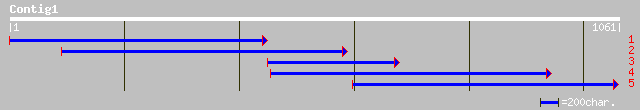
Nr search
BLASTX 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= KCC000011A_C01 KCC000011A_c01
(956 letters)
Database: nr
1,537,769 sequences; 498,525,298 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP73845.1| putative phospholipase [Oryza sativa (japonica cu... 233 5e-60
ref|NP_196024.1| patatin-related [Arabidopsis thaliana] gi|11282... 223 3e-57
dbj|BAB61223.1| contains EST AU057376(S21389)~similar to Arabido... 219 4e-56
ref|NP_191273.1| patatin-related [Arabidopsis thaliana] gi|11282... 207 2e-52
ref|XP_332099.1| hypothetical protein [Neurospora crassa] gi|289... 153 4e-36
>gb|AAP73845.1| putative phospholipase [Oryza sativa (japonica cultivar-group)]
Length = 820
Score = 233 bits (593), Expect = 5e-60
Identities = 143/326 (43%), Positives = 186/326 (56%), Gaps = 32/326 (9%)
Frame = +1
Query: 13 DLFSRLDEFD--GGFFSNSRAVELVQHLINKGSLQDMSDMIKKLRGLMGDATFLEAY*RT 186
D L FD GG F+ ++ V + G+L D+S M + LR L G+ TF EAY T
Sbjct: 285 DSLQTLQFFDRIGGIFAVTKRV------MTYGALHDISQMQRLLRDLTGNLTFQEAYDMT 338
Query: 187 GRILNVTVCPADTNEPTWVLNYLTAPNALIWSAGAASSAFPGLYPAQHILGRNSRGEIIR 366
GR+L VTVC NEP LNYLT+P+ +IWSA AS AFPGL+ AQ ++ +N GEI+
Sbjct: 339 GRVLGVTVCSPRKNEPPRCLNYLTSPHVVIWSAVTASCAFPGLFEAQELMAKNRFGEIVP 398
Query: 367 FSAQSTND------SLERRWRDGSLELDLPVQALGEMFNCNHFLVSQTNPHIVPLLNLKK 528
F A + D + +RRWRDGSLE+DLP+ L E+FN NHF+VSQTNPHI PLL LK+
Sbjct: 399 FHAPFSTDPEQGPGASKRRWRDGSLEMDLPMMQLKELFNVNHFIVSQTNPHISPLLRLKE 458
Query: 529 ALSR-------KWANVLEAELKHRC-QVAQWLLPEWVPSKWLMLFTQAWEGDITMTLPSA 684
++ K A + E E+K+RC Q+ + LP +K LF Q WEGD+TM +P+
Sbjct: 459 IVTTYGGRFAGKLARLAEMEVKYRCNQILEIGLPLGGLAK---LFAQDWEGDVTMVMPAT 515
Query: 685 LWHLSKTIVNPTTEELIRNVKVGEVDTWEKISAIECNCSIEATLDKCLA----------- 831
K I NPT EL G TWEKISAI NC+IE LD+ +A
Sbjct: 516 AAQYLKIIQNPTYAELQMAANQGRRCTWEKISAIRTNCAIELALDESIAVLNHKRRLKRS 575
Query: 832 --NIANQVRGYRLQ---RMHNRIPSW 894
+A+ +GY R R+PSW
Sbjct: 576 MERVASASQGYTCSSVIRTPRRVPSW 601
>ref|NP_196024.1| patatin-related [Arabidopsis thaliana] gi|11282314|pir||T48431
hypothetical protein F8F6.250 - Arabidopsis thaliana
gi|7406414|emb|CAB85524.1| putative protein [Arabidopsis
thaliana] gi|22531263|gb|AAM97135.1| putative protein
[Arabidopsis thaliana]
Length = 825
Score = 223 bits (569), Expect = 3e-57
Identities = 136/316 (43%), Positives = 183/316 (57%), Gaps = 15/316 (4%)
Frame = +1
Query: 43 GGFFSNSRAVELVQHLINKGSLQDMSDMIKKLRGLMGDATFLEAY*RTGRILNVTVCPAD 222
GG FS +V+ ++ +G+L D+ + LR L + TF EAY TGRIL +TVC
Sbjct: 300 GGVFS------IVKRVMTQGALHDIRQLQCMLRNLTSNLTFQEAYDMTGRILGITVCSPR 353
Query: 223 TNEPTWVLNYLTAPNALIWSAGAASSAFPGLYPAQHILGRNSRGEIIRFSAQ-------S 381
+EP LNYLT+P+ +IWSA AS AFPGL+ AQ ++ ++ GEI+ +
Sbjct: 354 KHEPPRCLNYLTSPHVVIWSAVTASCAFPGLFEAQELMAKDRSGEIVPYHPPFNLDPEVG 413
Query: 382 TNDSLERRWRDGSLELDLPVQALGEMFNCNHFLVSQTNPHIVPLLNLKKAL-------SR 540
T S RRWRDGSLE+DLP+ L E+FN NHF+VSQ NPHI PLL LK + +
Sbjct: 414 TKSSSGRRWRDGSLEVDLPMMQLKELFNVNHFIVSQANPHIAPLLRLKDLVRAYGGRFAA 473
Query: 541 KWANVLEAELKHRC-QVAQWLLPEWVPSKWLMLFTQAWEGDITMTLPSALWHLSKTIVNP 717
K A+++E E+KHRC QV + P +K LF Q WEGD+T+ +P+ L SK I NP
Sbjct: 474 KLAHLVEMEVKHRCNQVLELGFPLGGLAK---LFAQEWEGDVTVVMPATLAQYSKIIQNP 530
Query: 718 TTEELIRNVKVGEVDTWEKISAIECNCSIEATLDKCLANIANQVRGYRLQRMHNRIPSWL 897
T EL + G TWEK+SAI+ NC IE LD +A I N +R RL++ R +
Sbjct: 531 THVELQKAANQGRRCTWEKLSAIKSNCGIELALDDSVA-ILNHMR--RLKKSAERAATAT 587
Query: 898 HMSAVGLPAVVSWGDS 945
S GL + + S
Sbjct: 588 SSSHHGLASTTRFNAS 603
>dbj|BAB61223.1| contains EST AU057376(S21389)~similar to Arabidopsis thaliana
chromosome 5, F8F6.250~unknown protein [Oryza sativa
(japonica cultivar-group)] gi|20804680|dbj|BAB92368.1|
P0512C01.22 [Oryza sativa (japonica cultivar-group)]
Length = 1044
Score = 219 bits (559), Expect = 4e-56
Identities = 127/283 (44%), Positives = 172/283 (59%), Gaps = 14/283 (4%)
Frame = +1
Query: 76 LVQHLINKGSLQDMSDMIKKLRGLMGDATFLEAY*RTGRILNVTVCPADTNEPTWVLNYL 255
+V+ ++ G++ D+ + LR L + TF EAY TGRIL VTVC +EP LNYL
Sbjct: 306 VVKRILTHGAVHDIRHLQTLLRNLTSNLTFQEAYDMTGRILVVTVCSPRKHEPPRCLNYL 365
Query: 256 TAPNALIWSAGAASSAFPGLYPAQHILGRNSRGEIIRFSA------QSTNDSLERRWRDG 417
T+P+ LIWSA AS AFPGL+ AQ ++ ++ GE + F A + + RRWRDG
Sbjct: 366 TSPHVLIWSAVTASCAFPGLFEAQELMAKDRFGETVPFHAPFLLGLEERVGATTRRWRDG 425
Query: 418 SLELDLPVQALGEMFNCNHFLVSQTNPHIVPLLNLKK-------ALSRKWANVLEAELKH 576
SLE DLP++ L E+FN NHF+VSQ NPHI PLL LK+ + + K A + E E+KH
Sbjct: 426 SLESDLPMKQLKELFNVNHFIVSQANPHIAPLLRLKEIIRAYGGSFAAKLAELAEMEVKH 485
Query: 577 RC-QVAQWLLPEWVPSKWLMLFTQAWEGDITMTLPSALWHLSKTIVNPTTEELIRNVKVG 753
RC Q+ + P +K LF Q WEGD+T+ +P+ L SK I NP+ EL + G
Sbjct: 486 RCNQILELGFPLGGIAK---LFAQDWEGDVTVVMPATLAQYSKIIQNPSYAELQKAANQG 542
Query: 754 EVDTWEKISAIECNCSIEATLDKCLANIANQVRGYRLQRMHNR 882
TWEK+SAI NC+IE LD+C+A + N +R RL+R R
Sbjct: 543 RRCTWEKLSAIRANCAIELVLDECVA-LLNHMR--RLKRSAER 582
>ref|NP_191273.1| patatin-related [Arabidopsis thaliana] gi|11282313|pir||T47774
hypothetical protein F24I3.220 - Arabidopsis thaliana
gi|6911884|emb|CAB72184.1| putative protein [Arabidopsis
thaliana] gi|26450904|dbj|BAC42559.1| unknown protein
[Arabidopsis thaliana] gi|29029050|gb|AAO64904.1|
At3g57140 [Arabidopsis thaliana]
Length = 801
Score = 207 bits (528), Expect = 2e-52
Identities = 122/302 (40%), Positives = 175/302 (57%), Gaps = 13/302 (4%)
Frame = +1
Query: 43 GGFFSNSRAVELVQHLINKGSLQDMSDMIKKLRGLMGDATFLEAY*RTGRILNVTVCPAD 222
GG F+ V+ ++ +G++ ++ + KLR L + TF EAY TGRIL +TVC
Sbjct: 301 GGIFTT------VKRVMTQGAVHEIRHLQWKLRNLTNNLTFQEAYDITGRILGITVCSLR 354
Query: 223 TNEPTWVLNYLTAPNALIWSAGAASSAFPGLYPAQHILGRNSRGEIIRFSAQSTNDSLE- 399
+EP LNYLT+P+ +IWSA AS AFPGL+ AQ ++ ++ GEI+ + D E
Sbjct: 355 KHEPPRCLNYLTSPHVVIWSAVTASCAFPGLFEAQELMAKDRTGEIVPYHPPFNLDPEEG 414
Query: 400 ----RRWRDGSLELDLPVQALGEMFNCNHFLVSQTNPHIVPLLNLKK-------ALSRKW 546
RRWRDGSLE+DLP+ L E+FN NHF+VSQ NPHI P L +K+ + K
Sbjct: 415 SASVRRWRDGSLEMDLPMIQLKELFNVNHFIVSQANPHIAPFLRMKEFVRACGGRFAAKL 474
Query: 547 ANVLEAELKHRC-QVAQWLLPEWVPSKWLMLFTQAWEGDITMTLPSALWHLSKTIVNPTT 723
A + E E+KHRC QV + LP + LF Q WEGD+T+ +P+ K I NP+
Sbjct: 475 AQLAEMEVKHRCNQVLELGLP---LREVASLFAQEWEGDVTIVMPATFSQYLKIIQNPSN 531
Query: 724 EELIRNVKVGEVDTWEKISAIECNCSIEATLDKCLANIANQVRGYRLQRMHNRIPSWLHM 903
E+ + G TWEK++ I+ N IE LD+C+ + N +R RL+R R ++ +
Sbjct: 532 VEIQKAANQGRRCTWEKLAVIKANFGIELALDECV-TVLNHMR--RLKRSAERAAAFSAI 588
Query: 904 SA 909
S+
Sbjct: 589 SS 590
>ref|XP_332099.1| hypothetical protein [Neurospora crassa] gi|28920133|gb|EAA29514.1|
hypothetical protein [Neurospora crassa]
Length = 802
Score = 153 bits (387), Expect = 4e-36
Identities = 100/294 (34%), Positives = 150/294 (51%), Gaps = 24/294 (8%)
Frame = +1
Query: 19 FSRLDEFDGGFFSNSRAVELVQHLINKGSLQDMSDMIKKLRGLMGDATFLEAY*RTGRIL 198
+ LD F G N + ++ L+ +GS D++++ + +R ++GD TF EAY RT RI
Sbjct: 290 YGDLDVFKG---PNDGISDSLRRLLTQGSWADITNLTRVMRSMLGDLTFQEAYNRTRRIC 346
Query: 199 NVTVCPADTNEPTWVLNYLTAPNALIWSAGAASSAFPGLYPAQHILGRNSRGEIIRFSAQ 378
N+ V A E +LNY+TAPN +IWSA AAS + P ++ A +L ++ A
Sbjct: 347 NICVSTASIYELPRLLNYITAPNVMIWSAVAASCSVPLVFQAAPLLVKDPAT-----GAH 401
Query: 379 STNDSLERRWRDGSLELDLPVQALGEMFNCNHFLVSQTNPHIVPLLNLKKAL-------- 534
+ +RW DGS++ DLP+ L EMFN NHF+VSQ NPHIVP L+ L
Sbjct: 402 VPWNPTPQRWIDGSVDNDLPMTRLAEMFNVNHFIVSQVNPHIVPFLSKDDRLYPRNHPGR 461
Query: 535 -----------SRKWANVLEAELKHRCQVAQWLLPEW-----VPSKWLMLFTQAWEGDIT 666
+ +W L K L E+ + +K + +Q + GDIT
Sbjct: 462 LRQQKAAAQENNSEWLYYLTTLAKDEAVHRLHFLTEFGIFPGLLTKLRSILSQRYSGDIT 521
Query: 667 MTLPSALWHLSKTIVNPTTEELIRNVKVGEVDTWEKISAIECNCSIEATLDKCL 828
+ L L + + NPT+E ++RN +GE TW K+S I +IE LD+ +
Sbjct: 522 ILPELELQDLPRILKNPTSEFMMRNCLIGERATWPKLSRIRDRLAIELALDQAV 575