

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147875.9 + phase: 0
(122 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
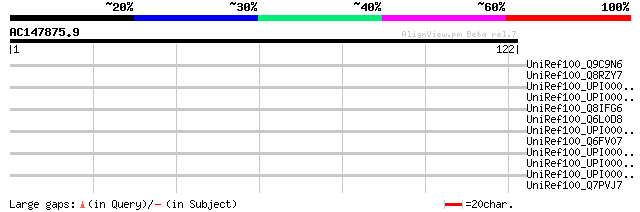
UniRef100_Q9C9N6 Hypothetical protein F4N21_22 [Arabidopsis thal... 37 0.13
UniRef100_Q8RZY7 Myosin heavy chain-like protein [Oryza sativa] 34 0.87
UniRef100_UPI00004999BB UPI00004999BB UniRef100 entry 33 1.5
UniRef100_UPI0000368EE6 UPI0000368EE6 UniRef100 entry 32 4.3
UniRef100_Q8IFG6 Hypothetical protein Tb927.1.760 [Trypanosoma b... 32 4.3
UniRef100_Q6L0D8 Iron-dependent repressor [Picrophilus torridus] 32 4.3
UniRef100_UPI000042DC3D UPI000042DC3D UniRef100 entry 31 7.4
UniRef100_Q6FV07 Similar to sp|P38991 Saccharomyces cerevisiae Y... 31 7.4
UniRef100_UPI00004328CB UPI00004328CB UniRef100 entry 30 9.6
UniRef100_UPI000042FCB7 UPI000042FCB7 UniRef100 entry 30 9.6
UniRef100_UPI00002F403D UPI00002F403D UniRef100 entry 30 9.6
UniRef100_Q7PVJ7 ENSANGP00000023831 [Anopheles gambiae str. PEST] 30 9.6
>UniRef100_Q9C9N6 Hypothetical protein F4N21_22 [Arabidopsis thaliana]
Length = 607
Score = 36.6 bits (83), Expect = 0.13
Identities = 23/63 (36%), Positives = 36/63 (56%), Gaps = 9/63 (14%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+K L+ IEES+ + K++ DIE + + R + N Y ++MRELE +K+EL
Sbjct: 71 VKELTLRIEESNRRLKSRRIDIEAVMNES---------RIDGNGGYVRIMRELEDMKQEL 121
Query: 61 FKL 63
KL
Sbjct: 122 SKL 124
>UniRef100_Q8RZY7 Myosin heavy chain-like protein [Oryza sativa]
Length = 1051
Score = 33.9 bits (76), Expect = 0.87
Identities = 17/62 (27%), Positives = 33/62 (52%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKELF 61
K L IE+++ KA ++ +++ + + G + YA+V++EL+ KKEL
Sbjct: 506 KELERQIEQTTAKATSQRSELQAMWAARTRRKGTDAPGAERDARYAEVVQELDQAKKELL 565
Query: 62 KL 63
+L
Sbjct: 566 RL 567
>UniRef100_UPI00004999BB UPI00004999BB UniRef100 entry
Length = 420
Score = 33.1 bits (74), Expect = 1.5
Identities = 17/48 (35%), Positives = 25/48 (51%)
Query: 6 SMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMREL 53
S ++SS K K++D ETL+ GKG +G V E+ RE+
Sbjct: 102 SKSDQSSNALKPKLEDFETLKLIGKGTYGKKAVVETNEVEHTMAEREV 149
>UniRef100_UPI0000368EE6 UPI0000368EE6 UniRef100 entry
Length = 517
Score = 31.6 bits (70), Expect = 4.3
Identities = 23/59 (38%), Positives = 31/59 (51%), Gaps = 10/59 (16%)
Query: 6 SMIEESS------YKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
SMIEE++ YK A K IE E K K + R+ EY +VMRE+E+L +
Sbjct: 163 SMIEEAAGTRMYEYKKIAAQKTIEKKEAKLKE----IKTERSSYLEYQKVMREIEHLSR 217
>UniRef100_Q8IFG6 Hypothetical protein Tb927.1.760 [Trypanosoma brucei]
Length = 761
Score = 31.6 bits (70), Expect = 4.3
Identities = 17/62 (27%), Positives = 29/62 (46%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+++L ++ SY KA + KK + + R NE +EY Q + L L++E
Sbjct: 543 LRHLQRQHQQDSYDLKALRLQVGEERKKRQDAEARLAERENETHEYQQRLERLTVLQEEC 602
Query: 61 FK 62
K
Sbjct: 603 EK 604
>UniRef100_Q6L0D8 Iron-dependent repressor [Picrophilus torridus]
Length = 131
Score = 31.6 bits (70), Expect = 4.3
Identities = 16/49 (32%), Positives = 30/49 (60%), Gaps = 2/49 (4%)
Query: 4 LSSMIEESSYKAKAKMKDIETLEKKG--KGQHGVMVVRRNENYEYAQVM 50
LS + + K ++ +E LE+KG K +HG++++ N N EY+++M
Sbjct: 25 LSDLAVGLNIKPPTAIELLERLERKGLIKKEHGMIILTDNGNNEYSKIM 73
>UniRef100_UPI000042DC3D UPI000042DC3D UniRef100 entry
Length = 805
Score = 30.8 bits (68), Expect = 7.4
Identities = 20/58 (34%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKE 59
K SS EE + +A+ + KD+E+ EKK + V +NEN E + + ++E +KE
Sbjct: 141 KETSSKNEEKAEEAEGQSKDLESTEKKELNPKDNVNVNKNEN-EKSGLKDKVETNEKE 197
>UniRef100_Q6FV07 Similar to sp|P38991 Saccharomyces cerevisiae YPL209c IPL1 ser/thr
protein kinase [Candida glabrata]
Length = 358
Score = 30.8 bits (68), Expect = 7.4
Identities = 14/47 (29%), Positives = 26/47 (54%)
Query: 8 IEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELE 54
+E+S+Y K +KD E K GKG+ G + R++ + ++ +E
Sbjct: 86 LEKSTYNNKLSLKDFEVGRKLGKGKFGKVYCVRHKKSGFICALKAIE 132
>UniRef100_UPI00004328CB UPI00004328CB UniRef100 entry
Length = 208
Score = 30.4 bits (67), Expect = 9.6
Identities = 16/52 (30%), Positives = 33/52 (62%), Gaps = 3/52 (5%)
Query: 9 EESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
E+ S K + K+ DI+ L++K + + RN++ +Y+Q++ E++ L KE+
Sbjct: 58 EKHSTKLQKKLDDIKLLKEK---ERELQEECRNKDQQYSQLVTEVQKLSKEV 106
>UniRef100_UPI000042FCB7 UPI000042FCB7 UniRef100 entry
Length = 399
Score = 30.4 bits (67), Expect = 9.6
Identities = 16/42 (38%), Positives = 24/42 (57%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNEN 43
K SS EE + +A+ + KD+E+ EKK + V +NEN
Sbjct: 141 KETSSKNEEKAEEAEGQSKDLESTEKKELNPKDNVNVNKNEN 182
>UniRef100_UPI00002F403D UPI00002F403D UniRef100 entry
Length = 250
Score = 30.4 bits (67), Expect = 9.6
Identities = 16/46 (34%), Positives = 25/46 (53%), Gaps = 2/46 (4%)
Query: 7 MIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRE 52
M E+S AK + +E E K G HGV +V +N N ++ +M +
Sbjct: 106 MGEDSKSAAKGLLTQVE--EMKQDGSHGVKMVLKNANADFPYIMSD 149
>UniRef100_Q7PVJ7 ENSANGP00000023831 [Anopheles gambiae str. PEST]
Length = 397
Score = 30.4 bits (67), Expect = 9.6
Identities = 16/26 (61%), Positives = 21/26 (80%), Gaps = 1/26 (3%)
Query: 37 VVRR-NENYEYAQVMRELEYLKKELF 61
VVRR ++ YE AQ+MR+L LK+ELF
Sbjct: 97 VVRRYHKRYEIAQLMRDLFQLKRELF 122
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.129 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 134,065,919
Number of Sequences: 2790947
Number of extensions: 3338006
Number of successful extensions: 9781
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 9773
Number of HSP's gapped (non-prelim): 13
length of query: 122
length of database: 848,049,833
effective HSP length: 98
effective length of query: 24
effective length of database: 574,537,027
effective search space: 13788888648
effective search space used: 13788888648
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147875.9


