

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149579.7 - phase: 0 /pseudo
(367 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
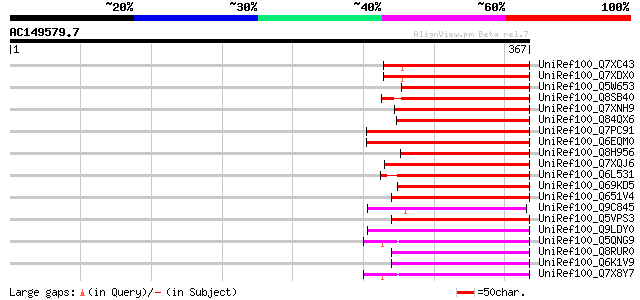
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XC43 Hypothetical protein [Oryza sativa] 107 4e-22
UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza... 107 5e-22
UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sat... 99 2e-19
UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa] 99 2e-19
UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa] 98 4e-19
UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sat... 98 4e-19
UniRef100_Q7PC91 Putative transposase [Oryza sativa] 97 5e-19
UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa] 97 5e-19
UniRef100_Q8H956 B protein [Oryza sativa] 97 7e-19
UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa] 97 9e-19
UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa] 96 2e-18
UniRef100_Q69KD5 Ribosomal protein-like [Oryza sativa] 96 2e-18
UniRef100_Q651V4 Ribosomal protein-like [Oryza sativa] 87 7e-16
UniRef100_Q9C845 Hypothetical protein F8D11.12 [Arabidopsis thal... 86 2e-15
UniRef100_Q5VPS3 Ribosomal protein-like [Oryza sativa] 85 4e-15
UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana] 84 8e-15
UniRef100_Q5QNG9 Hypothetical protein P0466B10.34 [Oryza sativa] 83 1e-14
UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa] 82 3e-14
UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa] 82 3e-14
UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa] 81 5e-14
>UniRef100_Q7XC43 Hypothetical protein [Oryza sativa]
Length = 513
Score = 107 bits (268), Expect = 4e-22
Identities = 52/106 (49%), Positives = 71/106 (66%), Gaps = 3/106 (2%)
Query: 265 RKQIEEGSGSRS---RKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRI 321
+ ++E G+ R+ RKY+ R+H GA+ +L DYFA +P Y A FRRR+RM++HVFL I
Sbjct: 39 KARLEGGTSRRTSGPRKYINRNHEGAHDQLFADYFAEDPLYSAATFRRRFRMRRHVFLHI 98
Query: 322 VGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
V +L +YFT RVD G SPL KCT +++ML YG AAD +D
Sbjct: 99 VDELGKWSSYFTHRVDCTGCLGHSPLQKCTAAIRMLAYGTAADTLD 144
>UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza sativa]
Length = 308
Score = 107 bits (267), Expect = 5e-22
Identities = 50/106 (47%), Positives = 72/106 (67%), Gaps = 3/106 (2%)
Query: 265 RKQIEEGSGSRS---RKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRI 321
+ ++E G+ R+ RKY+ R+H GA+ +L DYFA +P Y A FRRR+RM++HVFL I
Sbjct: 39 KARLEGGTSRRTSGPRKYINRNHEGAHDQLFADYFAKDPLYSAATFRRRFRMRRHVFLHI 98
Query: 322 VGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
V +L +YFT R++ + G SPL KCT +++ML YG AAD +D
Sbjct: 99 VDELGKWSSYFTHRINCTGRLGHSPLQKCTAAIRMLAYGTAADTLD 144
>UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sativa]
Length = 444
Score = 99.0 bits (245), Expect = 2e-19
Identities = 47/90 (52%), Positives = 60/90 (66%)
Query: 278 KYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQRVD 337
K ++RDH A+ RL+ DYFA P Y + MFR R+RM K +FLRIV LS YFTQR D
Sbjct: 59 KTIRRDHVDAHSRLVADYFAEHPLYPERMFRTRFRMHKPLFLRIVEALSQWSPYFTQRGD 118
Query: 338 AANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+ +SPL KCT +++ML YG ADA+D
Sbjct: 119 CSGHTSLSPLQKCTAALRMLAYGTPADALD 148
>UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa]
Length = 915
Score = 98.6 bits (244), Expect = 2e-19
Identities = 49/104 (47%), Positives = 68/104 (65%), Gaps = 4/104 (3%)
Query: 264 RRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVG 323
RR+Q + G R+Y+ R ++ L+ +YF+ P Y D FRRR+RM K +FLRIV
Sbjct: 519 RRRQRQSGP----RRYIPRPREKGHEDLVANYFSANPIYTDEQFRRRFRMNKPLFLRIVN 574
Query: 324 DLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
LS+ D +FTQRVDA ++ SPL KCT +++ML YG ADA+D
Sbjct: 575 ALSNWDQFFTQRVDATGRDSHSPLQKCTAAIRMLGYGTPADALD 618
>UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa]
Length = 450
Score = 97.8 bits (242), Expect = 4e-19
Identities = 47/95 (49%), Positives = 60/95 (62%)
Query: 273 GSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYF 332
G +R+Y+ RDH + RL YF+ P Y D FRRR+RM++H+FLRIV L YF
Sbjct: 61 GGYTRRYISRDHEDDHNRLFAKYFSESPLYTDDQFRRRFRMRRHLFLRIVQALGVWSPYF 120
Query: 333 TQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
R DA K G+SPL KCT +M+ML YG AD +D
Sbjct: 121 RLRRDAFGKVGLSPLQKCTAAMRMLAYGTPADLMD 155
>UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sativa]
Length = 626
Score = 97.8 bits (242), Expect = 4e-19
Identities = 46/94 (48%), Positives = 63/94 (66%)
Query: 274 SRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFT 333
S +R+Y+ R+ +N L+ +YF+ P Y D MFRRR+RM+K +FLRIV LS YFT
Sbjct: 238 SGTRRYIPRNREASNADLVANYFSESPIYTDKMFRRRFRMRKPLFLRIVSALSEWSPYFT 297
Query: 334 QRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
R+DA + G SPL KCT +++ML YG AD +D
Sbjct: 298 NRLDATGRAGHSPLQKCTAAIRMLAYGTPADQLD 331
>UniRef100_Q7PC91 Putative transposase [Oryza sativa]
Length = 482
Score = 97.4 bits (241), Expect = 5e-19
Identities = 49/115 (42%), Positives = 70/115 (60%)
Query: 253 EDTYMVNRFIQRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYR 312
E T +V I+ + RK++KR A+Q+L++DYF+ P Y +FRRR+R
Sbjct: 67 EATVVVQSTIEGLQNEASDHRHHPRKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFR 126
Query: 313 MQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
M + +FLRIV L YFTQRVDA N++G+SPL KCT +++ L G AD +D
Sbjct: 127 MSRPLFLRIVEALGQWSVYFTQRVDAVNRKGLSPLQKCTAAIRQLATGSGADELD 181
>UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa]
Length = 447
Score = 97.4 bits (241), Expect = 5e-19
Identities = 49/115 (42%), Positives = 70/115 (60%)
Query: 253 EDTYMVNRFIQRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYR 312
E T +V I+ + RK++KR A+Q+L++DYF+ P Y +FRRR+R
Sbjct: 32 EATVVVQSTIEGLQNEASDHRHHPRKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFR 91
Query: 313 MQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
M + +FLRIV L YFTQRVDA N++G+SPL KCT +++ L G AD +D
Sbjct: 92 MSRPLFLRIVEALGQWSVYFTQRVDAVNRKGLSPLQKCTAAIRQLATGSGADELD 146
>UniRef100_Q8H956 B protein [Oryza sativa]
Length = 455
Score = 97.1 bits (240), Expect = 7e-19
Identities = 45/91 (49%), Positives = 63/91 (68%)
Query: 277 RKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQRV 336
RK++KR A+Q L++DYF+ P Y +FRRR+RM + +FLRIV L +YFTQRV
Sbjct: 57 RKHIKRPREEAHQNLVNDYFSENPLYPSNIFRRRFRMYRPLFLRIVDALGQWSDYFTQRV 116
Query: 337 DAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
DAA ++G+SPL KCT +++ L G AD +D
Sbjct: 117 DAAGRQGLSPLQKCTAAIRQLATGSGADELD 147
>UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa]
Length = 454
Score = 96.7 bits (239), Expect = 9e-19
Identities = 48/102 (47%), Positives = 63/102 (61%)
Query: 266 KQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDL 325
K+ + G ++ RKY++R+ A RL YF P Y+D FR RYRM+KH+FL IV L
Sbjct: 56 KKQQRGPPTQQRKYIRRERQDAEDRLKAYYFLENPMYNDTQFRWRYRMKKHLFLHIVQTL 115
Query: 326 SSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
S F QR DA K G SPL KCT +++ML YG +AD +D
Sbjct: 116 SHWSPVFQQRKDAFGKVGFSPLLKCTAALRMLAYGTSADILD 157
>UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa]
Length = 443
Score = 95.9 bits (237), Expect = 2e-18
Identities = 48/105 (45%), Positives = 68/105 (64%), Gaps = 6/105 (5%)
Query: 263 QRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIV 322
+RR+Q S R+Y+ R+ + L+ +YF+ P Y D MFRRR+RM++ +FLRIV
Sbjct: 51 RRRRQ------SGPRRYIPRNRERGHDDLVANYFSANPIYTDEMFRRRFRMRRPLFLRIV 104
Query: 323 GDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+LS YFTQRVDA + +SPL KC +++ML YG AAD +D
Sbjct: 105 HELSEWSPYFTQRVDATGRNSLSPLQKCIAAIRMLGYGTAADQLD 149
>UniRef100_Q69KD5 Ribosomal protein-like [Oryza sativa]
Length = 510
Score = 95.9 bits (237), Expect = 2e-18
Identities = 44/93 (47%), Positives = 59/93 (63%)
Query: 275 RSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQ 334
R R+Y +RD G + R++ DYF P Y D FRRR+RM++ +FLRI+ L YFTQ
Sbjct: 58 RRRRYCRRDRIGGHARIMADYFCENPVYSDRDFRRRFRMRRPLFLRILNALGEWSEYFTQ 117
Query: 335 RVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
RVD + G+SP+ KCT +ML Y +AD VD
Sbjct: 118 RVDGIGQLGLSPIQKCTAVNRMLAYAASADQVD 150
>UniRef100_Q651V4 Ribosomal protein-like [Oryza sativa]
Length = 373
Score = 87.0 bits (214), Expect = 7e-16
Identities = 41/97 (42%), Positives = 63/97 (64%)
Query: 271 GSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 330
G K KRD A+ +L +DYF P Y++ FRRR+RM++ +FLRIV ++++ +
Sbjct: 40 GGSVPGHKTYKRDRIAADWQLNEDYFVERPLYNEEHFRRRFRMRRELFLRIVDEVTAKNR 99
Query: 331 YFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+FTQR +AA + G S L KCT +++ML YG A+ +D
Sbjct: 100 FFTQRRNAAGQLGFSALHKCTVALKMLAYGGPANELD 136
>UniRef100_Q9C845 Hypothetical protein F8D11.12 [Arabidopsis thaliana]
Length = 333
Score = 85.9 bits (211), Expect = 2e-15
Identities = 48/119 (40%), Positives = 68/119 (56%), Gaps = 7/119 (5%)
Query: 254 DTYMVNRFIQRRKQIEEGSGSRSRK-------YLKRDHAGANQRLIDDYFANEPTYDDAM 306
D ++F Q Q E G+R R Y++R+ + +L++DYF PTY +
Sbjct: 11 DDMFDDKFDQCFDQALESYGNRQRVKPRKKKLYIERNREEGHIQLVNDYFTENPTYPPHI 70
Query: 307 FRRRYRMQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADA 365
FRRR+RM K +F+RIV S+ YF QR DA + G S L K T +++ML YG+AADA
Sbjct: 71 FRRRFRMNKSLFMRIVERFSNEVPYFKQRRDATRRLGFSALQKSTAAIRMLAYGIAADA 129
>UniRef100_Q5VPS3 Ribosomal protein-like [Oryza sativa]
Length = 436
Score = 84.7 bits (208), Expect = 4e-15
Identities = 41/97 (42%), Positives = 61/97 (62%)
Query: 271 GSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 330
G K KRD A+ +L DYF P Y++ FRRR+RM++ +FLRIV ++++ +
Sbjct: 40 GGSVPGHKTYKRDRIAADWQLNQDYFVERPLYNEEHFRRRFRMRRELFLRIVDEVTAKNR 99
Query: 331 YFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+FTQR +AA + G S L KCT +++ML Y AD +D
Sbjct: 100 FFTQRRNAAGQLGFSALHKCTVALKMLAYVGPADELD 136
>UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana]
Length = 415
Score = 83.6 bits (205), Expect = 8e-15
Identities = 45/114 (39%), Positives = 64/114 (55%)
Query: 254 DTYMVNRFIQRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRM 313
D N F++ E +R R Y R + L +DYF++ + FRRR+RM
Sbjct: 22 DQQFENVFVELSNSEEVAKKNRKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRM 81
Query: 314 QKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+K +FLRIV LSS +F R DA + G SP+ KCT ++++L YG A+DAVD
Sbjct: 82 KKPLFLRIVDRLSSELMFFQHRRDATGRFGHSPIQKCTAAIRLLAYGYASDAVD 135
>UniRef100_Q5QNG9 Hypothetical protein P0466B10.34 [Oryza sativa]
Length = 417
Score = 83.2 bits (204), Expect = 1e-14
Identities = 45/122 (36%), Positives = 72/122 (58%), Gaps = 6/122 (4%)
Query: 251 DAEDTYMVNRFI-----QRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDA 305
D +D Y+V + +R K+ GS +R+ + RD N R++ DYFA+ P Y+
Sbjct: 105 DDDDYYIVAALLTDIEHKRTKRKRRGSVP-AREIIHRDRFAGNLRIVADYFADPPVYNAK 163
Query: 306 MFRRRYRMQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADA 365
+FRRR+RM + +FLRIV + + D+YF QR +A G + L K +++ML Y + AD+
Sbjct: 164 LFRRRFRMSRELFLRIVASVEAHDDYFRQRPNAMGLLGATALQKVYGAIRMLAYDIPADS 223
Query: 366 VD 367
+D
Sbjct: 224 LD 225
>UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa]
Length = 626
Score = 81.6 bits (200), Expect = 3e-14
Identities = 40/97 (41%), Positives = 57/97 (58%)
Query: 271 GSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 330
G + + R+ + RL DYF+N PTY +FRRR RM + +FLRI+ + D+
Sbjct: 243 GGSVMGHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDD 302
Query: 331 YFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
YF Q+ +AA G S K T +M+ L YG+AADA+D
Sbjct: 303 YFVQKRNAAGLIGFSCHQKVTAAMRQLAYGIAADALD 339
>UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa]
Length = 916
Score = 81.6 bits (200), Expect = 3e-14
Identities = 40/97 (41%), Positives = 57/97 (58%)
Query: 271 GSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 330
G + + R+ + RL DYF+N PTY +FRRR RM + +FLRI+ + D+
Sbjct: 533 GGSVMGHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDD 592
Query: 331 YFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
YF Q+ +AA G S K T +M+ L YG+AADA+D
Sbjct: 593 YFVQKRNAAGLIGFSCHQKVTAAMRQLAYGIAADALD 629
>UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa]
Length = 440
Score = 80.9 bits (198), Expect = 5e-14
Identities = 44/122 (36%), Positives = 69/122 (56%), Gaps = 6/122 (4%)
Query: 251 DAEDTYMVNRFI-----QRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDA 305
D +D Y+ + +R K+ GS R+ + RD N R++ DYFA+ P Y
Sbjct: 30 DDDDYYIAAALLTDIEHKRTKRKHHGSVP-GREIIHRDRFAGNLRIVADYFADPPIYSAK 88
Query: 306 MFRRRYRMQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADA 365
+FRRR+RM + +FLRIV + + D+YF QR +A G + L K +++ML Y + AD+
Sbjct: 89 LFRRRFRMSRELFLRIVASVEAHDDYFRQRPNAVGLLGATALQKVYGAIRMLAYDIPADS 148
Query: 366 VD 367
+D
Sbjct: 149 LD 150
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.333 0.143 0.475
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 641,441,196
Number of Sequences: 2790947
Number of extensions: 28120306
Number of successful extensions: 186271
Number of sequences better than 10.0: 648
Number of HSP's better than 10.0 without gapping: 219
Number of HSP's successfully gapped in prelim test: 467
Number of HSP's that attempted gapping in prelim test: 162292
Number of HSP's gapped (non-prelim): 7315
length of query: 367
length of database: 848,049,833
effective HSP length: 129
effective length of query: 238
effective length of database: 488,017,670
effective search space: 116148205460
effective search space used: 116148205460
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 75 (33.5 bits)
Medicago: description of AC149579.7


