYLSPYLDSLGTNFSRGASFATAGSTIIPQQ----SFRSSP 430
Query: 128 FSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRM 186
FSL +Q+ QF++FK T+ + ++ + + +P E F +ALYTFDIGQNDL+ G F
Sbjct: 431 FSLGVQYSQFQRFKPTTQFIREQG--GVFATLMPKEEYFHEALYTFDIGQNDLTAGFFGN 604
Query: 187 MNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP 230
M Q ++PDI+ S +KNIY +G R+FWIHNT PIGCLP
Sbjct: 605 MTLQQFNATIPDIIKSFTSNIKNIYNMGARSFWIHNTGPIGCLP 736
>TC220456 weakly similar to UP|O81262 (O81262) Early nodule-specific protein,
partial (43%)
Length = 587
Score = 172 bits (435), Expect = 3e-43
Identities = 86/156 (55%), Positives = 109/156 (69%)
Frame = +1
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+NFGDSNSDTGG++A+ PPYGE++ +P+ R DGRL++DFIA+ LPY
Sbjct: 118 CVFPAIFNFGDSNSDTGGLAASLIAPTPPYGETYFHRPAGRFSDGRLVIDFIAKSFGLPY 297
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LSAYL+SLGTN+ HGANFAT STIR I Q G SPF LD+Q+ QF+ K+RT+ +
Sbjct: 298 LSAYLDSLGTNFSHGANFATSASTIRLPTSIIPQGGFSPFYLDIQYTQFRDSKSRTQFIR 477
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGF 184
+ + S +P E F KALYTFDIGQ L + F
Sbjct: 478 HQG--GVFASLMPKEEYFDKALYTFDIGQKILVLAF 579
>TC232240 weakly similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (38%)
Length = 469
Score = 160 bits (406), Expect = 7e-40
Identities = 82/142 (57%), Positives = 100/142 (69%)
Frame = +2
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+N GDSNSDTGG+SAAF PPP G ++ P+ R DGRLI+DFIAE L Y
Sbjct: 50 CIFPAIFNLGDSNSDTGGLSAAFGQAPPPNGITYFHSPNGRFSDGRLIIDFIAESSGLAY 229
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
L AYL+S+ +N+ HGANFAT GST+R QN TI Q G SP SLD+QFVQF FK R+K +
Sbjct: 230 LRAYLDSVASNFTHGANFATAGSTVRPQNTTISQSGYSPISLDVQFVQFSDFKTRSKLVR 409
Query: 149 QEAKTALERSKLPVPEEFSKAL 170
Q+ + + LP E FS+AL
Sbjct: 410 QQG--GVFKELLPKEEYFSQAL 469
>TC217532 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E60850), partial
(54%)
Length = 1372
Score = 156 bits (394), Expect = 2e-38
Identities = 113/360 (31%), Positives = 167/360 (46%), Gaps = 20/360 (5%)
Frame = +1
Query: 31 FPAIYNFGDSNSDTGG---ISAAFEP--IPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
+ ++++FGDS +DTG IS P + PPYG++ +P+ R DGRLI+DF+AE L
Sbjct: 112 YTSLFSFGDSLTDTGNLYFISPRQSPDCLLPPYGQTHFHRPNGRCSDGRLILDFLAESLG 291
Query: 86 LPYLSAYLNSLGT-----NYRHGANFATGGSTIRRQN---ETIFQYGISP-FSLDMQFVQ 136
LPY+ YL N G NFA G+T + E F ++ FSL +Q
Sbjct: 292 LPYVKPYLGFKNGAVKRGNIEQGVNFAVAGATALDRGFFEEKGFAVDVTANFSLGVQLDW 471
Query: 137 FKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMN-FDQMRES 195
FK+ K + S V E IG ND F +
Sbjct: 472 FKELLPSLCNSSSSCKKVIGSSLFIVGE----------IGGNDYGYPLSETTAFGDLVTY 621
Query: 196 MPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNV 255
+P +++ + SA++ + +LG TF + + P+GC P L + D GC+K N
Sbjct: 622 IPQVISVITSAIRELIDLGAVTFMVPGSLPLGCNPAYLTIFATIDKEEYDQAGCLKWLNT 801
Query: 256 MAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGF-VDPLKICCG---- 310
N+ L+ + +LR P I Y D + A ++ + GF + LK+CCG
Sbjct: 802 FYEYHNELLQIEINRLRVLYPLTNIIYADYFNAALEFYNSPEQFGFGGNVLKVCCGGGGP 981
Query: 311 YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDP 370
Y+ N+T + CG G AC+ PS YVSWDG H EAA W+ +L+G +T P
Sbjct: 982 YNYNETAM-CGDAGVV--------ACDDPSQYVSWDGYHLTEAAYRWMTKGLLDGPYTIP 1134
>TC233041 weakly similar to UP|Q8W0Y7 (Q8W0Y7) Enod8.3 (Fragment), partial
(35%)
Length = 748
Score = 109 bits (273), Expect(2) = 2e-38
Identities = 55/114 (48%), Positives = 72/114 (62%), Gaps = 5/114 (4%)
Frame = +3
Query: 271 LRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCG----YHVNDTHIWCGTIGTA 326
LR ELP AAITYVD+Y KY LIS+ GF + CCG Y+ N+T CG
Sbjct: 255 LRKELPGAAITYVDVYTVKYTLISHAHKYGFEQGVIACCGHGGKYNFNNTER-CGATKRV 431
Query: 327 NGKD-VFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACY 379
NG + V N+C+ S+ + WDG+HY EAAN W+ +I+NGSF+DPP + +ACY
Sbjct: 432 NGTEIVIANSCKDLSVRIIWDGIHYTEAANKWIFQQIVNGSFSDPPHSLKRACY 593
Score = 67.4 bits (163), Expect(2) = 2e-38
Identities = 31/77 (40%), Positives = 50/77 (64%)
Frame = +2
Query: 198 DIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMA 257
D++ Q ++ +K +Y GG +FWIHNT P+GCLP + ++ + +D +GC K N +A
Sbjct: 38 DVLGQXSNVIKGVYGEGGXSFWIHNTGPLGCLPY-MLDRYPMKPTQMDEFGCAKPFNEVA 214
Query: 258 VEFNKQLKDRVVKLRTE 274
FN++LK+ VV+L E
Sbjct: 215 QYFNRKLKE-VVELTAE 262
>TC209518 weakly similar to UP|O82076 (O82076) Lipase homolog (Fragment),
partial (56%)
Length = 874
Score = 148 bits (373), Expect = 5e-36
Identities = 77/199 (38%), Positives = 115/199 (57%), Gaps = 5/199 (2%)
Frame = +1
Query: 177 QNDLSVGF-RMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFY 235
QNDL+ F + +++ Q+ + +P ++ ++ +AVKN+Y G R FW+HNT P+GCLP L
Sbjct: 1 QNDLADSFAKNLSYAQVIKKIPAVITEIENAVKNLYNDGARKFWVHNTGPLGCLPKVLAL 180
Query: 236 KHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISN 295
LD GC+ N A FN+ L KLR+E +A + YVD+YA KY LI+N
Sbjct: 181 AQKKD---LDSLGCLSSYNSAARLFNEALLHSSQKLRSEFKDATLVYVDIYAIKYDLITN 351
Query: 296 TKNEGFVDPLKICCGY----HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYA 351
GF +PL +CCGY + D + CG G C++ + YVSWDG+H
Sbjct: 352 AAKYGFSNPLMVCCGYGGPPYNFDVRVTCGQPGY--------QVCDEGARYVSWDGIHQT 507
Query: 352 EAANHWVANRILNGSFTDP 370
EAAN +A++IL+ +++ P
Sbjct: 508 EAANTLIASKILSMAYSTP 564
>BE059369 weakly similar to PIR|T09416|T094 coil protein PO22
microspore/pollen-specific - alfalfa, partial (15%)
Length = 411
Score = 144 bits (363), Expect = 7e-35
Identities = 69/138 (50%), Positives = 95/138 (68%), Gaps = 4/138 (2%)
Frame = +1
Query: 180 LSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNL 239
L+ G + + +Q+ +S+P+I+NQ AV+ +Y + R FWIHNT PIGCLP + Y +
Sbjct: 1 LAFGLQHTSQEQVIKSIPEILNQFFQAVQQLYNVRARVFWIHNTGPIGCLPYSYIY-YEP 177
Query: 240 PAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNE 299
G +D GCVK QN +A EFN+QLKD+V +LR + P A TYVD+Y AKY LI+NT+N+
Sbjct: 178 KKGNIDANGCVKPQNDLAQEFNRQLKDQVFQLRRKFPLAKFTYVDVYTAKYELINNTRNQ 357
Query: 300 GFVDPLKICC----GYHV 313
GFV PL+ CC GYH+
Sbjct: 358 GFVSPLEFCCGSYYGYHI 411
>TC231116 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrolase-like
protein, partial (94%)
Length = 1541
Score = 131 bits (330), Expect = 4e-31
Identities = 110/386 (28%), Positives = 166/386 (42%), Gaps = 18/386 (4%)
Frame = +1
Query: 1 MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIP---PP 57
MG F+A FL L + + N P PAI+ FGDS D G + + PP
Sbjct: 364 MGYSRSFLASFLLAVL----LNVTNGQPLV-PAIFTFGDSIVDVGNNNHQLTIVKANFPP 528
Query: 58 YGESFPQK-PSARDCDGRLIVDFIAEKLNLP-YLSAYLN--SLGTNYRHGANFATGGSTI 113
YG F P+ R C+G+L DFIA+ L Y AYLN + G N +GANFA+ S
Sbjct: 529 YGRDFENHFPTGRFCNGKLATDFIADILGFTSYQPAYLNLKTKGKNLLNGANFASASSGY 708
Query: 114 RRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERS--KLPVPEEFSKALY 171
+ Y P S +++ Y+E +T L + + S A+Y
Sbjct: 709 FELTSKL--YSSIPLSKQLEY-------------YKECQTKLVEAAGQSSASSIISDAIY 843
Query: 172 TFDIGQNDLSVGF-------RMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTA 224
G +D + ++ DQ +++ + ++ ++++Y LG R + +
Sbjct: 844 LISAGTSDFVQNYYINPLLNKLYTTDQFSDTL---LRCYSNFIQSLYALGARRIGVTSLP 1014
Query: 225 PIGCLP--VNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITY 282
PIGCLP + LF H CV N A+ FN++L L+ LP +
Sbjct: 1015PIGCLPAVITLFGAHINE--------CVTSLNSDAINFNEKLNTTSQNLKNMLPGLNLVV 1170
Query: 283 VDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMY 342
D+Y Y L + GF + K CCG + + I C N K + C S Y
Sbjct: 1171FDIYQPLYDLATKPSENGFFEARKACCGTGLIEVSILC------NKKSI--GTCANASEY 1326
Query: 343 VSWDGVHYAEAANHWVANRILNGSFT 368
V WDG H +EAAN +A+ ++ +
Sbjct: 1327VFWDGFHPSEAANKVLADELITSGIS 1404
>TC207591 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E60850), partial
(57%)
Length = 1290
Score = 130 bits (327), Expect = 1e-30
Identities = 98/313 (31%), Positives = 146/313 (46%), Gaps = 13/313 (4%)
Frame = +1
Query: 72 DGRLIVDFIAEKLNLPYLSAYLN-SLGTNYRHGANFATGGSTIRRQNETIFQYGISPF-- 128
DGRL++DFIAE +LPYL YL + + + G NFA G+T + + G++ +
Sbjct: 7 DGRLMIDFIAEAYDLPYLPPYLALTKDKDIQRGVNFAVAGATAL-DAKFFIEAGLAKYLW 183
Query: 129 ---SLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFR 185
SL++Q FK+ K Q+ + +RS V E IG ND +
Sbjct: 184 TNNSLNIQLGWFKKLKPSLCTTKQDCDSYFKRSLFLVGE----------IGGNDYNYAAI 333
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGC--LPVNLFYKHNLPAGY 243
N Q++ ++P +V + +A+ + G R + PIGC L + LF N
Sbjct: 334 AGNITQLQATVPPVVEAITAAINELIAEGARELLVPGNFPIGCSALYLTLFRSENKED-- 507
Query: 244 LDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVD 303
D GC+K N A NK+LK + LR + P A I Y D Y A + GF +
Sbjct: 508 YDESGCLKTFNGFAEYHNKELKLALETLRKKNPHARILYADYYGAAKRFFHAPGHHGFTN 687
Query: 304 -PLKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWV 358
L+ CCG ++ N + CG G+ AC PS Y +WDG+H EAA ++
Sbjct: 688 GALRACCGGGGPFNFNIS-ARCGHTGS--------KACADPSTYANWDGIHLTEAAYRYI 840
Query: 359 ANRILNGSFTDPP 371
A ++ G F+ PP
Sbjct: 841 AKGLIYGPFSYPP 879
>CD409722
Length = 605
Score = 126 bits (317), Expect = 1e-29
Identities = 63/149 (42%), Positives = 86/149 (57%), Gaps = 4/149 (2%)
Frame = -2
Query: 227 GCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLY 286
GCLP N+ K + LD GCV N A FN QL+ KL+ + P++ +TYVD++
Sbjct: 604 GCLPQNIA-KFGTDSSKLDGLGCVSSHNQAAKTFNLQLRALCTKLQGQYPDSNVTYVDIF 428
Query: 287 AAKYGLISNTKNEGFVDPLKICCGYH---VN-DTHIWCGTIGTANGKDVFGNACEKPSMY 342
K LI+N GF P+ CCGY +N D+ + CG T NG + AC S Y
Sbjct: 427 TIKSSLIANYSRYGFEQPIMACCGYGGPPLNYDSRVSCGETKTFNGTTITAKACNDSSEY 248
Query: 343 VSWDGVHYAEAANHWVANRILNGSFTDPP 371
+SWDG+HY E AN +VA++IL G ++DPP
Sbjct: 247 ISWDGIHYTETANQYVASQILTGKYSDPP 161
>TC212934 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetylesterase
precursor, partial (30%)
Length = 442
Score = 81.3 bits (199), Expect(2) = 2e-29
Identities = 36/71 (50%), Positives = 53/71 (73%)
Frame = +3
Query: 76 IVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFV 135
++DF+A+ L LP+LS YL S+G++Y+HGAN+AT ST+ N ++F GISPFSL +Q
Sbjct: 225 LLDFLAQALGLPFLSPYLQSIGSDYKHGANYATMASTVLMPNTSLFVTGISPFSLAIQLN 404
Query: 136 QFKQFKARTKQ 146
Q KQFK + ++
Sbjct: 405 QMKQFKTKVEE 437
Score = 65.9 bits (159), Expect(2) = 2e-29
Identities = 33/70 (47%), Positives = 44/70 (62%)
Frame = +2
Query: 10 FFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSAR 69
FF+ T+ + + + C F AI+NFGDSNSDTGG AAF PYG ++ +KP+ R
Sbjct: 26 FFVIVTIVLLCLFSLSHSECNFKAIFNFGDSNSDTGGFYAAFPGESGPYGMTYFKKPAGR 205
Query: 70 DCDGRLIVDF 79
DGRLI+ F
Sbjct: 206ASDGRLIIGF 235
>TC232972 weakly similar to UP|O82681 (O82681) Lanatoside
15'-O-acetylesterase precursor, partial (45%)
Length = 570
Score = 115 bits (289), Expect = 3e-26
Identities = 60/169 (35%), Positives = 88/169 (51%), Gaps = 5/169 (2%)
Frame = +3
Query: 215 GRTFWIHNTAPIGCLPVNLF-YKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRT 273
GRTF + N AP+GC P L + H+ + +D +GC+ N + +N LK+ + + R
Sbjct: 3 GRTFMVLNLAPVGCYPAFLVEFPHD--SSNIDDFGCLISYNNAVLNYNNMLKETLKQTRE 176
Query: 274 ELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTH----IWCGTIGTANGK 329
L +A++ YVD ++ L + + G K CCGY D + + CG NG
Sbjct: 177 SLSDASVIYVDTHSVLLELFQHPTSHGLQYGTKACCGYGGGDYNFDPKVSCGNTKEINGS 356
Query: 330 DVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
+ C P YVSWDG+H EAAN + ILNGSF+DPP + + C
Sbjct: 357 IMPATTCNDPYNYVSWDGIHSTEAANKLITFAILNGSFSDPPFIFQEHC 503
>BE329784
Length = 454
Score = 112 bits (281), Expect = 2e-25
Identities = 54/102 (52%), Positives = 70/102 (67%)
Frame = +1
Query: 206 AVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLK 265
A + +Y +G R FWIHNT PIGCLP + Y + G +D GCVK QN +A EFN+QLK
Sbjct: 1 AFQQLYNVGARVFWIHNTGPIGCLPYSYIY-YEPKKGNIDANGCVKPQNDLAQEFNRQLK 177
Query: 266 DRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKI 307
D+V +LR + P A TYVD+Y AKY LI+NT+N+G LK+
Sbjct: 178 DQVFQLRRKFPLAKFTYVDVYTAKYELINNTRNQGGRQVLKV 303
>TC218564
Length = 1398
Score = 110 bits (274), Expect = 1e-24
Identities = 95/362 (26%), Positives = 158/362 (43%), Gaps = 11/362 (3%)
Frame = +1
Query: 13 SCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGG---ISAAFEPIPPPYGESFP-QKPSA 68
S + V + E KN+ PA+ FGDS+ D+G I+ + PYG F +P+
Sbjct: 43 SLLVAVTTSEAKNN----VPAVIVFGDSSVDSGNNNVIATVLKSNFKPYGRDFEGDRPTG 210
Query: 69 RDCDGRLIVDFIAEKLNLPY-LSAYLNSLGT--NYRHGANFATGGSTIRRQNETIFQYGI 125
R C+GR+ DFIAE + + AYL+ T ++ G FA+ G+ N T +
Sbjct: 211 RFCNGRVPPDFIAEAFGIKRTVPAYLDPAYTIQDFATGVCFASAGTGY--DNATSAVLNV 384
Query: 126 SPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFR 185
P ++++ + Q K RT ++A + S+ALY +G ND +
Sbjct: 385 IPLWKEIEYYKEYQAKLRTHLGVEKANKII-----------SEALYLMSLGTNDFLENYY 531
Query: 186 MMNFDQMRESMP---DIVNQLA-SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPA 241
+ ++ ++ D + ++A + V+ +Y LG R I P+GCLP+
Sbjct: 532 VFPTRRLHFTVSQYQDFLLRIAENFVRELYALGVRKLSITGLGPVGCLPLER------AT 693
Query: 242 GYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGF 301
L +GC ++ N +A+ FN++L++ + KL ELP + Y+ +I+ GF
Sbjct: 694 NILGDHGCNQEYNDVALSFNRKLENVITKLNRELPRLKALSANAYSIVNDIITKPSTYGF 873
Query: 302 VDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANR 361
K CC + C D C YV WD H E N V++
Sbjct: 874 EVVEKACCSTGTFEMSYLC--------SDKNPLTCTDAEKYVFWDAFHPTEKTNRIVSSY 1029
Query: 362 IL 363
++
Sbjct: 1030LI 1035
>TC217084
Length = 1310
Score = 109 bits (273), Expect = 2e-24
Identities = 103/374 (27%), Positives = 158/374 (41%), Gaps = 18/374 (4%)
Frame = +3
Query: 11 FLSCTLCVNSVELKNSPPC-AFPAIYNFGDSNSDTGG---ISAAFEPIPPPYG-ESFPQK 65
FL L + + + +P A A + FGDS D G ++ PYG +S +
Sbjct: 96 FLCLCLLITLISIIVAPQAEAARAFFVFGDSLVDNGNNNFLATTARADSYPYGIDSASHR 275
Query: 66 PSARDCDGRLIVDFIAEKLN----LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIF 121
S R +G + D I+EK+ LPYLS LN G GANFA+ G I N+T
Sbjct: 276 ASGRFSNGLNMPDLISEKIGSEPTLPYLSPQLN--GERLLVGANFASAGIGIL--NDTGI 443
Query: 122 QYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLS 181
Q+ I+ + Q FKQ++ R L E +T +KAL +G ND
Sbjct: 444 QF-INIIRITEQLAYFKQYQQRVSALIGEEQTR---------NLVNKALVLITLGGNDFV 593
Query: 182 VGFRMMNFD-QMRE-SMPD----IVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFY 235
+ ++ F + RE ++PD ++++ + N+YELG R + T P+GC+P L
Sbjct: 594 NNYYLVPFSARSREYALPDYVVFLISEYRKILANLYELGARRVLVTGTGPLGCVPAEL-- 767
Query: 236 KHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISN 295
A + C + FN QL + +L T++ + + +SN
Sbjct: 768 -----AMHSQNGECATELQRAVNLFNPQLVQLLHELNTQIGSDVFISANAFTMHLDFVSN 932
Query: 296 TKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDV---FGNACEKPSMYVSWDGVHYAE 352
+ GFV CCG G NG + N C +Y WD H +E
Sbjct: 933 PQAYGFVTSKVACCGQ------------GAYNGIGLCTPASNLCPNRDLYAFWDPFHPSE 1076
Query: 353 AANHWVANRILNGS 366
AN + ++ + GS
Sbjct: 1077RANRLIVDKFMTGS 1118
>TC207761 weakly similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, partial (46%)
Length = 1609
Score = 82.0 bits (201), Expect(2) = 5e-24
Identities = 64/232 (27%), Positives = 102/232 (43%), Gaps = 1/232 (0%)
Frame = +1
Query: 136 QFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRES 195
Q FK +K L QE A + L +KA+Y +IG ND V + E
Sbjct: 727 QLSYFKKVSKILSQELGDAETTTLL------AKAVYLINIGSNDYLVSLTENSSVFTAEK 888
Query: 196 MPD-IVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQN 254
D +V L + +K I++ GGR F + N + +GC+P+ + CV++ +
Sbjct: 889 YVDMVVGNLTTVIKGIHKTGGRKFGVLNQSALGCIPLVKALLNGSKGS------CVEEAS 1050
Query: 255 VMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN 314
+A N L + KL+ +L +YVD + + L++N G + CCG
Sbjct: 1051ALAKLHNGVLSVELEKLKKQLEGFKYSYVDFFNLSFDLMNNPSKYGLKEGGMACCGSGPY 1230
Query: 315 DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGS 366
+ CG G KD CE PS YV +D +H E N ++ + +G+
Sbjct: 1231RRYYSCG--GKRAVKDY--ELCENPSDYVFFDSIHPTERFNQIISQLMWSGN 1374
Score = 47.0 bits (110), Expect(2) = 5e-24
Identities = 25/55 (45%), Positives = 30/55 (54%)
Frame = +3
Query: 57 PYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGS 111
PYGE+F PS R DGR+I D IA+ LP YL Y G NFA+ G+
Sbjct: 507 PYGETFFNYPSGRFSDGRVIPDLIADYAKLPLSPPYLFPGYQRYLDGVNFASAGA 671
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,712,885
Number of Sequences: 63676
Number of extensions: 258141
Number of successful extensions: 1574
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 1454
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1464
length of query: 381
length of database: 12,639,632
effective HSP length: 99
effective length of query: 282
effective length of database: 6,335,708
effective search space: 1786669656
effective search space used: 1786669656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0001.6


