|
|
|||||
| identifier | direction | position | No. of exon |
No. of EST |
Definition of the most similar sequence |
|---|---|---|---|---|---|
|
|
|||||
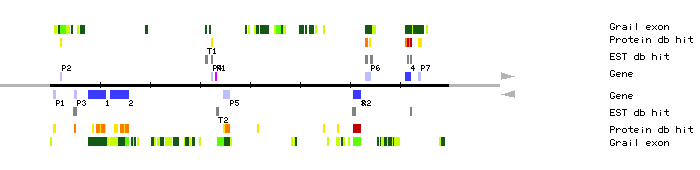
| MBK23.1 | 7617..11018 | 4 | 0 | TMV resistance protein N | |
| MBK23.2 | 12190..15740 | 4 | 0 | TMV resistance protein N | |
| MBK23.3 | 60605..62065 | 1 | 2 | phosphogluconate dehydrogenase | |
| MBK23.4 | 71027..72175 | 4 | 23 | ubiquitin-conjugating enzyme e2-17 kd 8 (EC 6.3.2.19) | |
|
|
|||||
|
|
|||||
| identifier | direction | position | No. of exon |
No. of EST |
Definition of the most similar sequence |
|---|---|---|---|---|---|
|
|
|||||
| MBK23.P1 | 749..1006 | 1 | 0 | reverse transcriptase | |
| MBK23.P2 | 2094..2204 | 1 | 0 | Caenorhabditis elegans cosmid C26E6 | |
| MBK23.P3 | 4991..5254 | 2 | 3 | 40s ribosomal protein s10 | |
| MBK23.P4 | 32203..32370 | 1 | 1 | Gallus gallus chS-Rex-s | |
| MBK23.P5 | 34643..35820 | 4 | 0 | hypothetical 97.1 kd protein in bck1-tok1 intergenic region | |
| MBK23.P6 | 63191..64162 | 2 | 2 | Petunia inflata receptor kinase gene exons 1-3 | |
| MBK23.P7 | 73709..74518 | 2 | 0 | hypothetical protein 612 | |
|
|
|||||
|
|
|||||
| identifier | direction | position | Definition of the most similar sequence | ||
|---|---|---|---|---|---|
|
|
|||||
| MBK23.T1 | 31132..31404 | clone 134I23T7 clone 228A6T7 | |||
| MBK23.T2 | 33331..33657 | clone YAP143T7, 3' end | |||
|
|
|||||
|
|
|||||
| identifier | direction | position | No. of exon |
No. of EST |
Definition of the most similar sequence |
|---|---|---|---|---|---|
|
|
|||||
| MBK23.R1 | 33121..33204 | 1 | 0 | tRNA-Leu(CAA) | |
| MBK23.R2 | 62462..62391 | 1 | 0 | tRNA-Met(CAU) | |
|
|
|||||
Structural Analysis of Arabidopsis thaliana Chromosome 5. I. Sequence Features of the 1.6 Mb Regions Covered by Physically assigned Twenty P1 Clones. (1997) Shusei Sato, Hirokazu Kotani, Yasukazu Nakamura, Takakazu Kaneko, Erika Asamizu, Masanobu Fukami, Nobuyuki Miyajima, and Satoshi Tabata DNA Research, 4, 215-230
 Kazusa Status
Kazusa Status