

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148758.1 - phase: 0 /pseudo
(143 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
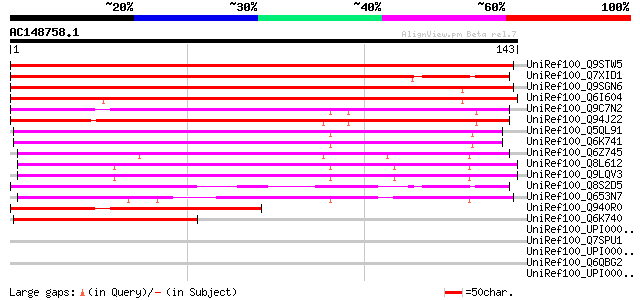
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis tha... 182 2e-45
UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa] 155 2e-37
UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana] 142 2e-33
UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa] 124 3e-28
UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thal... 109 1e-23
UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa] 106 9e-23
UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa] 105 3e-22
UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa] 105 3e-22
UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa] 97 1e-19
UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis tha... 93 1e-18
UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana] 93 1e-18
UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa] 84 7e-16
UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa] 80 1e-14
UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana] 70 1e-11
UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa] 54 7e-07
UniRef100_UPI00002347C9 UPI00002347C9 UniRef100 entry 35 0.46
UniRef100_Q7SPU1 Envelope glycoprotein [Human immunodeficiency v... 35 0.46
UniRef100_UPI0000275224 UPI0000275224 UniRef100 entry 32 2.3
UniRef100_Q6QBG2 Envelope glycoprotein [Human immunodeficiency v... 32 3.9
UniRef100_UPI00002C6EC4 UPI00002C6EC4 UniRef100 entry 31 5.1
>UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis thaliana]
Length = 606
Score = 182 bits (461), Expect = 2e-45
Identities = 89/142 (62%), Positives = 108/142 (75%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV E+ LG RK Q L+FS GPKLY+NTTPVDVG RP+
Sbjct: 325 IEELHQFLEFQLPRQWAPVFSELPLGPQRKQQSCASLQFSFFGPKLYVNTTPVDVGKRPI 384
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
TG+RL LEG RSNRLAIHLQHL+SLPK L D+ N + +S+ +++KV +S+V
Sbjct: 385 TGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLNRSIRQESHDRRYYEKVNWKNYSHV 444
Query: 121 CTAPVESDDSLSIVTGAQLQVE 142
CT PVESDD LS+VTGAQL VE
Sbjct: 445 CTEPVESDDDLSVVTGAQLHVE 466
>UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa]
Length = 608
Score = 155 bits (391), Expect = 2e-37
Identities = 84/144 (58%), Positives = 106/144 (73%), Gaps = 6/144 (4%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV ++ LG RK Q L +++GPKLY+ T VDVG RPV
Sbjct: 325 IEELHQFLEFQLPRQWAPVYSDLPLGPQRKRQSTVSLPVNLIGPKLYVCTNMVDVGKRPV 384
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKV---KRNCF 117
TG+RL LEG RSN+LAIHLQHL SLP+ L L D+ + ++Y +++ + KR F
Sbjct: 385 TGIRLFLEGKRSNKLAIHLQHLCSLPQILQLEDDPYNDQTPEAYDRKYYEPIGSWKR--F 442
Query: 118 SYVCTAPVESDDSLSIVTGAQLQV 141
S+VCTAPVESDDS SIVTGAQL+V
Sbjct: 443 SHVCTAPVESDDS-SIVTGAQLEV 465
>UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana]
Length = 612
Score = 142 bits (358), Expect = 2e-33
Identities = 76/147 (51%), Positives = 100/147 (67%), Gaps = 5/147 (3%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV G++ LG R Q + L+FS++GPKLY+NT+ VD G RPV
Sbjct: 331 IEELHQFLEFQLPRQWAPVYGDLPLGLRRSKQSSPSLQFSLMGPKLYVNTSKVDSGERPV 390
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
TGLR LEG + N LAIHLQHL++ P SL L+ + + ++ VK FS+V
Sbjct: 391 TGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTYEPIEEPVEKGYYVPVKWGIFSHV 450
Query: 121 CTAPVE-----SDDSLSIVTGAQLQVE 142
CT PV+ SDD+ SIVT A L+V+
Sbjct: 451 CTYPVQYNGARSDDTASIVTKAWLEVK 477
>UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa]
Length = 628
Score = 124 bits (312), Expect = 3e-28
Identities = 65/150 (43%), Positives = 96/150 (63%), Gaps = 7/150 (4%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHL--GSYRKHQVNTWLRFSILGPKLYINTTPVDVGNR 58
+E+LH+FLEFQ+PRQWAP GE+ L G +K L+F+++GPKL++ T D GNR
Sbjct: 338 IEELHQFLEFQVPRQWAPEFGELPLALGPRKKKNSLPSLQFTLMGPKLHVTTAKADSGNR 397
Query: 59 PVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFS 118
PVTG+RL LEG ++NRL +HLQHL++ P ++ +A + + + +K S
Sbjct: 398 PVTGIRLFLEGKKNNRLGVHLQHLSATPGTITIAGEVASAEDATVRERDYIEPIKSPLLS 457
Query: 119 YVCTAPVE-----SDDSLSIVTGAQLQVEK 143
+VCTAPV+ DD +IVT A L+V++
Sbjct: 458 HVCTAPVQYNGARIDDCAAIVTRAWLEVQE 487
>UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thaliana]
Length = 561
Score = 109 bits (273), Expect = 1e-23
Identities = 67/154 (43%), Positives = 93/154 (59%), Gaps = 17/154 (11%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+EDL FL++Q+ R WAP + RK V + L+FS++GPKL+I+ V VG +PV
Sbjct: 307 IEDLQYFLDYQIARAWAPEQSNLQ----RKEPVCSSLQFSLMGPKLFISADQVTVGRKPV 362
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSL-PLADN-----ANAYLSCDSYSCTFHKKVKR 114
TGLRL LEGS+ NRL+IHLQHL SLPK L P D+ A + + + + +K
Sbjct: 363 TGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVPIGAPKWQGPEEQDSRWFEPIKW 422
Query: 115 NCFSYVCTAPVESDDS-------LSIVTGAQLQV 141
FS+V T+P+E ++ + IVTGAQL V
Sbjct: 423 KNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGV 456
>UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa]
Length = 553
Score = 106 bits (265), Expect = 9e-23
Identities = 66/154 (42%), Positives = 94/154 (60%), Gaps = 14/154 (9%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+L FL+FQ+ WAPV I G +RK V L+FS++GPKL+++T + VG RPV
Sbjct: 303 IEELQYFLDFQVQLVWAPVPPGI-AGQHRKEPVCPSLQFSLMGPKLFVSTEQISVGRRPV 361
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPK-SLPLADN-----ANAYLSCDSYSCTFHKKVKR 114
TGL+L LEG++ NRLAIHLQHL SLPK +P D+ + + + + +K
Sbjct: 362 TGLKLCLEGAKQNRLAIHLQHLGSLPKIFVPHWDSHITIGPPKWQGPEEQDSRWFEPIKW 421
Query: 115 NCFSYVCTAPVESDDS-------LSIVTGAQLQV 141
F++V TAP+E ++ + IVTGAQL V
Sbjct: 422 RNFAHVSTAPIEYTETSITDLSGVYIVTGAQLGV 455
>UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa]
Length = 551
Score = 105 bits (261), Expect = 3e-22
Identities = 58/150 (38%), Positives = 87/150 (57%), Gaps = 12/150 (8%)
Query: 2 EDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
EDL FLEFQ+P QWAP+ E+ LG ++ L+F LGPKL ++T+ V ++PV
Sbjct: 241 EDLQHFLEFQVPLQWAPLFNELILGPQKRKGSYPSLQFRFLGPKLQVSTSQVSSSHKPVV 300
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSL--PLADNANAYLSCDSYSCTFHKKVKRNCFSY 119
GLRL LEG + NRLAIH+QHL+S P L L+ + + + + + + ++ +S
Sbjct: 301 GLRLYLEGRKCNRLAIHVQHLSSAPSMLGDSLSSSMSEWRESEEVGAGYIEPIQWKSYSC 360
Query: 120 VCTAPVESDD----------SLSIVTGAQL 139
VCT+ V+ + + +VTGAQL
Sbjct: 361 VCTSKVDYNPEWLKRVPGGRGVFVVTGAQL 390
>UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa]
Length = 634
Score = 105 bits (261), Expect = 3e-22
Identities = 58/150 (38%), Positives = 87/150 (57%), Gaps = 12/150 (8%)
Query: 2 EDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
EDL FLEFQ+P QWAP+ E+ LG ++ L+F LGPKL ++T+ V ++PV
Sbjct: 324 EDLQHFLEFQVPLQWAPLFNELILGPQKRKGSYPSLQFRFLGPKLQVSTSQVSSSHKPVV 383
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSL--PLADNANAYLSCDSYSCTFHKKVKRNCFSY 119
GLRL LEG + NRLAIH+QHL+S P L L+ + + + + + + ++ +S
Sbjct: 384 GLRLYLEGRKCNRLAIHVQHLSSAPSMLGDSLSSSMSEWRESEEVGAGYIEPIQWKSYSC 443
Query: 120 VCTAPVESDD----------SLSIVTGAQL 139
VCT+ V+ + + +VTGAQL
Sbjct: 444 VCTSKVDYNPEWLKRVPGGRGVFVVTGAQL 473
>UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa]
Length = 620
Score = 96.7 bits (239), Expect = 1e-19
Identities = 62/154 (40%), Positives = 88/154 (56%), Gaps = 15/154 (9%)
Query: 3 DLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNT-WLRFSILGPKLYINTTPVDVGNRPVT 61
DL FL+FQ WAPVLGE+ LG Q ++ L FS+LG KLY+++T V V PVT
Sbjct: 324 DLRYFLDFQHHCVWAPVLGELPLGPCSHRQGSSPALHFSLLGSKLYVSSTEVVVPKLPVT 383
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPK--SLPLADNANAYLSCDSYS-CTFHKKVKRNCFS 118
G+RL LEG ++NRL IHLQHL++ P + AD + ++ + +++ V+ +
Sbjct: 384 GMRLHLEGKKNNRLGIHLQHLSTTPTFVAAARADKPPVWRGTEAVTDDRYYEPVQWRMLA 443
Query: 119 YVCTAPVESD-----------DSLSIVTGAQLQV 141
VCTAPV+ D + +V GAQL V
Sbjct: 444 RVCTAPVKYDPRWCAGDRRRRPAACVVAGAQLHV 477
>UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis thaliana]
Length = 627
Score = 93.2 bits (230), Expect = 1e-18
Identities = 60/159 (37%), Positives = 83/159 (51%), Gaps = 18/159 (11%)
Query: 3 DLHRFLEFQLPRQWAPVLGEIHLGSY-RKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
DL FL+F PR WAPV ++ G+ L + +GPKLY+NTTPV PVT
Sbjct: 334 DLQYFLDFSGPRAWAPVHNDLPFGAAPNMASAYPALHINFMGPKLYVNTTPVTSEKNPVT 393
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSL-PLADNANAYLSCDSYSCT--FHKKVKRNCFS 118
G+R LEG + NRLAIHLQHL + ++ + + + D + + + + FS
Sbjct: 394 GMRFFLEGKKCNRLAIHLQHLDNTRTTVGEKITDEHIWRGSDQITDNDRYFEPLNGKKFS 453
Query: 119 YVCTAPVESD--------------DSLSIVTGAQLQVEK 143
+VCT PV+ D D IVTGAQL+V+K
Sbjct: 454 HVCTVPVKYDPNWIKTTSNHKSQNDVAFIVTGAQLEVKK 492
>UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana]
Length = 645
Score = 93.2 bits (230), Expect = 1e-18
Identities = 60/159 (37%), Positives = 83/159 (51%), Gaps = 18/159 (11%)
Query: 3 DLHRFLEFQLPRQWAPVLGEIHLGSY-RKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
DL FL+F PR WAPV ++ G+ L + +GPKLY+NTTPV PVT
Sbjct: 352 DLQYFLDFSGPRAWAPVHNDLPFGAAPNMASAYPALHINFMGPKLYVNTTPVTSEKNPVT 411
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSL-PLADNANAYLSCDSYSCT--FHKKVKRNCFS 118
G+R LEG + NRLAIHLQHL + ++ + + + D + + + + FS
Sbjct: 412 GMRFFLEGKKCNRLAIHLQHLDNTRTTVGEKITDEHIWRGSDQITDNDRYFEPLNGKKFS 471
Query: 119 YVCTAPVESD--------------DSLSIVTGAQLQVEK 143
+VCT PV+ D D IVTGAQL+V+K
Sbjct: 472 HVCTVPVKYDPNWIKTTSNHKSQNDVAFIVTGAQLEVKK 510
>UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa]
Length = 573
Score = 84.0 bits (206), Expect = 7e-16
Identities = 58/141 (41%), Positives = 75/141 (53%), Gaps = 35/141 (24%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV ++ LG RK Q + L +++GPKLY+ T +
Sbjct: 326 VEELHQFLEFQLPRQWAPVYSDLPLGPQRKRQSSASLPVNLIGPKLYVCTNMI------- 378
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
+QLE N P A+ Y S+ KR FS+V
Sbjct: 379 ----IQLEDDTYN-------------PQTPEAEIRKYYEPIGSW--------KR--FSHV 411
Query: 121 CTAPVESDDSLSIVTGAQLQV 141
CTAPV+SDDS SIVTGA L+V
Sbjct: 412 CTAPVDSDDS-SIVTGAHLEV 431
>UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa]
Length = 614
Score = 80.1 bits (196), Expect = 1e-14
Identities = 57/162 (35%), Positives = 84/162 (51%), Gaps = 38/162 (23%)
Query: 3 DLHRFLEFQLPRQWAPVLGEIHLGSYRKHQ-VNTWLRFS-ILGPKLYINTTPVDVGNRPV 60
+L FL+FQ R WAPVL ++ LG Q N L FS ++ PKL P+
Sbjct: 331 ELRYFLDFQHHRLWAPVLSDLPLGLCSNRQGTNPALHFSLVIVPKL------------PI 378
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSL-------PLADNANAYLSCDSYSCTFHKKVK 113
TG+RL LEG ++NRL IHLQHL++ P + P A + ++ + Y ++ V+
Sbjct: 379 TGMRLHLEGKKNNRLGIHLQHLSTTPTFIAGGWSGRPPAWRGSEAIADERY----YEPVQ 434
Query: 114 RNCFSYVCTAPVESD-------------DSLSIVTGAQLQVE 142
R F++VCT PV+ D + +V+GAQL V+
Sbjct: 435 RRMFAHVCTVPVKHDPRWLAAGDGGGGRPAAYVVSGAQLHVK 476
>UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana]
Length = 383
Score = 70.1 bits (170), Expect = 1e-11
Identities = 35/71 (49%), Positives = 48/71 (67%), Gaps = 4/71 (5%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+EDL FL++Q+ R WAP + RK V + L+FS++GPKL+I+ V VG +PV
Sbjct: 307 IEDLQYFLDYQIARAWAPEQSNLQ----RKEPVCSSLQFSLMGPKLFISADQVTVGRKPV 362
Query: 61 TGLRLQLEGSR 71
TGLRL LEGS+
Sbjct: 363 TGLRLSLEGSK 373
>UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa]
Length = 406
Score = 53.9 bits (128), Expect = 7e-07
Identities = 25/52 (48%), Positives = 34/52 (65%)
Query: 2 EDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPV 53
EDL FLEFQ+P QWAP+ E+ LG ++ L+F LGPKL ++T+ V
Sbjct: 324 EDLQHFLEFQVPLQWAPLFNELILGPQKRKGSYPSLQFRFLGPKLQVSTSQV 375
>UniRef100_UPI00002347C9 UPI00002347C9 UniRef100 entry
Length = 870
Score = 34.7 bits (78), Expect = 0.46
Identities = 23/64 (35%), Positives = 31/64 (47%), Gaps = 6/64 (9%)
Query: 44 PKLYINTTPVDVGNRPVT-GLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCD 102
P L NTTP +P T L L +R N A+H + P P+AD + Y +CD
Sbjct: 675 PLLAENTTPSSPTTQPGTLHPELPLPAARGNTSALH-----ATPTIQPIADGRSMYGNCD 729
Query: 103 SYSC 106
+ SC
Sbjct: 730 NESC 733
>UniRef100_Q7SPU1 Envelope glycoprotein [Human immunodeficiency virus 1]
Length = 843
Score = 34.7 bits (78), Expect = 0.46
Identities = 19/65 (29%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 38 RFSILGPKLYINTTPVDVGNRPVTGLRLQLEGS-RSNRLAIHLQHLASLPKSLPLADNAN 96
+F+ GP ++T GN+PV +L L GS +R+ I Q+++S K++ + N +
Sbjct: 232 KFNGTGPCTNVSTVQCTHGNKPVVSTQLLLNGSLAEDRIVIRSQNISSNTKNIIVHLNES 291
Query: 97 AYLSC 101
++C
Sbjct: 292 VQINC 296
>UniRef100_UPI0000275224 UPI0000275224 UniRef100 entry
Length = 242
Score = 32.3 bits (72), Expect = 2.3
Identities = 16/38 (42%), Positives = 22/38 (57%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLR 38
+ED+H +E +L P+ G+IH G R QV T LR
Sbjct: 78 LEDIHMNIESRLYDLIGPIAGKIHTGKSRNDQVATSLR 115
>UniRef100_Q6QBG2 Envelope glycoprotein [Human immunodeficiency virus 1]
Length = 352
Score = 31.6 bits (70), Expect = 3.9
Identities = 16/64 (25%), Positives = 32/64 (50%)
Query: 38 RFSILGPKLYINTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANA 97
+F+ GP ++T G RPV +L L GS + + I ++ + K++ + N +
Sbjct: 113 KFNGTGPCTNVSTVQCTHGIRPVVSTQLLLNGSLAEEIVIRSENFTNNAKTIIVQLNESV 172
Query: 98 YLSC 101
++C
Sbjct: 173 VINC 176
>UniRef100_UPI00002C6EC4 UPI00002C6EC4 UniRef100 entry
Length = 143
Score = 31.2 bits (69), Expect = 5.1
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGN 57
+ED+H +E +L P G+IH G R QV+T R + K IN + + + N
Sbjct: 79 LEDIHMNIESRLHDMVGPFAGKIHSGKSRNDQVSTIERMYV---KNAINESLIKIEN 132
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 245,242,954
Number of Sequences: 2790947
Number of extensions: 9608508
Number of successful extensions: 21561
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 21515
Number of HSP's gapped (non-prelim): 32
length of query: 143
length of database: 848,049,833
effective HSP length: 119
effective length of query: 24
effective length of database: 515,927,140
effective search space: 12382251360
effective search space used: 12382251360
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148758.1


