

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.6 - phase: 0 /pseudo
(974 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
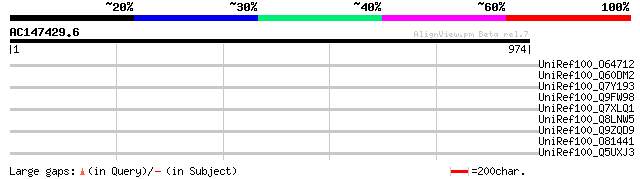
UniRef100_O64712 Putative reverse transcriptase [Arabidopsis tha... 44 0.024
UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa] 40 0.45
UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa] 40 0.45
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 39 0.59
UniRef100_Q7XLQ1 OSJNBa0044M19.5 protein [Oryza sativa] 38 1.3
UniRef100_Q8LNW5 Hypothetical protein OSJNBa0012L23.45 [Oryza sa... 38 1.7
UniRef100_Q9ZQD9 Putative non-LTR retroelement reverse transcrip... 38 1.7
UniRef100_O81441 T24H24.17 protein [Arabidopsis thaliana] 37 2.9
UniRef100_Q5UXJ3 Ribonuclease H-like protein [Haloarcula marismo... 35 8.5
>UniRef100_O64712 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 365
Score = 43.9 bits (102), Expect = 0.024
Identities = 25/73 (34%), Positives = 36/73 (49%), Gaps = 2/73 (2%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTA 956
+G R V FE+D++SL+ I G D S LG + +I L C F+ R+ N A
Sbjct: 274 HGLRYVWFESDSKSLVTLINNG--EDHSLLGTLIYDIRHWMLKLPYCSLEFVNRERNSAA 331
Query: 957 HSLAQLAHSEPNL 969
+LA H+ L
Sbjct: 332 DALASHVHARDPL 344
>UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa]
Length = 855
Score = 39.7 bits (91), Expect = 0.45
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 1/71 (1%)
Query: 893 LLKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDG 952
L K+ G + + +D ++I+RI+ DRS +G + +I +L F +C F + R
Sbjct: 761 LAKEEGLQHIVMASDCLTVIRRIQTS-GRDRSGVGCVIEDIKKLASTFVLCSFMHVNRLS 819
Query: 953 NHTAHSLAQLA 963
N AHSLA+ A
Sbjct: 820 NLAAHSLARNA 830
>UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa]
Length = 404
Score = 39.7 bits (91), Expect = 0.45
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 1/71 (1%)
Query: 893 LLKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDG 952
L K+ G + + +D ++I+RI+ DRS +G + +I +L F +C F + R
Sbjct: 310 LAKEEGLQHIVMASDCLTVIRRIQTS-GRDRSGVGCVIEDIKKLASTFVLCSFMHVNRLS 368
Query: 953 NHTAHSLAQLA 963
N AHSLA+ A
Sbjct: 369 NLAAHSLARNA 379
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 39.3 bits (90), Expect = 0.59
Identities = 24/71 (33%), Positives = 38/71 (52%), Gaps = 1/71 (1%)
Query: 893 LLKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDG 952
L K+ G + +D ++I+RI+ DRS +G + +I +L F +C F + R
Sbjct: 1288 LAKEEGLEHIVMASDCLTVIRRIQTS-GRDRSGVGCVIEDIKKLASTFVLCSFMHVNRLS 1346
Query: 953 NHTAHSLAQLA 963
N AHSLA+ A
Sbjct: 1347 NLAAHSLARNA 1357
>UniRef100_Q7XLQ1 OSJNBa0044M19.5 protein [Oryza sativa]
Length = 140
Score = 38.1 bits (87), Expect = 1.3
Identities = 22/66 (33%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLG-LFLSEIHQLQLNFDICHFSFIPRDGNHTA 956
G +VQFE D+ +L+Q +K + G L L + ++F++ F + PR+ N A
Sbjct: 43 GMMRVQFETDSLTLVQGLKSSNGYRLAATGGLCLDILQPCVISFNVFSFHYCPRNCNRVA 102
Query: 957 HSLAQL 962
H+LA L
Sbjct: 103 HALAAL 108
>UniRef100_Q8LNW5 Hypothetical protein OSJNBa0012L23.45 [Oryza sativa]
Length = 243
Score = 37.7 bits (86), Expect = 1.7
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 3/77 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G +Q E+D+ +LIQ ++ GL L + + +F C FS+ PR+ N A
Sbjct: 147 GMANIQLESDSLTLIQALREENFRYAPLGGLLLEAKNIITSSFVSCQFSYCPRECNKVAD 206
Query: 958 SLAQLAHSEP---NLVW 971
++A + + P +LVW
Sbjct: 207 AIASIGCNSPLNTDLVW 223
>UniRef100_Q9ZQD9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 280
Score = 37.7 bits (86), Expect = 1.7
Identities = 21/64 (32%), Positives = 34/64 (52%), Gaps = 2/64 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G +++ F +D+ +L++ I L L +IH L ++FD C F FI R N A
Sbjct: 208 GIKRISFASDSLTLVKAINRKFLTKE--LHRILHDIHSLSISFDACSFFFISRKHNSEAD 265
Query: 958 SLAQ 961
+LA+
Sbjct: 266 ALAK 269
>UniRef100_O81441 T24H24.17 protein [Arabidopsis thaliana]
Length = 1077
Score = 37.0 bits (84), Expect = 2.9
Identities = 20/64 (31%), Positives = 36/64 (56%), Gaps = 2/64 (3%)
Query: 900 RQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSL 959
+ + F +D++ +++ + + V L L +I L +FD C F+FI R+ NH A +L
Sbjct: 1009 KAISFASDSQIIVKALNQKLQVKE--LHGILHDILSLSSSFDACTFTFINREFNHQADAL 1066
Query: 960 AQLA 963
A+ A
Sbjct: 1067 AKRA 1070
>UniRef100_Q5UXJ3 Ribonuclease H-like protein [Haloarcula marismortui]
Length = 198
Score = 35.4 bits (80), Expect = 8.5
Identities = 18/66 (27%), Positives = 32/66 (48%)
Query: 895 KDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNH 954
+DYGF V D+E ++++++ D + L + +L +FD +PR+ N
Sbjct: 126 RDYGFDDVHIRGDSELIVKQVRGEWDTNDPELREHRVRVRELLTDFDDWQIEHVPREIND 185
Query: 955 TAHSLA 960
A LA
Sbjct: 186 RADELA 191
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.363 0.165 0.602
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,339,473,952
Number of Sequences: 2790947
Number of extensions: 47803232
Number of successful extensions: 162937
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 162933
Number of HSP's gapped (non-prelim): 10
length of query: 974
length of database: 848,049,833
effective HSP length: 137
effective length of query: 837
effective length of database: 465,690,094
effective search space: 389782608678
effective search space used: 389782608678
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 80 (35.4 bits)
Medicago: description of AC147429.6


