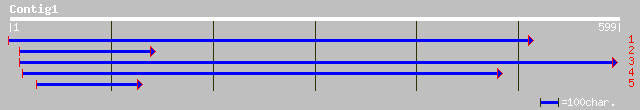
Nr search
BLASTX 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= KMC011365A_C01 KMC011365A_c01
(598 letters)
Database: nr
1,393,205 sequences; 448,689,247 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_181271.2| unknown protein; protein id: At2g37340.1, suppo... 296 1e-79
emb|CAB67657.2| splicing factor-like protein [Arabidopsis thaliana] 295 3e-79
emb|CAC03605.1| splicing factor RSZ33 [Arabidopsis thaliana] 295 3e-79
gb|AAO16998.1| Unknown protein [Oryza sativa (japonica cultivar-... 270 1e-71
pir||T45890 splicing factor-like protein - Arabidopsis thaliana 256 2e-67
>ref|NP_181271.2| unknown protein; protein id: At2g37340.1, supported by cDNA:
gi_18252178 [Arabidopsis thaliana]
gi|18252179|gb|AAL61922.1| unknown protein [Arabidopsis
thaliana] gi|21386957|gb|AAM47882.1| unknown protein
[Arabidopsis thaliana]
Length = 290
Score = 296 bits (759), Expect = 1e-79
Identities = 141/179 (78%), Positives = 153/179 (84%)
Frame = +1
Query: 61 MPRYDDKYGNTRLYVGRLSSRTRSRDLERVFSRYGRVRDVDMKRDYAFVEFSDPRDADDA 240
MPRYDD+YGNTRLYVGRLSSRTR+RDLER+FSRYGRVRDVDMKRDYAFVEF DPRDADDA
Sbjct: 1 MPRYDDRYGNTRLYVGRLSSRTRTRDLERLFSRYGRVRDVDMKRDYAFVEFGDPRDADDA 60
Query: 241 RYNLDGRDVDGSRLIVEFAKGVPRGSREGGGGRDRDREYMGRGPPPGSGRCFNCGIDGHW 420
R+ LDGRD DGSR+ VEF++G PRGS R++ RGPPPG+GRCFNCG+DGHW
Sbjct: 61 RHYLDGRDFDGSRITVEFSRGAPRGS----------RDFDSRGPPPGAGRCFNCGVDGHW 110
Query: 421 ARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSRHPRSDSRSPVRSRSPRRGRSRDRS 597
ARDC AGDWKNKCYRCGERGHIE+NCKNSPKKL R S SRSPVRSRSPRR RS RS
Sbjct: 111 ARDCTAGDWKNKCYRCGERGHIERNCKNSPKKL-RRSGSYSRSPVRSRSPRRRRSPSRS 168
>emb|CAB67657.2| splicing factor-like protein [Arabidopsis thaliana]
Length = 284
Score = 295 bits (755), Expect = 3e-79
Identities = 141/180 (78%), Positives = 152/180 (84%), Gaps = 1/180 (0%)
Frame = +1
Query: 61 MPRYDDKYGNTRLYVGRLSSRTRSRDLERVFSRYGRVRDVDMKRDYAFVEFSDPRDADDA 240
MPRYDD+YGNTRLYVGRLSSRTR+RDLER+FSRYGRVRDVDMKRDYAFVEFSDPRDADDA
Sbjct: 1 MPRYDDRYGNTRLYVGRLSSRTRTRDLERLFSRYGRVRDVDMKRDYAFVEFSDPRDADDA 60
Query: 241 RYNLDGRDVDGSRLIVEFAKGVPRGSREGGGGRDRDREYMGRGPPPGSGRCFNCGIDGHW 420
RY LDGRD DGSR+ VE ++G PRGSR+ G RGPPPGSGRCFNCG+DGHW
Sbjct: 61 RYYLDGRDFDGSRITVEASRGAPRGSRDNG----------SRGPPPGSGRCFNCGVDGHW 110
Query: 421 ARDCKAGDWKNKCYRCGERGHIEKNCKNSPK-KLSRHPRSDSRSPVRSRSPRRGRSRDRS 597
ARDC AGDWKNKCYRCGERGHIE+NCKNSP K +R S SRSPV+SRSPRR RS RS
Sbjct: 111 ARDCTAGDWKNKCYRCGERGHIERNCKNSPSPKKARQGGSYSRSPVKSRSPRRRRSPSRS 170
>emb|CAC03605.1| splicing factor RSZ33 [Arabidopsis thaliana]
Length = 290
Score = 295 bits (755), Expect = 3e-79
Identities = 140/179 (78%), Positives = 152/179 (84%)
Frame = +1
Query: 61 MPRYDDKYGNTRLYVGRLSSRTRSRDLERVFSRYGRVRDVDMKRDYAFVEFSDPRDADDA 240
MPRYDD+YGNTRLYVGRLSSRTR+RDLER+FSRYGRVRDVDMKRDYAFVEF DPRDADDA
Sbjct: 1 MPRYDDRYGNTRLYVGRLSSRTRTRDLERLFSRYGRVRDVDMKRDYAFVEFGDPRDADDA 60
Query: 241 RYNLDGRDVDGSRLIVEFAKGVPRGSREGGGGRDRDREYMGRGPPPGSGRCFNCGIDGHW 420
R+ LDGRD DGSR+ VEF++G PRGS R++ RGPPPG+GRCFNCG+DGHW
Sbjct: 61 RHYLDGRDFDGSRITVEFSRGAPRGS----------RDFDSRGPPPGAGRCFNCGVDGHW 110
Query: 421 ARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSRHPRSDSRSPVRSRSPRRGRSRDRS 597
ARDC AGDWKNKCYRCGERGHIE+NCKN PKKL R S SRSPVRSRSPRR RS RS
Sbjct: 111 ARDCTAGDWKNKCYRCGERGHIERNCKNQPKKL-RRSGSYSRSPVRSRSPRRRRSPSRS 168
>gb|AAO16998.1| Unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 389
Score = 270 bits (690), Expect = 1e-71
Identities = 137/223 (61%), Positives = 152/223 (67%), Gaps = 46/223 (20%)
Frame = +1
Query: 61 MPRYDDKYGNTRLYVGRLSSRTRSRDLERVFSRYGR------------------------ 168
MPRYDD YG+TRLYVGRLSSRTRSRDLE +F RYGR
Sbjct: 46 MPRYDDHYGSTRLYVGRLSSRTRSRDLEYLFGRYGRSCVFDLNDLVEGFSVFYTTREWKL 105
Query: 169 ----------------------VRDVDMKRDYAFVEFSDPRDADDARYNLDGRDVDGSRL 282
+R+V++KRDYAF+EFSDPRDAD+ARYNLDGRDVDGSR+
Sbjct: 106 QVGDAFEYLSHDGLLFGNHWCRIREVELKRDYAFIEFSDPRDADEARYNLDGRDVDGSRI 165
Query: 283 IVEFAKGVPRGSREGGGGRDRDREYMGRGPPPGSGRCFNCGIDGHWARDCKAGDWKNKCY 462
+VEFAKGVPRG+ G REYMGRGPPPG+GRCFNCGIDGHWARDCKAGDWKNKCY
Sbjct: 166 LVEFAKGVPRGAAGGS------REYMGRGPPPGTGRCFNCGIDGHWARDCKAGDWKNKCY 219
Query: 463 RCGERGHIEKNCKNSPKKLSRHPRSDSRSPVRSRSPRRGRSRD 591
RCGERGHIE+NC+NSP+ LSR SRS S SPR R RD
Sbjct: 220 RCGERGHIERNCQNSPRNLSR-----SRSRSLSGSPRGRRDRD 257
>pir||T45890 splicing factor-like protein - Arabidopsis thaliana
Length = 302
Score = 256 bits (654), Expect = 2e-67
Identities = 129/198 (65%), Positives = 148/198 (74%), Gaps = 19/198 (9%)
Frame = +1
Query: 61 MPRYDDKYGNTRLYVGRLSSRTRSRDLERVFSRYGR---------VRDVDMKRDY----- 198
MPRYDD+YGNTRLYVGRLSSRTR+RDLER+FSRYGR + +++M ++
Sbjct: 1 MPRYDDRYGNTRLYVGRLSSRTRTRDLERLFSRYGRGYIPKVSGGLPNLEMFDEWVDCGV 60
Query: 199 ----AFVEFSDPRDADDARYNLDGRDVDGSRLIVEFAKGVPRGSREGGGGRDRDREYMGR 366
+ +EFSDPRDADDARY LDGRD DGSR+ VE ++G PRGSR+ G R
Sbjct: 61 IGGNSRLEFSDPRDADDARYYLDGRDFDGSRITVEASRGAPRGSRDNGS----------R 110
Query: 367 GPPPGSGRCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPK-KLSRHPRSDS 543
GPPPGSGRCFNCG+DGHWARDC AGDWKNKCYRCGERGHIE+NCKNSP K +R S S
Sbjct: 111 GPPPGSGRCFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCKNSPSPKKARQGGSYS 170
Query: 544 RSPVRSRSPRRGRSRDRS 597
RSPV+SRSPRR RS RS
Sbjct: 171 RSPVKSRSPRRRRSPSRS 188
Database: nr
Posted date: Apr 1, 2003 2:05 AM
Number of letters in database: 448,689,247
Number of sequences in database: 1,393,205
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 567,539,266
Number of Sequences: 1393205
Number of extensions: 14741820
Number of successful extensions: 101561
Number of sequences better than 10.0: 4254
Number of HSP's better than 10.0 without gapping: 67996
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 93124
length of database: 448,689,247
effective HSP length: 117
effective length of database: 285,684,262
effective search space used: 23140425222
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)