

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148218.5 - phase: 0
(164 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
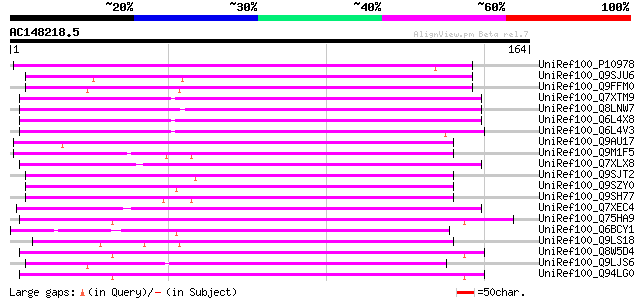
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_P10978 Retrovirus-related Pol polyprotein from transpo... 90 2e-17
UniRef100_Q9SJU6 Putative retroelement pol polyprotein [Arabidop... 87 1e-16
UniRef100_Q9FFM0 Copia-like retrotransposable element [Arabidops... 86 3e-16
UniRef100_Q7XTM9 OSJNBa0033G05.13 protein [Oryza sativa] 86 4e-16
UniRef100_Q8LNW7 Putative polyprotein [Oryza sativa] 85 8e-16
UniRef100_Q6L4X8 Putative polyprotein [Oryza sativa] 85 8e-16
UniRef100_Q6L4V3 Putative polyprotein [Oryza sativa] 84 1e-15
UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense] 84 1e-15
UniRef100_Q9M1F5 Copia-like polyprotein [Arabidopsis thaliana] 83 3e-15
UniRef100_Q7XLX8 OSJNBa0042I15.12 protein [Oryza sativa] 82 6e-15
UniRef100_Q9SJT2 Putative retroelement pol polyprotein [Arabidop... 79 4e-14
UniRef100_Q9SZY0 Putative retrotransposon [Arabidopsis thaliana] 78 7e-14
UniRef100_Q9SH77 Putative retroelement pol polyprotein [Arabidop... 77 1e-13
UniRef100_Q7XEC4 Putative polyprotein [Oryza sativa] 74 1e-12
UniRef100_Q75HA9 Putative polyprotein [Oryza sativa] 72 4e-12
UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas] 72 5e-12
UniRef100_Q9LS18 Retroelement pol polyprotein-like [Arabidopsis ... 70 1e-11
UniRef100_Q8W5D4 Putative retrotransposon-related protein [Oryza... 70 1e-11
UniRef100_Q9LJS6 Copia retroelement pol polyprotein-like [Arabid... 70 2e-11
UniRef100_Q94LG0 Putative retroelement pol polyprotein [Oryza sa... 69 3e-11
>UniRef100_P10978 Retrovirus-related Pol polyprotein from transposon TNT 1-94
[Contains: Protease (EC 3.4.23.-); Reverse transcriptase
(EC 2.7.7.49); Endonuclease] [Nicotiana tabacum]
Length = 1328
Score = 90.1 bits (222), Expect = 2e-17
Identities = 51/147 (34%), Positives = 81/147 (54%), Gaps = 2/147 (1%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSA 61
G K+++ KF GDN F W+ +M +LIQQ K L +S P TM + ++ ++A SA
Sbjct: 3 GVKYEVAKFNGDNGFSTWQRRMRDLLIQQGLHKVLDVDSKKPDTMKAEDWADLDERAASA 62
Query: 62 IVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELL 121
I L L + + E TA +W++ E LYM+K+L ++ LK+QLY+ M + + L
Sbjct: 63 IRLHLSDDVVNNIIDEDTARGIWTRLESLYMSKTLTNKLYLKKQLYALHMSEGTNFLSHL 122
Query: 122 VEFNKIIGDFEN--IKLHLEDAGALMV 146
FN +I N +K+ ED L++
Sbjct: 123 NVFNGLITQLANLGVKIEEEDKAILLL 149
>UniRef100_Q9SJU6 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 838
Score = 87.4 bits (215), Expect = 1e-16
Identities = 56/153 (36%), Positives = 83/153 (53%), Gaps = 12/153 (7%)
Query: 6 DIEKFTGDNDFRLWKVKMEA------VLIQQKCEKTLKGESSLPVTMSRAEKTE------ 53
++EK G+ D+ LWK K+ A +L + ++ ++ E S T S KTE
Sbjct: 7 EVEKLDGEGDYVLWKEKLLAHIELLGLLEGLEEDEAIEEEESTAETDSLLTKTEDKVLKE 66
Query: 54 MVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVK 113
KARS ++L LGN LR+ KE TA M + L+M KSL +R LKQ+LY +KM
Sbjct: 67 KRGKARSTVILSLGNHVLRKVIKEKTAAGMIRVLDKLFMAKSLPNRIYLKQRLYGYKMSD 126
Query: 114 SKVVVELLVEFNKIIGDFENIKLHLEDAGALMV 146
S + E + +F K+I D EN+K+ + D +V
Sbjct: 127 SMTIEENVNDFFKLISDLENVKVSVPDEDQAIV 159
>UniRef100_Q9FFM0 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1342
Score = 85.9 bits (211), Expect = 3e-16
Identities = 56/161 (34%), Positives = 81/161 (49%), Gaps = 20/161 (12%)
Query: 6 DIEKFTGDNDFRLWKVKM-----------------EAVLIQQKCEKTLKGESSLPVTMSR 48
++EKF GD D+ LWK K+ EAV+ E + G S+
Sbjct: 7 EVEKFDGDGDYILWKEKLLAHMEMLGLLEGLGEEEEAVVEDSTTEISDGGNQDPETATSK 66
Query: 49 AEKT---EMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQ 105
E E KARS I+L LGN LR+ K+ TA M + L+M KSL +R LKQ+
Sbjct: 67 LEDKILKEKRGKARSTIILSLGNNVLRKVIKQKTAAGMIKVLDQLFMAKSLPNRIYLKQR 126
Query: 106 LYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLEDAGALMV 146
LY +KM ++ + E + +F K+I D EN+K+ + D +V
Sbjct: 127 LYGYKMSENMTMEENVNDFFKLISDLENVKVVVPDEDQAIV 167
>UniRef100_Q7XTM9 OSJNBa0033G05.13 protein [Oryza sativa]
Length = 1181
Score = 85.5 bits (210), Expect = 4e-16
Identities = 51/146 (34%), Positives = 76/146 (51%), Gaps = 1/146 (0%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIV 63
K+D+ D F LW+VKM AVL QQ+ + L G S EK + KA S I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQELDDALSGFDKRTQDWSNDEK-KRDRKAMSYIH 63
Query: 64 LCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVE 123
L L N L+E KE TA +W K E + MTK L + LKQ+L+ K+ V++ L
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKLQDDGSVMDHLSA 123
Query: 124 FNKIIGDFENIKLHLEDAGALMVWCC 149
F +I+ D E++++ ++ ++ C
Sbjct: 124 FKEIVADLESMEVKYDEKDLALILLC 149
>UniRef100_Q8LNW7 Putative polyprotein [Oryza sativa]
Length = 1280
Score = 84.7 bits (208), Expect = 8e-16
Identities = 52/146 (35%), Positives = 75/146 (50%), Gaps = 1/146 (0%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIV 63
K+D+ D F LW+VKM AVL QQ + L G S EK + KA S I
Sbjct: 40 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEKKKD-RKAMSYIH 98
Query: 64 LCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVE 123
L L N L+E KE TA +W K E + MTK L + LKQ+L+ K+ V++ L
Sbjct: 99 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKLQDDGSVMDHLST 158
Query: 124 FNKIIGDFENIKLHLEDAGALMVWCC 149
F +I+ D E+I++ ++ ++ C
Sbjct: 159 FKEIVADLESIEVKYDEEDLGLILLC 184
>UniRef100_Q6L4X8 Putative polyprotein [Oryza sativa]
Length = 1211
Score = 84.7 bits (208), Expect = 8e-16
Identities = 51/146 (34%), Positives = 74/146 (49%), Gaps = 1/146 (0%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIV 63
K+D+ D F LW+VKM AVL QQ + L G S EK + K S I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEK-KRDRKTMSYIH 63
Query: 64 LCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVE 123
L L N L+E KE TA +W K E + MTK L + LKQ+L+ K+ V++ L
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKLQDDGSVMDHLSA 123
Query: 124 FNKIIGDFENIKLHLEDAGALMVWCC 149
F KI+ D E++++ ++ ++ C
Sbjct: 124 FKKIVADLESMEVKYDEEDLCLILLC 149
>UniRef100_Q6L4V3 Putative polyprotein [Oryza sativa]
Length = 1243
Score = 84.3 bits (207), Expect = 1e-15
Identities = 55/148 (37%), Positives = 79/148 (53%), Gaps = 2/148 (1%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIV 63
K+D+ D F LW+VKM AVL QQ + L G S EK + KA S I
Sbjct: 5 KYDLPLLDRDTRFSLWQVKMRAVLAQQDLDDALSGFDKRTQDWSNDEK-KRDRKAISYIH 63
Query: 64 LCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVE 123
L L N L+E KE TA +W K E + MTK L + LKQ+L+ K+ + V++ L
Sbjct: 64 LHLSNNILQEVLKEETAAGLWLKLEQICMTKDLTSKMHLKQKLFLHKLQDDESVMDHLSA 123
Query: 124 FNKIIGDFENIKL-HLEDAGALMVWCCL 150
F +I+ D E++++ + ED L++ C L
Sbjct: 124 FKEIVADLESMEVKYDEDDLGLILLCSL 151
>UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense]
Length = 1328
Score = 84.0 bits (206), Expect = 1e-15
Identities = 48/140 (34%), Positives = 79/140 (56%), Gaps = 1/140 (0%)
Query: 2 GSKWDIEKFTGDND-FRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARS 60
G K+++ KF GD F +W+ +M+ +LIQQ K L G+S P +M + E+ +KA S
Sbjct: 3 GVKYEVAKFNGDKPVFSMWQRRMKDLLIQQGLHKALGGKSKKPESMKLEDWEELDEKAAS 62
Query: 61 AIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVEL 120
AI L L + + E +A +W+K E LYM+K+L ++ LK+QLY+ M + +
Sbjct: 63 AIRLHLTDDVVNNIVDEESACGIWTKLENLYMSKTLTNKLYLKKQLYTLHMDEGTNFLSH 122
Query: 121 LVEFNKIIGDFENIKLHLED 140
L N +I N+ + +E+
Sbjct: 123 LNVLNGLITQLANLGVKIEE 142
>UniRef100_Q9M1F5 Copia-like polyprotein [Arabidopsis thaliana]
Length = 1363
Score = 82.8 bits (203), Expect = 3e-15
Identities = 54/155 (34%), Positives = 84/155 (53%), Gaps = 17/155 (10%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSR------------A 49
G++ ++EKF G D+ +WK K+ A + L+ ES P+ R
Sbjct: 3 GARIEVEKFDGRGDYTMWKEKLLAHIDMLGLSAVLR-ESETPMGKERDSEKSDEDEKEER 61
Query: 50 EKTEMVD----KARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQ 105
EK E + KARS IVL + ++ LR+ KET+A +M + LYM+K+L +R LKQ+
Sbjct: 62 EKMEAFEEKKRKARSTIVLSVSDRVLRKIKKETSAAAMLEALDRLYMSKALPNRIYLKQK 121
Query: 106 LYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
LYSFKM ++ + + EF I+ D EN+ + + D
Sbjct: 122 LYSFKMSENLSIEGNIDEFLHIVADLENLNVLVSD 156
>UniRef100_Q7XLX8 OSJNBa0042I15.12 protein [Oryza sativa]
Length = 432
Score = 81.6 bits (200), Expect = 6e-15
Identities = 44/146 (30%), Positives = 80/146 (54%), Gaps = 2/146 (1%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIV 63
K+D+ KF G +F LW+ +++ +L QQ+ K LKG P M + EM +A + I
Sbjct: 43 KFDLVKFDGFGNFGLWQTRVKDLLAQQEVSKALKGVK--PAKMEDDDCEEMQLQAAATIR 100
Query: 64 LCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVE 123
CL ++ + +ET+ +W K E +M+K+L +++ LKQ+LY KM + + +
Sbjct: 101 RCLSDQIMYHVMEETSPKIIWEKLEAQFMSKTLTNKRYLKQKLYGLKMQEGADLTAHVNV 160
Query: 124 FNKIIGDFENIKLHLEDAGALMVWCC 149
FN+++ D + + ++D +V C
Sbjct: 161 FNQLVTDLVKMDVEVDDEDKAIVLLC 186
>UniRef100_Q9SJT2 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1333
Score = 79.0 bits (193), Expect = 4e-14
Identities = 52/150 (34%), Positives = 79/150 (52%), Gaps = 15/150 (10%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDK-------- 57
++EKF G D+ +WK K+ A L LK E L ++ + TE +K
Sbjct: 7 EVEKFDGRGDYTMWKEKLMAHLDILGLSVALKEEDDLVEKVAEMQLTEEEEKEEVLRREL 66
Query: 58 -------ARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFK 110
ARSAIVL + ++ LR+ KE +A +M + LYM+K+L +R KQ+LYSFK
Sbjct: 67 LEEKRRKARSAIVLSVTDRVLRKIKKEQSAAAMLGVLDKLYMSKALPNRIYQKQKLYSFK 126
Query: 111 MVKSKVVVELLVEFNKIIGDFENIKLHLED 140
M ++ + + EF +II D EN + + D
Sbjct: 127 MSENLSIEGNIDEFLRIIADLENTNVLVSD 156
>UniRef100_Q9SZY0 Putative retrotransposon [Arabidopsis thaliana]
Length = 1230
Score = 78.2 bits (191), Expect = 7e-14
Identities = 51/148 (34%), Positives = 78/148 (52%), Gaps = 13/148 (8%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEK-------------T 52
++EKF G D+ LWK K+ A + L+ S+ + E+
Sbjct: 7 EMEKFDGHGDYTLWKEKLMAHMDLLGLTVALRETQSVSDPLESEEEGKESEKGDKEALME 66
Query: 53 EMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMV 112
E KARS IVL + ++ LR++ KE TA SM + LYM+K+L +R LKQ+LYS+KM
Sbjct: 67 EKRQKARSTIVLSVSDQVLRKSKKEKTAPSMLEALDKLYMSKALPNRIYLKQKLYSYKMQ 126
Query: 113 KSKVVVELLVEFNKIIGDFENIKLHLED 140
++ V + EF ++I D EN + + D
Sbjct: 127 ENLSVEGNIDEFLRLIADLENTNVLVSD 154
>UniRef100_Q9SH77 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1356
Score = 77.4 bits (189), Expect = 1e-13
Identities = 54/150 (36%), Positives = 81/150 (54%), Gaps = 15/150 (10%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMS-----------RAEKTEM 54
++EKF G D+ +WK K+ A + LK S S + EK E
Sbjct: 7 EVEKFDGRGDYTMWKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEKFEA 66
Query: 55 VD----KARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFK 110
++ KARSAIVL + ++ LR+ KE+TA +M + LYM+K+L +R KQ+LYSFK
Sbjct: 67 LEEKKKKARSAIVLSVTDRVLRKIKKESTAAAMLLALDKLYMSKALPNRIYPKQKLYSFK 126
Query: 111 MVKSKVVVELLVEFNKIIGDFENIKLHLED 140
M ++ V + EF +II D EN+ + + D
Sbjct: 127 MSENLSVEGNIDEFLQIITDLENMNVIISD 156
>UniRef100_Q7XEC4 Putative polyprotein [Oryza sativa]
Length = 382
Score = 74.3 bits (181), Expect = 1e-12
Identities = 40/147 (27%), Positives = 79/147 (53%), Gaps = 2/147 (1%)
Query: 3 SKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAI 62
SK+++ KF G +F LW+++++ +L QQ K L + ++P + + EM +A + I
Sbjct: 7 SKFEVVKFDGTGNFVLWQMRLKDLLAQQGISKAL--QETMPEKIDADKWNEMKAQAAATI 64
Query: 63 VLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLV 122
L L + + + E + +W K L+M+KSL + LKQQLY ++ + + + +
Sbjct: 65 RLSLSDSVMYQVMDEKSPKEIWDKLASLHMSKSLTSKLYLKQQLYGLQVQEESDLRKHVD 124
Query: 123 EFNKIIGDFENIKLHLEDAGALMVWCC 149
FN+++ D + + L+D ++ C
Sbjct: 125 VFNQLVVDLSKLDVKLDDEDKAIILLC 151
>UniRef100_Q75HA9 Putative polyprotein [Oryza sativa]
Length = 1322
Score = 72.4 bits (176), Expect = 4e-12
Identities = 47/158 (29%), Positives = 78/158 (48%), Gaps = 2/158 (1%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQK-CEKTLKGESSLPVTMSRAEKTEMVDKARSAI 62
K+D+ F LW+VKM AVL Q ++ L+ T AE+ KA S I
Sbjct: 5 KYDLPLLDYKTRFSLWQVKMRAVLAQTSDLDEALESFGKKKTTEWTAEEKRKDRKALSLI 64
Query: 63 VLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLV 122
L L N L+E ++ TA +W K E + M+K L + +K +L+S K+ +S V+ +
Sbjct: 65 QLHLSNDILQEVLQKKTAAELWLKLESICMSKDLTSKMHIKMKLFSHKLHESGSVLNHIS 124
Query: 123 EFNKIIGDFENIKLHLEDAG-ALMVWCCLEDEEGDVSH 159
F +I+ D ++++ +D L++ C L + H
Sbjct: 125 VFKEIVADLVSMEVQFDDEDLGLLLLCSLPSSYANFRH 162
>UniRef100_Q6BCY1 Gag-Pol [Ipomoea batatas]
Length = 1298
Score = 72.0 bits (175), Expect = 5e-12
Identities = 45/140 (32%), Positives = 75/140 (53%), Gaps = 5/140 (3%)
Query: 1 MGSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEK-TEMVDKAR 59
M +K++IEKF G N F LWK+K++A+L + C L S PV + +K +EM + A
Sbjct: 1 MAAKFEIEKFNGKN-FSLWKLKVKAILRKDNC---LAAISERPVDFTDDKKWSEMNEDAM 56
Query: 60 SAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVE 119
+ + L + + L ++ TA +W LY KSL ++ LK++LY+ +M +S V E
Sbjct: 57 ADLYLSIADGVLSSIEEKKTANEIWDHLNRLYEAKSLHNKIFLKRKLYTLRMSESTSVTE 116
Query: 120 LLVEFNKIIGDFENIKLHLE 139
L N + ++ +E
Sbjct: 117 HLNTLNTLFSQLTSLSCKIE 136
>UniRef100_Q9LS18 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1338
Score = 70.5 bits (171), Expect = 1e-11
Identities = 49/153 (32%), Positives = 80/153 (52%), Gaps = 20/153 (13%)
Query: 8 EKFTGDNDFRLWKVKMEAVL--------IQQKCEKTLKGESS-LPVTMSRAEKT------ 52
E+F G D+ LWK K+ A L +++K EK E+ + V S +E+
Sbjct: 18 ERFDGRGDYTLWKRKLLAQLEVMGISDALKEKEEKKEAVETERVKVVSSSSERRREEHKK 77
Query: 53 -----EMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLY 107
E +KARS I+L + + LR E TA M S + LY++ L+ R LK++L+
Sbjct: 78 DHSREEKENKARSVIILSVADNILRRIRTEETAAGMISVLDKLYLSDPLSSRISLKRKLF 137
Query: 108 SFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
FKM ++K V E + +F +I+ D E + +++ D
Sbjct: 138 EFKMSENKAVEENIEDFFRIVEDLEKLDVYVSD 170
>UniRef100_Q8W5D4 Putative retrotransposon-related protein [Oryza sativa]
Length = 1229
Score = 70.5 bits (171), Expect = 1e-11
Identities = 47/149 (31%), Positives = 76/149 (50%), Gaps = 2/149 (1%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQK-CEKTLKGESSLPVTMSRAEKTEMVDKARSAI 62
K+D+ F LW+VKM AVL Q ++ L+ T AE+ KA S I
Sbjct: 2 KYDLPLQDYKTRFSLWQVKMRAVLAQTSDLDEALESFGKKKTTEWTAEEKRKDRKALSLI 61
Query: 63 VLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLV 122
L L N L++ +E TA +W K E + M+K L + +K +L+S K+ +S V+ +
Sbjct: 62 QLHLSNDILQKVLQEKTAAELWFKLESICMSKDLTSKMHIKMKLFSHKLQESGSVLNHIS 121
Query: 123 EFNKIIGDFENIKLHLEDAG-ALMVWCCL 150
F +II D ++++ +D L++ C L
Sbjct: 122 VFKEIIADLVSMEVQFDDEDLGLLLLCSL 150
>UniRef100_Q9LJS6 Copia retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 526
Score = 70.1 bits (170), Expect = 2e-11
Identities = 47/160 (29%), Positives = 75/160 (46%), Gaps = 28/160 (17%)
Query: 6 DIEKFTGDNDFRLWKVKM---------------------------EAVLIQQKCEKTLKG 38
++++F G DF LWKV+M EA + + + G
Sbjct: 8 EVDRFDGTGDFSLWKVRMLAHFGVLGLKGILNEEQLLRDPPATEEEAAVAGRDYHVGIAG 67
Query: 39 ESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAH 98
+ LP T+ K E +KA+ IVL +GN+ LR+ T +MWS LYM SL +
Sbjct: 68 DFELPPTVDPV-KFEKSEKAKDLIVLNVGNQVLRKIKNCETVAAMWSTLNKLYMETSLPN 126
Query: 99 RQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHL 138
R L + Y++KM SK + + +F K++ D NI +++
Sbjct: 127 RIYLHLKFYTYKMTDSKSIDGNVDDFLKLVTDLNNIGVNV 166
>UniRef100_Q94LG0 Putative retroelement pol polyprotein [Oryza sativa]
Length = 1326
Score = 69.3 bits (168), Expect = 3e-11
Identities = 46/149 (30%), Positives = 75/149 (49%), Gaps = 2/149 (1%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVLIQQK-CEKTLKGESSLPVTMSRAEKTEMVDKARSAI 62
K+D+ F LW+VKM A+L Q ++ L+ T AE+ KA I
Sbjct: 5 KYDLPLLDYKTRFSLWQVKMRAILAQTSDLDEALESFGKKKSTEWTAEEKRKDRKALLLI 64
Query: 63 VLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLV 122
L L N L+E +E TA +W K E + M+K L + +K +L+S K+ +S V+ +
Sbjct: 65 QLHLSNDILQEVLQEKTAAELWLKLESICMSKDLTSKMHIKMKLFSHKLQESGSVLNHIS 124
Query: 123 EFNKIIGDFENIKLHLEDAG-ALMVWCCL 150
F +I+ D +I++ +D L++ C L
Sbjct: 125 VFKEIVVDLVSIEVQFDDEDLGLLLLCSL 153
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.131 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 257,747,452
Number of Sequences: 2790947
Number of extensions: 8793626
Number of successful extensions: 22405
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 110
Number of HSP's that attempted gapping in prelim test: 22286
Number of HSP's gapped (non-prelim): 193
length of query: 164
length of database: 848,049,833
effective HSP length: 117
effective length of query: 47
effective length of database: 521,509,034
effective search space: 24510924598
effective search space used: 24510924598
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC148218.5


