

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148758.11 + phase: 0
(358 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
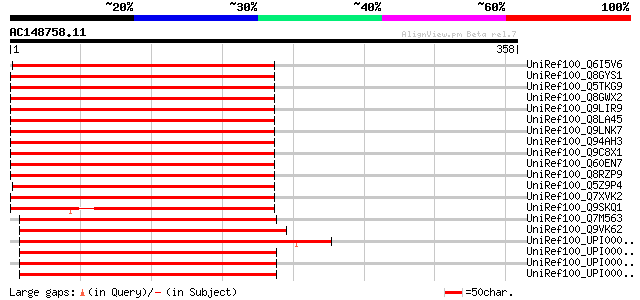
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6I5V6 Hypothetical protein OJ1378_A04.2 [Oryza sativa] 280 6e-74
UniRef100_Q8GYS1 Hypothetical protein At4g38730/T9A14_10 [Arabid... 263 6e-69
UniRef100_Q5TKG9 Hypothetical protein B1155G07.13 [Oryza sativa] 256 5e-67
UniRef100_Q8GWX2 Hypothetical protein At2g21120 [Arabidopsis tha... 254 3e-66
UniRef100_Q9LIR9 Gb|AAF34307.1 [Arabidopsis thaliana] 250 4e-65
UniRef100_Q8LA45 Hypothetical protein [Arabidopsis thaliana] 248 2e-64
UniRef100_Q9LNK7 F12K21.21 [Arabidopsis thaliana] 247 4e-64
UniRef100_Q94AH3 Hypothetical protein At1g71900 [Arabidopsis tha... 245 1e-63
UniRef100_Q9C8X1 Hypothetical protein F17M19.5 [Arabidopsis thal... 245 1e-63
UniRef100_Q60EN7 Unknow protein [Oryza sativa] 244 3e-63
UniRef100_Q8RZP9 B1065E10.15 protein [Oryza sativa] 244 3e-63
UniRef100_Q5Z9P4 Permease-like [Oryza sativa] 242 1e-62
UniRef100_Q7XVK2 OSJNBa0069D17.4 protein [Oryza sativa] 225 2e-57
UniRef100_Q9SKQ1 Hypothetical protein At2g21120 [Arabidopsis tha... 209 1e-52
UniRef100_Q7M563 Non-imprinted in Prader-Willi/Angelman syndrome... 197 5e-49
UniRef100_Q9VK62 CG12292-PA [Drosophila melanogaster] 196 1e-48
UniRef100_UPI00003AD972 UPI00003AD972 UniRef100 entry 195 1e-48
UniRef100_UPI000024CB22 UPI000024CB22 UniRef100 entry 195 2e-48
UniRef100_UPI0000249C5C UPI0000249C5C UniRef100 entry 195 2e-48
UniRef100_UPI00003A9E57 UPI00003A9E57 UniRef100 entry 194 2e-48
>UniRef100_Q6I5V6 Hypothetical protein OJ1378_A04.2 [Oryza sativa]
Length = 355
Score = 280 bits (715), Expect = 6e-74
Identities = 133/185 (71%), Positives = 160/185 (85%)
Query: 3 STNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTMIV 62
+ NL G LLA+ SSAFIG SFI+KKKGL+ A GP A VGGYGYLL+PLWWVGM+TM++
Sbjct: 14 AANLKGSLLAVASSAFIGVSFIVKKKGLRRAGAAGPRAGVGGYGYLLEPLWWVGMITMLI 73
Query: 63 GEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVL 122
GEIANFVAYM+APAVLVTPLGALSIIVSAVLAHF+L EKLQ+MG+LGC++CI+GST+I+L
Sbjct: 74 GEIANFVAYMFAPAVLVTPLGALSIIVSAVLAHFILNEKLQRMGVLGCVLCIVGSTVIIL 133
Query: 123 HAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIF 182
HAP+E + SSV+QIW LA QPAFL Y A+ ++L L+ +CAPRYGQ+NI VYIGICS+
Sbjct: 134 HAPEEETPSSVEQIWHLATQPAFLCYVAFALVVSLILMAHCAPRYGQTNIAVYIGICSVI 193
Query: 183 GSLTV 187
GSLTV
Sbjct: 194 GSLTV 198
>UniRef100_Q8GYS1 Hypothetical protein At4g38730/T9A14_10 [Arabidopsis thaliana]
Length = 326
Score = 263 bits (672), Expect = 6e-69
Identities = 129/187 (68%), Positives = 155/187 (81%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
M S N G +LA+ SS FIGSSFI+KKKGL+ A NG A GGY YLL+PLWWVG+VTM
Sbjct: 1 MVSDNEMGLVLAVSSSVFIGSSFILKKKGLKRAAANGTRAGFGGYTYLLEPLWWVGLVTM 60
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
GEIANFVAY+YAPAVLVTPLGALSII+SAVLAHFLL EKL+KMG+ GC+ CI+GS +I
Sbjct: 61 TFGEIANFVAYVYAPAVLVTPLGALSIIISAVLAHFLLDEKLRKMGVWGCVCCIVGSVMI 120
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
V+HAPQE + +SV++IWKLA+QPAFL+Y +++I L L+LYC P GQ+NILVYIGICS
Sbjct: 121 VIHAPQEQTPNSVEEIWKLAMQPAFLIYVAISMSIVLALILYCEPLCGQTNILVYIGICS 180
Query: 181 IFGSLTV 187
+ GSLTV
Sbjct: 181 LMGSLTV 187
>UniRef100_Q5TKG9 Hypothetical protein B1155G07.13 [Oryza sativa]
Length = 354
Score = 256 bits (655), Expect = 5e-67
Identities = 122/188 (64%), Positives = 155/188 (81%), Gaps = 1/188 (0%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARV-NGPSASVGGYGYLLQPLWWVGMVT 59
+S+ N+ G +LAL+SS FIG+SFIIKKKGL+ A V +G A VGGY YLL+PLWWVGM+T
Sbjct: 11 LSTDNVKGIVLALLSSGFIGASFIIKKKGLRRAAVASGIRAGVGGYSYLLEPLWWVGMIT 70
Query: 60 MIVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTI 119
MIVGE+ANFVAY +APAVLVTPLGALSIIVSAVLAHF+L E+L +G+LGC++CI GS +
Sbjct: 71 MIVGEVANFVAYAFAPAVLVTPLGALSIIVSAVLAHFILNERLHALGVLGCVMCIAGSVV 130
Query: 120 IVLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGIC 179
IV+HAPQE ++SV++IW +AIQPAFL+Y S I + LV + +P YGQSN+L+Y IC
Sbjct: 131 IVIHAPQEQEITSVREIWNMAIQPAFLLYVASVIVVVFVLVFHFSPLYGQSNVLIYTAIC 190
Query: 180 SIFGSLTV 187
S+ GSL+V
Sbjct: 191 SLMGSLSV 198
>UniRef100_Q8GWX2 Hypothetical protein At2g21120 [Arabidopsis thaliana]
Length = 328
Score = 254 bits (648), Expect = 3e-66
Identities = 125/187 (66%), Positives = 152/187 (80%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
M + N G +LA+ SS FIGSSFI+KKKGL+ A G A GGY YLL+PLWW GMVTM
Sbjct: 1 METDNGKGLILAVASSVFIGSSFILKKKGLKRAGAIGTRAGYGGYTYLLEPLWWAGMVTM 60
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGE ANFVAY+YAPAVLVTPLGALSII+SAVLAHFLLKEKL+KMG+LGC+ CI+GS +I
Sbjct: 61 IVGEAANFVAYIYAPAVLVTPLGALSIIISAVLAHFLLKEKLKKMGVLGCVSCIVGSVVI 120
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
V+HAP+E + +SV++IW LA QPAFL+Y ++I L L+L+ P GQ+NILVYIGICS
Sbjct: 121 VIHAPKEQTPNSVEEIWNLATQPAFLIYVAITMSIVLALILHFEPLCGQTNILVYIGICS 180
Query: 181 IFGSLTV 187
+ G+LTV
Sbjct: 181 LMGALTV 187
>UniRef100_Q9LIR9 Gb|AAF34307.1 [Arabidopsis thaliana]
Length = 335
Score = 250 bits (639), Expect = 4e-65
Identities = 116/187 (62%), Positives = 148/187 (79%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MS N+ G +LA+ SS FIGSSFIIKKKGL+ A +G A GGYGYL +P WW GM+TM
Sbjct: 4 MSPDNINGVILAVSSSIFIGSSFIIKKKGLKKAGASGVRAGEGGYGYLKEPWWWAGMITM 63
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGE+ANF AY +APA+LVTPLGALSII SAVLAHF+LKEKL G+LGC++C++GST I
Sbjct: 64 IVGEVANFAAYAFAPAILVTPLGALSIIFSAVLAHFILKEKLHMFGILGCILCVVGSTTI 123
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAP E + SV+QIW+LAI+P FL+Y+ + + L+ Y PRYG+++++VY+GICS
Sbjct: 124 VLHAPHEQKIESVKQIWQLAIEPGFLVYSAVIVIVVAILIFYYEPRYGKTHMIVYVGICS 183
Query: 181 IFGSLTV 187
+ GSLTV
Sbjct: 184 LMGSLTV 190
>UniRef100_Q8LA45 Hypothetical protein [Arabidopsis thaliana]
Length = 333
Score = 248 bits (633), Expect = 2e-64
Identities = 116/187 (62%), Positives = 147/187 (78%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MS N+ G +LA+ SS FIGSSFIIKKKGL+ A V+G A GGYGYL +P WW GM+TM
Sbjct: 1 MSPDNIHGVILAVSSSIFIGSSFIIKKKGLKKAGVSGARAGEGGYGYLYEPWWWAGMITM 60
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGEIANF AY +APA+LVTPLGALSII SAVLAHF+L+EKL G+LGC++C++GST I
Sbjct: 61 IVGEIANFAAYAFAPAILVTPLGALSIIFSAVLAHFILEEKLHMFGILGCVLCVVGSTTI 120
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAP E + SV+Q+W LA +P FL Y+ + + L L+ Y PRYG+++++VY+GICS
Sbjct: 121 VLHAPHEQGIESVKQVWHLATEPGFLAYSAVVLVVVLALIFYYEPRYGKTHMIVYVGICS 180
Query: 181 IFGSLTV 187
+ GSLTV
Sbjct: 181 LMGSLTV 187
>UniRef100_Q9LNK7 F12K21.21 [Arabidopsis thaliana]
Length = 368
Score = 247 bits (630), Expect = 4e-64
Identities = 114/187 (60%), Positives = 147/187 (77%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MSS N+ G +LAL SS FIG+SFI+KKKGL+ A +G A GGY YLL+PLWWVGM+TM
Sbjct: 15 MSSDNIKGLVLALSSSLFIGASFIVKKKGLKRAGASGLRAGSGGYSYLLEPLWWVGMITM 74
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGEIANF AY +APA+LVTPLGALSII+SA LAH +L EKL G+LGC++C++GS I
Sbjct: 75 IVGEIANFAAYAFAPAILVTPLGALSIIISAALAHVILHEKLHTFGLLGCVLCVVGSITI 134
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAPQE + SV Q+W LA +PAFL+Y + + + L++ P+YGQS+++VYIG+CS
Sbjct: 135 VLHAPQEQEIDSVLQVWNLATEPAFLLYAAAVVGAAIILIVQFVPQYGQSHVMVYIGVCS 194
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 195 LVGSLSV 201
>UniRef100_Q94AH3 Hypothetical protein At1g71900 [Arabidopsis thaliana]
Length = 343
Score = 245 bits (626), Expect = 1e-63
Identities = 113/187 (60%), Positives = 147/187 (78%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MSS N+ G +LAL SS FIG+SFI+KKKGL+ A G A VGGY YL +PLWW+GM TM
Sbjct: 15 MSSDNIKGLVLALSSSLFIGASFIVKKKGLKKAASTGTRAGVGGYSYLYEPLWWIGMTTM 74
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
++GEIANF AY +APA+LVTPLGA+SII+SAVLAH +L+EKL G+LGC +C++GST I
Sbjct: 75 LLGEIANFAAYAFAPAILVTPLGAVSIIISAVLAHIILREKLHIFGILGCALCVVGSTTI 134
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAPQE + SV ++W LA +PAF+ Y + I +FL++ P+YGQ+N++VYIGICS
Sbjct: 135 VLHAPQEREIDSVIEVWNLATEPAFMFYASLVIGAAVFLIIRFVPQYGQTNVMVYIGICS 194
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 195 LVGSLSV 201
>UniRef100_Q9C8X1 Hypothetical protein F17M19.5 [Arabidopsis thaliana]
Length = 347
Score = 245 bits (626), Expect = 1e-63
Identities = 113/187 (60%), Positives = 147/187 (78%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MSS N+ G +LAL SS FIG+SFI+KKKGL+ A G A VGGY YL +PLWW+GM TM
Sbjct: 15 MSSDNIKGLVLALSSSLFIGASFIVKKKGLKKAASTGTRAGVGGYSYLYEPLWWIGMTTM 74
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
++GEIANF AY +APA+LVTPLGA+SII+SAVLAH +L+EKL G+LGC +C++GST I
Sbjct: 75 LLGEIANFAAYAFAPAILVTPLGAVSIIISAVLAHIILREKLHIFGILGCALCVVGSTTI 134
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAPQE + SV ++W LA +PAF+ Y + I +FL++ P+YGQ+N++VYIGICS
Sbjct: 135 VLHAPQEREIDSVIEVWNLATEPAFMFYASLVIGAAVFLIIRFVPQYGQTNVMVYIGICS 194
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 195 LVGSLSV 201
>UniRef100_Q60EN7 Unknow protein [Oryza sativa]
Length = 358
Score = 244 bits (622), Expect = 3e-63
Identities = 114/187 (60%), Positives = 146/187 (77%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MS+ N+ G LLAL SS FIG+SFI+KKKGL+ A +G A VGGY YLL+PLWW GM M
Sbjct: 13 MSADNVKGLLLALSSSLFIGASFIVKKKGLKKAGASGVRAGVGGYSYLLEPLWWAGMTAM 72
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGEIANF AY +APA+LVTPLGALSII+SAVLAH +L+EKL G+LGC++C++GST I
Sbjct: 73 IVGEIANFAAYAFAPAILVTPLGALSIIISAVLAHIILREKLHIFGILGCILCVVGSTSI 132
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAP E + SV ++W LA +PAFL+Y +A L+ + P+YGQ++I+VYIG+CS
Sbjct: 133 VLHAPPERQIESVAEVWDLATEPAFLLYAAIVLAAAFVLIFHFVPQYGQTHIMVYIGVCS 192
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 193 LVGSLSV 199
>UniRef100_Q8RZP9 B1065E10.15 protein [Oryza sativa]
Length = 592
Score = 244 bits (622), Expect = 3e-63
Identities = 114/187 (60%), Positives = 147/187 (77%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MS+ N+ G +LAL SS FIG+SFI+KKKGL+ A +G A VGGY YL +PLWW GM+TM
Sbjct: 22 MSADNIKGLVLALSSSFFIGASFIVKKKGLKKAGASGVRAGVGGYSYLYEPLWWAGMITM 81
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGE+ANF AY +APA+LVTPLGALSII+SAVLA +LKEKL G+LGC++C++GST I
Sbjct: 82 IVGEVANFAAYAFAPAILVTPLGALSIIISAVLADIMLKEKLHIFGILGCVLCVVGSTTI 141
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAPQE + SV ++W LA +PAFL Y + +A T L+ P+YGQ++I+VYIG+CS
Sbjct: 142 VLHAPQEREIDSVAEVWALATEPAFLFYAVTVLAATFVLIFRFIPQYGQTHIMVYIGVCS 201
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 202 LVGSLSV 208
>UniRef100_Q5Z9P4 Permease-like [Oryza sativa]
Length = 344
Score = 242 bits (617), Expect = 1e-62
Identities = 117/185 (63%), Positives = 146/185 (78%)
Query: 3 STNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTMIV 62
S N G LA+ SSAFIG+SFI+KK GL A G A GGY YLL+PLWW GM+TM++
Sbjct: 4 SDNTVGLSLAVASSAFIGASFILKKIGLIRAGKGGVRAGGGGYTYLLEPLWWAGMMTMLL 63
Query: 63 GEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVL 122
GEIANFVAY +APAVLVTPLGALSIIVS+ LAHF+LKE+L+K+G+LGC+ CI+GS I+V+
Sbjct: 64 GEIANFVAYTFAPAVLVTPLGALSIIVSSFLAHFVLKERLEKLGVLGCVSCIVGSVIVVI 123
Query: 123 HAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIF 182
HAPQE +SV++IW LAIQP FL Y + + + LVL+ PRYGQ+NI++Y+GICS
Sbjct: 124 HAPQEHMPNSVEEIWNLAIQPGFLTYAVATLVVVAALVLFFEPRYGQTNIMIYLGICSSM 183
Query: 183 GSLTV 187
GSLTV
Sbjct: 184 GSLTV 188
>UniRef100_Q7XVK2 OSJNBa0069D17.4 protein [Oryza sativa]
Length = 317
Score = 225 bits (573), Expect = 2e-57
Identities = 108/187 (57%), Positives = 137/187 (72%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MS N+ GF LA SSAFIGSSF+IKK GL+ A G A GGY YL +PLWW+GM M
Sbjct: 3 MSIDNVRGFALATSSSAFIGSSFVIKKIGLKKAGDAGVRAGSGGYSYLYEPLWWIGMTAM 62
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
I+GE+ANF AY +APA+LVTPLGALSII SAVLAHF+LKE+L G++GC++C++GS I
Sbjct: 63 ILGEVANFAAYAFAPAILVTPLGALSIIFSAVLAHFILKERLHMFGIVGCILCVVGSVGI 122
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAP+E + SV +IW LA QP F++Y+ A+ + L L+ + R Q +L YI ICS
Sbjct: 123 VLHAPKEKKIDSVNEIWHLATQPGFIVYSCMAVVVALILIFWVVHRTEQRKMLAYIAICS 182
Query: 181 IFGSLTV 187
+ GSLTV
Sbjct: 183 LMGSLTV 189
>UniRef100_Q9SKQ1 Hypothetical protein At2g21120 [Arabidopsis thaliana]
Length = 323
Score = 209 bits (531), Expect = 1e-52
Identities = 111/192 (57%), Positives = 141/192 (72%), Gaps = 15/192 (7%)
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSAS-----VGGYGYLLQPLWWV 55
M + N G +LA+ SS FIGSSFI+KKKGL+ A G A + + + L
Sbjct: 1 METDNGKGLILAVASSVFIGSSFILKKKGLKRAGAIGTRADCNNKIISNFKFCL------ 54
Query: 56 GMVTMIVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICIL 115
+IVGE ANFVAY+YAPAVLVTPLGALSII+SAVLAHFLLKEKL+KMG+LGC+ CI+
Sbjct: 55 ----VIVGEAANFVAYIYAPAVLVTPLGALSIIISAVLAHFLLKEKLKKMGVLGCVSCIV 110
Query: 116 GSTIIVLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVY 175
GS +IV+HAP+E + +SV++IW LA QPAFL+Y ++I L L+L+ P GQ+NILVY
Sbjct: 111 GSVVIVIHAPKEQTPNSVEEIWNLATQPAFLIYVAITMSIVLALILHFEPLCGQTNILVY 170
Query: 176 IGICSIFGSLTV 187
IGICS+ G+LTV
Sbjct: 171 IGICSLMGALTV 182
>UniRef100_Q7M563 Non-imprinted in Prader-Willi/Angelman syndrome 2 [Xenopus
tropicalis]
Length = 362
Score = 197 bits (500), Expect = 5e-49
Identities = 96/182 (52%), Positives = 130/182 (70%), Gaps = 1/182 (0%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G +LA+ SS FIG SFI+KKKGL +LAR A GG+ YL + LWW G+++M GE+A
Sbjct: 13 GLVLAISSSLFIGGSFILKKKGLLRLARSGSMRAGQGGHAYLKEWLWWAGLLSMGAGEVA 72
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS++VSA+L+ + L EKL G +GCL+ ILGST++V+HAPQ
Sbjct: 73 NFAAYAFAPATLVTPLGALSVLVSAILSSYFLNEKLNLHGKIGCLLSILGSTVMVIHAPQ 132
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E + S+ ++ P FL++ T + +L L+ PR+GQSNILVYI ICS+ G+L+
Sbjct: 133 EEEIGSLNEMSIKLADPGFLLFATGVVIASLILIFVVGPRHGQSNILVYISICSVIGALS 192
Query: 187 VS 188
VS
Sbjct: 193 VS 194
>UniRef100_Q9VK62 CG12292-PA [Drosophila melanogaster]
Length = 385
Score = 196 bits (497), Expect = 1e-48
Identities = 97/189 (51%), Positives = 132/189 (69%), Gaps = 1/189 (0%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G LA+ S FIGSSFIIKKK L +L+R AS GG+GYL + +WW G++TM VGE A
Sbjct: 48 GVGLAISSCFFIGSSFIIKKKALIRLSRYGEVRASAGGFGYLREWIWWAGLLTMGVGEAA 107
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS+I+SAV+A L EKL +G +GC +CILGSTIIV+H+P+
Sbjct: 108 NFAAYAFAPASLVTPLGALSVIISAVMASRFLNEKLNLLGKIGCFLCILGSTIIVIHSPK 167
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E + +Q ++ + + P F++Y + T+F+ + APR+G +N++VYI +CS GSLT
Sbjct: 168 EKEIEDLQLLFDMLLDPVFILYVICIVGSTVFVACFIAPRHGHTNVVVYIFLCSGIGSLT 227
Query: 187 VSDSTYLSL 195
V L L
Sbjct: 228 VMSCKALGL 236
>UniRef100_UPI00003AD972 UPI00003AD972 UniRef100 entry
Length = 335
Score = 195 bits (496), Expect = 1e-48
Identities = 101/225 (44%), Positives = 149/225 (65%), Gaps = 5/225 (2%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G LA+ SS FIGSSFI+KKKGL +LA P A GGY YL + LWW G+++M +GE A
Sbjct: 43 GLALAVSSSIFIGSSFILKKKGLLKLATKGAPRAGQGGYSYLKEWLWWAGLLSMGLGEAA 102
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS+++SA+L+ + LKEKL G LGC++C+LGST++V+HAP+
Sbjct: 103 NFAAYAFAPATLVTPLGALSVLISAILSSYFLKEKLNIHGKLGCVLCVLGSTVMVIHAPE 162
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E ++S+ ++ PAF+ + +++ L L+ APR GQ+NIL+YI ICS+ G+ +
Sbjct: 163 EEEITSLDEMEIKLQDPAFVAFAVLLMSVALVLIFVVAPRRGQTNILIYILICSLIGAFS 222
Query: 187 VSDSTYLSLFQYSCM----LYRLIFHYTIYHLPILPLRTPHNFVS 227
VS L + + +YR Y + + +L + T N+++
Sbjct: 223 VSSVKGLGIAIKEMLERKPVYRHPLVYILVGILLLSVSTQINYLN 267
>UniRef100_UPI000024CB22 UPI000024CB22 UniRef100 entry
Length = 331
Score = 195 bits (495), Expect = 2e-48
Identities = 95/182 (52%), Positives = 129/182 (70%), Gaps = 1/182 (0%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G LA+ S+ FIG SFI+KKKGL +LA A GGY YL + LWW G+++M +GE A
Sbjct: 8 GLALAVSSTIFIGGSFILKKKGLLRLASKGSTRAGQGGYAYLKEWLWWAGLISMGIGEAA 67
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS++VSAVL+ + L E+L G +GCL+CI GST++VLHAPQ
Sbjct: 68 NFAAYAFAPATLVTPLGALSVLVSAVLSSYFLSERLNIHGKIGCLLCIFGSTVMVLHAPQ 127
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E ++S+ + + P F+ + + +L L+++ APRYGQ N+LVYI ICS+ GSL+
Sbjct: 128 EEEVASLSAMAEKLKDPGFIAFAVCIVVSSLVLIIFVAPRYGQKNVLVYILICSVIGSLS 187
Query: 187 VS 188
VS
Sbjct: 188 VS 189
>UniRef100_UPI0000249C5C UPI0000249C5C UniRef100 entry
Length = 331
Score = 195 bits (495), Expect = 2e-48
Identities = 95/182 (52%), Positives = 129/182 (70%), Gaps = 1/182 (0%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G LA+ S+ FIG SFI+KKKGL +LA A GGY YL + LWW G+++M +GE A
Sbjct: 8 GLALAVSSTIFIGGSFILKKKGLLRLATKGSTRAGQGGYAYLKEWLWWAGLISMGIGEAA 67
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS++VSAVL+ + L E+L G +GCL+CI GST++VLHAPQ
Sbjct: 68 NFAAYAFAPATLVTPLGALSVLVSAVLSSYFLSERLNIHGKIGCLLCIFGSTVMVLHAPQ 127
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E ++S+ + + P F+ + + +L L+++ APRYGQ N+LVYI ICS+ GSL+
Sbjct: 128 EEEVASLSAMAEKLKDPGFIAFAVCIVVSSLVLIIFVAPRYGQKNVLVYILICSVIGSLS 187
Query: 187 VS 188
VS
Sbjct: 188 VS 189
>UniRef100_UPI00003A9E57 UPI00003A9E57 UniRef100 entry
Length = 361
Score = 194 bits (494), Expect = 2e-48
Identities = 94/182 (51%), Positives = 132/182 (71%), Gaps = 1/182 (0%)
Query: 8 GFLLALISSAFIGSSFIIKKKGL-QLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIA 66
G +LA+ SS FIG SFI+KKKGL +LAR A GG+ YL + LWW G+++M GE+A
Sbjct: 13 GLVLAMSSSIFIGGSFILKKKGLLRLARKGSMRAGQGGHAYLKEWLWWAGLLSMGAGEVA 72
Query: 67 NFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQ 126
NF AY +APA LVTPLGALS++VSA+L+ F L EKL G +GCL+ ILGST++V+HAPQ
Sbjct: 73 NFAAYAFAPATLVTPLGALSVLVSAILSSFFLNEKLNLHGKIGCLLSILGSTVMVIHAPQ 132
Query: 127 EMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLT 186
E + ++ ++ P F+++ T + ++L L++ PR+GQ+NILVYI ICS+ G+L+
Sbjct: 133 EEEVETLDEMSNKLRDPGFVVFATLVVIVSLILIVVVGPRHGQTNILVYITICSVIGALS 192
Query: 187 VS 188
VS
Sbjct: 193 VS 194
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 723,064,164
Number of Sequences: 2790947
Number of extensions: 44005141
Number of successful extensions: 1686452
Number of sequences better than 10.0: 22388
Number of HSP's better than 10.0 without gapping: 13390
Number of HSP's successfully gapped in prelim test: 9904
Number of HSP's that attempted gapping in prelim test: 625524
Number of HSP's gapped (non-prelim): 235596
length of query: 358
length of database: 848,049,833
effective HSP length: 128
effective length of query: 230
effective length of database: 490,808,617
effective search space: 112885981910
effective search space used: 112885981910
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148758.11


