

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149306.10 - phase: 0 /pseudo
(138 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
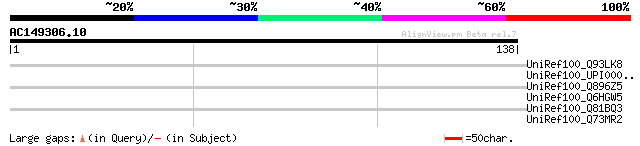
Sequences producing significant alignments: (bits) Value
UniRef100_Q93LK8 Spore germination protein GerQB [Bacillus cereus] 31 5.2
UniRef100_UPI00003CB21F UPI00003CB21F UniRef100 entry 31 6.8
UniRef100_Q896Z5 Ethanolamine utilization protein [Clostridium t... 31 6.8
UniRef100_Q6HGW5 Spore germination protein, GerB family [Bacillu... 30 8.9
UniRef100_Q81BQ3 Spore germination protein QB [Bacillus cereus] 30 8.9
UniRef100_Q73MR2 Membrane protein, putative [Treponema denticola] 30 8.9
>UniRef100_Q93LK8 Spore germination protein GerQB [Bacillus cereus]
Length = 360
Score = 31.2 bits (69), Expect = 5.2
Identities = 15/38 (39%), Positives = 21/38 (54%)
Query: 98 LFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIVVV 135
LF I FFLN R + L G IG Y+Y+++ V+
Sbjct: 311 LFIFILSFFLNNRNKINLLNTWTGQIGFVYIYVYLPVL 348
>UniRef100_UPI00003CB21F UPI00003CB21F UniRef100 entry
Length = 360
Score = 30.8 bits (68), Expect = 6.8
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 98 LFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIVVV 135
LF + FFLN R + L G IG Y+Y+++ V+
Sbjct: 311 LFIFVLSFFLNNRNKINLLNTWTGQIGFVYIYVYLPVL 348
>UniRef100_Q896Z5 Ethanolamine utilization protein [Clostridium tetani]
Length = 265
Score = 30.8 bits (68), Expect = 6.8
Identities = 16/45 (35%), Positives = 25/45 (55%)
Query: 81 LSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFLRKLLGSIGI 125
++ GLL + +G +YN+ P I F FL G++K KL+ I
Sbjct: 58 IAGGLLCDVDKGYMLYNMIPIILFSFLLSIGLIKIPDKLINIFNI 102
>UniRef100_Q6HGW5 Spore germination protein, GerB family [Bacillus thuringiensis]
Length = 360
Score = 30.4 bits (67), Expect = 8.9
Identities = 13/38 (34%), Positives = 21/38 (55%)
Query: 98 LFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIVVV 135
LF + FFLN R + L G IG Y+Y+++ ++
Sbjct: 311 LFIFVLSFFLNNRNKINLLNTWTGQIGFVYIYVYLPIL 348
>UniRef100_Q81BQ3 Spore germination protein QB [Bacillus cereus]
Length = 360
Score = 30.4 bits (67), Expect = 8.9
Identities = 13/38 (34%), Positives = 21/38 (55%)
Query: 98 LFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIVVV 135
LF + FFLN R + L G IG Y+Y+++ ++
Sbjct: 311 LFIFVLSFFLNNRNKINLLNTWTGQIGFVYIYVYLPIL 348
>UniRef100_Q73MR2 Membrane protein, putative [Treponema denticola]
Length = 209
Score = 30.4 bits (67), Expect = 8.9
Identities = 22/71 (30%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Query: 65 ISIFWWPSGSFEICPILSSGLLSFLFQGSYMYNLFP*IFFFF----LNIRGVVKFLRKLL 120
I + ++P S +I LSS L F FQ ++ LF FFF L+ + L L
Sbjct: 42 IIVSFFPFSSLQISSYLSSHLCRFFFQYFFLNALFGLAFFFLISWSLSEETLSNSLSALF 101
Query: 121 GSIGIQYVYLW 131
G + YL+
Sbjct: 102 GIFSAVFAYLF 112
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.368 0.170 0.704
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 179,112,628
Number of Sequences: 2790947
Number of extensions: 5774423
Number of successful extensions: 34618
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 34613
Number of HSP's gapped (non-prelim): 6
length of query: 138
length of database: 848,049,833
effective HSP length: 114
effective length of query: 24
effective length of database: 529,881,875
effective search space: 12717165000
effective search space used: 12717165000
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149306.10


