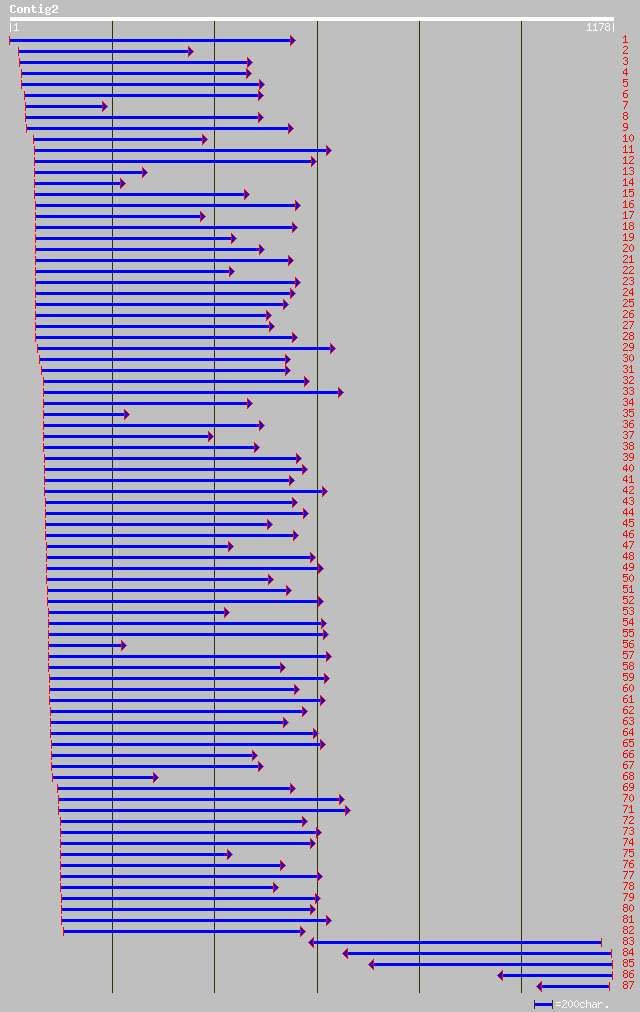
Nr search
BLASTX 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= KMC004544A_C02 KMC004544A_c02
(1162 letters)
Database: nr
1,393,205 sequences; 448,689,247 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO39008.1| plasma intrinsic protein 2,2 [Juglans regia] 496 e-139
gb|AAO39007.1| plasma intrinsic protein 2,1 [Juglans regia] 494 e-138
pir||T12440 mipC protein - common ice plant gi|1657948|gb|AAB182... 494 e-138
gb|AAF71816.1|AF141642_1 putative aquaporin PIP2-1 [Vitis berlan... 489 e-137
gb|AAC17529.1| aquaporin 2 [Samanea saman] 487 e-136
>gb|AAO39008.1| plasma intrinsic protein 2,2 [Juglans regia]
Length = 287
Score = 496 bits (1278), Expect = e-139
Identities = 244/286 (85%), Positives = 256/286 (89%)
Frame = -1
Query: 1120 MAKDIETEAPRGDQLAHKDYHDPPPAPLFDTTELTKWSFYRALIAEFVATLLFLYITVLT 941
MAKDIE G + KDYHDPPPAPL D E T+WSFYRA+IAEF+ATLLFLYITVLT
Sbjct: 1 MAKDIEAAGQGG--FSAKDYHDPPPAPLIDAEEFTQWSFYRAIIAEFIATLLFLYITVLT 58
Query: 940 VIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 761
VIGY SQTD A D CGGVGILGIAWAFGGMIF+LVYCTAGISGGHINPAVTFGLFLAR
Sbjct: 59 VIGYKSQTDKAKGGDDCGGVGILGIAWAFGGMIFVLVYCTAGISGGHINPAVTFGLFLAR 118
Query: 760 KVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHGGANMLADGYSKGTGLGAEIIGT 581
KVSLVRA+FYM AQCLGA+ G GLVKAFQK+YY++Y GGAN LADGYSKGTGL AEIIGT
Sbjct: 119 KVSLVRAVFYMAAQCLGAVCGCGLVKAFQKAYYSKYGGGANELADGYSKGTGLAAEIIGT 178
Query: 580 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 401
FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI
Sbjct: 179 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 238
Query: 400 YNNEKAWDDQWIFWVGPFIGATIAAIYHQYVLRAQAAKALGSFRSS 263
Y +KAWDDQWIFWVGPFIGA IAA YHQY+LRA AAKALGSFRSS
Sbjct: 239 YGKDKAWDDQWIFWVGPFIGAAIAAFYHQYILRAAAAKALGSFRSS 284
>gb|AAO39007.1| plasma intrinsic protein 2,1 [Juglans regia]
Length = 287
Score = 494 bits (1272), Expect = e-138
Identities = 243/286 (84%), Positives = 256/286 (88%)
Frame = -1
Query: 1120 MAKDIETEAPRGDQLAHKDYHDPPPAPLFDTTELTKWSFYRALIAEFVATLLFLYITVLT 941
MAKDIE G + KDYHDPPPAPL D E T+WSFYRA+IAEF+ATLLFLYITVLT
Sbjct: 1 MAKDIEAAGQGG--FSAKDYHDPPPAPLIDAEEFTQWSFYRAIIAEFIATLLFLYITVLT 58
Query: 940 VIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 761
VIGY SQTD A D CGGVGILGIAWAFGGMIF+LVYCTAGISGGHINPAVTFGLFLAR
Sbjct: 59 VIGYKSQTDKAKGGDDCGGVGILGIAWAFGGMIFVLVYCTAGISGGHINPAVTFGLFLAR 118
Query: 760 KVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHGGANMLADGYSKGTGLGAEIIGT 581
KVSLVRA+FYM AQCLGA+ G GLVKAFQK+YY++Y GGAN LADGYSKGTGL AEIIGT
Sbjct: 119 KVSLVRAVFYMAAQCLGAVCGCGLVKAFQKAYYSKYGGGANELADGYSKGTGLAAEIIGT 178
Query: 580 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 401
FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI
Sbjct: 179 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 238
Query: 400 YNNEKAWDDQWIFWVGPFIGATIAAIYHQYVLRAQAAKALGSFRSS 263
Y +KAW+DQWIFWVGPFIGA IAA YHQY+LRA AAKALGSFRSS
Sbjct: 239 YGKDKAWNDQWIFWVGPFIGAAIAAFYHQYILRAGAAKALGSFRSS 284
>pir||T12440 mipC protein - common ice plant gi|1657948|gb|AAB18227.1| MipC
[Mesembryanthemum crystallinum]
Length = 287
Score = 494 bits (1272), Expect = e-138
Identities = 239/286 (83%), Positives = 254/286 (88%)
Frame = -1
Query: 1120 MAKDIETEAPRGDQLAHKDYHDPPPAPLFDTTELTKWSFYRALIAEFVATLLFLYITVLT 941
M KD+E + + + KDYHDPPPAPL D EL KWSFYRA+IAEF+ATLLFLYITVLT
Sbjct: 1 MTKDVEVAEQQAPEFSAKDYHDPPPAPLIDFEELRKWSFYRAVIAEFIATLLFLYITVLT 60
Query: 940 VIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 761
VIGY SQTDP D CGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR
Sbjct: 61 VIGYKSQTDPNTNADQCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 120
Query: 760 KVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHGGANMLADGYSKGTGLGAEIIGT 581
KVSL++A+ YMVAQCLGAI GVG VKAFQK+YY RY GGAN +A GY+KGTGLGAEIIGT
Sbjct: 121 KVSLIKAVLYMVAQCLGAICGVGFVKAFQKAYYVRYGGGANEMASGYTKGTGLGAEIIGT 180
Query: 580 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 401
FVLVYTVF+ATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI
Sbjct: 181 FVLVYTVFAATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 240
Query: 400 YNNEKAWDDQWIFWVGPFIGATIAAIYHQYVLRAQAAKALGSFRSS 263
YN +KAWDD WIFWVGPFIGA IAA YHQY+LRA A KALGSFRSS
Sbjct: 241 YNQQKAWDDHWIFWVGPFIGAAIAAFYHQYILRAAAIKALGSFRSS 286
>gb|AAF71816.1|AF141642_1 putative aquaporin PIP2-1 [Vitis berlandieri x Vitis rupestris]
Length = 284
Score = 489 bits (1259), Expect = e-137
Identities = 245/286 (85%), Positives = 258/286 (89%)
Frame = -1
Query: 1120 MAKDIETEAPRGDQLAHKDYHDPPPAPLFDTTELTKWSFYRALIAEFVATLLFLYITVLT 941
M KD+E A G A KDYHDPPPAPLFD+ ELTKWSFYRALIAEF+ATLLFLYITVLT
Sbjct: 1 MTKDVEV-AEHGSFSA-KDYHDPPPAPLFDSVELTKWSFYRALIAEFIATLLFLYITVLT 58
Query: 940 VIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 761
VIGY SQT DPCGGVGILGIAW+FGGMIFILVYCTAGISGGHINPAVTFGLFLAR
Sbjct: 59 VIGYKSQTAGG---DPCGGVGILGIAWSFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 115
Query: 760 KVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHGGANMLADGYSKGTGLGAEIIGT 581
KVSL+RA+ YMVAQCLGAI GVGLVKAFQ +YY+RY GGAN L+ GYSKGTGLGAEIIGT
Sbjct: 116 KVSLIRAILYMVAQCLGAICGVGLVKAFQSAYYDRYGGGANELSTGYSKGTGLGAEIIGT 175
Query: 580 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 401
FVLVYTVFSATDPKR+ARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARS GAAVI
Sbjct: 176 FVLVYTVFSATDPKRSARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSLGAAVI 235
Query: 400 YNNEKAWDDQWIFWVGPFIGATIAAIYHQYVLRAQAAKALGSFRSS 263
YNNEKAWDDQWIFWVGPFIGA IAA YHQ++LRA A KALGSFRS+
Sbjct: 236 YNNEKAWDDQWIFWVGPFIGAAIAAFYHQFILRAGAIKALGSFRST 281
>gb|AAC17529.1| aquaporin 2 [Samanea saman]
Length = 287
Score = 487 bits (1254), Expect = e-136
Identities = 242/286 (84%), Positives = 257/286 (89%)
Frame = -1
Query: 1120 MAKDIETEAPRGDQLAHKDYHDPPPAPLFDTTELTKWSFYRALIAEFVATLLFLYITVLT 941
MAKD+E A RG A KDYHDPPPAPL D EL KWSFYRALIAEF+ATLLFLYITVLT
Sbjct: 1 MAKDVEV-AERGSYSA-KDYHDPPPAPLIDAEELGKWSFYRALIAEFIATLLFLYITVLT 58
Query: 940 VIGYSSQTDPANTTDPCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 761
VIGY SQ+D D CGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR
Sbjct: 59 VIGYKSQSDTKAGGDVCGGVGILGIAWAFGGMIFILVYCTAGISGGHINPAVTFGLFLAR 118
Query: 760 KVSLVRALFYMVAQCLGAISGVGLVKAFQKSYYNRYHGGANMLADGYSKGTGLGAEIIGT 581
KVSL+RA+ YMVAQCLGAI GVGLVKAFQK+YY+RY GGAN L+DGYS GTGLGAEIIGT
Sbjct: 119 KVSLIRAILYMVAQCLGAICGVGLVKAFQKAYYSRYGGGANTLSDGYSTGTGLGAEIIGT 178
Query: 580 FVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVFMVHLATIPITGTGINPARSFGAAVI 401
FVLVYTVFSATDPKR+ARDSHVPVLAPLPIGFAVFMVHLATIP+TGTGINPARS GAAVI
Sbjct: 179 FVLVYTVFSATDPKRSARDSHVPVLAPLPIGFAVFMVHLATIPVTGTGINPARSLGAAVI 238
Query: 400 YNNEKAWDDQWIFWVGPFIGATIAAIYHQYVLRAQAAKALGSFRSS 263
+N +KAWDD WIFWVGPFIGA IAA YHQ++LRA AAKALGSFRS+
Sbjct: 239 FNQQKAWDDHWIFWVGPFIGAAIAAFYHQFILRAGAAKALGSFRSN 284
Database: nr
Posted date: Apr 1, 2003 2:05 AM
Number of letters in database: 448,689,247
Number of sequences in database: 1,393,205
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,177,057,719
Number of Sequences: 1393205
Number of extensions: 32480033
Number of successful extensions: 291670
Number of sequences better than 10.0: 4393
Number of HSP's better than 10.0 without gapping: 145173
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 245923
length of database: 448,689,247
effective HSP length: 125
effective length of database: 274,538,622
effective search space used: 71654580342
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)