

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146571.7 + phase: 0 /pseudo
(2018 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
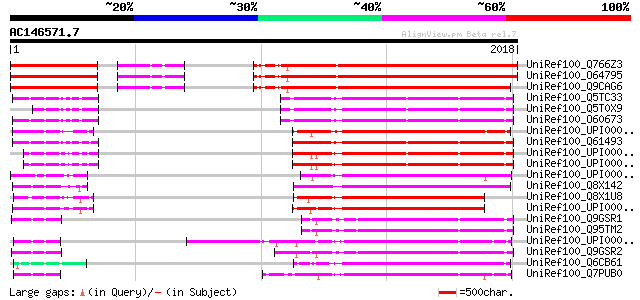
UniRef100_Q766Z3 Catalytic subunit of polymerase zeta [Arabidops... 1325 0.0
UniRef100_O64795 T1F15.3 protein [Arabidopsis thaliana] 1325 0.0
UniRef100_Q9CAG6 Putative DNA polymerase zeta catalytic subunit ... 1252 0.0
UniRef100_Q5TC33 REV3-like, catalytic subunit of DNA polymerase ... 692 0.0
UniRef100_Q5T0X9 REV3-like, catalytic subunit of DNA polymerase ... 692 0.0
UniRef100_O60673 DNA polymerase zeta catalytic subunit [Homo sap... 692 0.0
UniRef100_UPI000023CD58 UPI000023CD58 UniRef100 entry 691 0.0
UniRef100_Q61493 DNA polymerase zeta catalytic subunit [Mus musc... 691 0.0
UniRef100_UPI00003ACFBF UPI00003ACFBF UniRef100 entry 682 0.0
UniRef100_UPI00003ACFBE UPI00003ACFBE UniRef100 entry 682 0.0
UniRef100_UPI00003C1736 UPI00003C1736 UniRef100 entry 676 0.0
UniRef100_Q8X142 Polymerase zeta subunit [Emericella nidulans] 660 0.0
UniRef100_Q8X1U8 Catalytic subunit of DNA polymerase zeta [Neuro... 632 e-179
UniRef100_UPI000021B964 UPI000021B964 UniRef100 entry 625 e-177
UniRef100_Q9GSR1 DNA polymerase zeta catalytic subunit [Drosophi... 612 e-173
UniRef100_Q95TM2 LD40801p [Drosophila melanogaster] 612 e-173
UniRef100_UPI0000243912 UPI0000243912 UniRef100 entry 612 e-173
UniRef100_Q9GSR2 DNA polymerase zeta catalytic subunit [Drosophi... 612 e-173
UniRef100_Q6CB61 Similar to sp|P14284 Saccharomyces cerevisiae D... 608 e-172
UniRef100_Q7PUB0 ENSANGP00000009889 [Anopheles gambiae str. PEST] 602 e-170
>UniRef100_Q766Z3 Catalytic subunit of polymerase zeta [Arabidopsis thaliana]
Length = 1890
Score = 1325 bits (3428), Expect = 0.0
Identities = 682/1064 (64%), Positives = 824/1064 (77%), Gaps = 42/1064 (3%)
Query: 970 EDDEMPGNALDV-FLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLL 1028
+ DE GN+ D+ F P+ +++++KK + + ++ G+ ++ NDGS+LYLL
Sbjct: 853 QHDEYEGNSNDIPFFPLE--DNKEEKKHFFQGTSL---------GIPLHHLNDGSNLYLL 901
Query: 1029 TPNILPPSAGSVQRWLFCDERDQEPDAEDQDVPKCTSGPLRHTPDQMRQEPGAKDKDISK 1088
TP PPS SV +W+ D+ D D+E Q PLR + GA D++
Sbjct: 902 TPAFSPPSVDSVLQWISNDKGDSNIDSEKQ--------PLRDN----HNDRGASFTDLAS 949
Query: 1089 CASGPTLRPELHQD------------TEKKLPCINEGQTERIKAHMDHSQDISQISGPGE 1136
++ ++ + Q TE ++ +G + SQ++SQISGP
Sbjct: 950 ASNVVSVSEHVEQHNNLFVNSESNAYTESEIDLKPKGTFLNLNLQASVSQELSQISGPDG 1009
Query: 1137 KSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
KS TPLSQ+GF+DPAS G GQQL +LSIEV AESRGDL PDP+FD+VN++AL Q D
Sbjct: 1010 KSGPTPLSQMGFRDPASMGAGQQLTILSIEVHAESRGDLRPDPRFDSVNVIALVVQNDDS 1069
Query: 1197 ATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQG 1256
EV VLL S QR++DGLS CK+ F +E+ L + F + + DPD+L+GW+IQG
Sbjct: 1070 FVAEVFVLLFSPDSIDQRNVDGLSGCKLSVFLEERQLFRYFIETLCKWDPDVLLGWDIQG 1129
Query: 1257 SSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARES 1316
S+GFLAERA+ LG LN++SRTPS + N+ D K + L +PD + E
Sbjct: 1130 GSIGFLAERAAQLGIRFLNNISRTPSPTTTNNSDNKRKLGNNL---LPDPLVANPAQVEE 1186
Query: 1317 SIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVL 1376
+IEDEWGRTHASGVHVGGRIVLN WRLIRGEVKLN+Y++EAV+EAVLR+KVPS+ + VL
Sbjct: 1187 VVIEDEWGRTHASGVHVGGRIVLNAWRLIRGEVKLNMYTIEAVSEAVLRQKVPSIPYKVL 1246
Query: 1377 TKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQY 1436
T+WFS GP ARY+CI+Y++ RA LNLEI++QLDM+NRTSELARVFGIDFFSVLSRGSQY
Sbjct: 1247 TEWFSSGPAGARYRCIEYVIRRANLNLEIMSQLDMINRTSELARVFGIDFFSVLSRGSQY 1306
Query: 1437 RVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPS 1496
RVESM LRLAHTQNY+AISPGNQQVASQPAMEC+PLVMEPES FY DPV+VLDFQSLYPS
Sbjct: 1307 RVESMLLRLAHTQNYLAISPGNQQVASQPAMECVPLVMEPESAFYDDPVIVLDFQSLYPS 1366
Query: 1497 MIIAYNLCFCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRG 1556
MIIAYNLCF TCLGK+ K NTLGVS + + +LQDL +QIL TPN VM+VP ++RRG
Sbjct: 1367 MIIAYNLCFSTCLGKLAHLKMNTLGVSSYSLDLDVLQDL-NQILQTPNSVMYVPPEVRRG 1425
Query: 1557 VLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRM 1616
+LPRLLEEILSTRIMVK+AMKKL+PSE VL RIFNARQLALKLI+NVTYGYTAAGFSGRM
Sbjct: 1426 ILPRLLEEILSTRIMVKKAMKKLTPSEAVLHRIFNARQLALKLIANVTYGYTAAGFSGRM 1485
Query: 1617 PCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSE 1676
PCAELADSIVQCGR TLEKAISFVN ++ WNA+V+YGDTDSMFVLLKGRT KEAF +G E
Sbjct: 1486 PCAELADSIVQCGRSTLEKAISFVNANDNWNARVVYGDTDSMFVLLKGRTVKEAFVVGQE 1545
Query: 1677 IASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTC 1736
IASAIT MNP PVTLKMEKVYHPCFL+TKKRYVGYSYE+PNQ EP+FDAKGIETVRRDTC
Sbjct: 1546 IASAITEMNPHPVTLKMEKVYHPCFLLTKKRYVGYSYESPNQREPIFDAKGIETVRRDTC 1605
Query: 1737 EAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARIS 1796
EAVAK MEQSLRLFFE +++ +VK+YL RQWKRILSGRVSL+DFIFAKEVRLGTYS R S
Sbjct: 1606 EAVAKTMEQSLRLFFEQKNISKVKSYLYRQWKRILSGRVSLQDFIFAKEVRLGTYSTRDS 1665
Query: 1797 S-LPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRI 1855
S LPPAAIVATK+M+ D R EPRYAER+PYVVIHGEPGARLVDMVVDPL +L +D+P+R+
Sbjct: 1666 SLLPPAAIVATKSMKADPRTEPRYAERVPYVVIHGEPGARLVDMVVDPLVLLDVDTPYRL 1725
Query: 1856 NDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKHAF-TPNFQRTRIDYYYL 1914
NDLYYINKQIIPALQRVFGLVGADLNQWF EMPR R + + + N +TRIDY+YL
Sbjct: 1726 NDLYYINKQIIPALQRVFGLVGADLNQWFLEMPRLTRSSLGQRPLNSKNSHKTRIDYFYL 1785
Query: 1915 SKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLL 1974
SKHC+LCG +VQ SA+LC++C +N+ AAA ++ KT+KLE+EMQHL +IC+HCGGGD ++
Sbjct: 1786 SKHCILCGEVVQESAQLCNRCLQNKSAAAATIVWKTSKLEREMQHLATICRHCGGGDWVV 1845
Query: 1975 ESGVKCTSISCLVFYERRKVQKELLAATHVAADKGFYPRCTVEW 2018
+SGVKC S++C VFYERRKVQKEL + +A + YP+C EW
Sbjct: 1846 QSGVKCNSLACSVFYERRKVQKELRGLSSIATESELYPKCMAEW 1889
Score = 405 bits (1041), Expect = e-111
Identities = 206/362 (56%), Positives = 263/362 (71%), Gaps = 14/362 (3%)
Query: 1 MSDSESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTC 60
M+DS+S S+VFS+RIVS+D+YMASPIP +I +S+F +VNEVPVIR+YGSTPAGQKTC
Sbjct: 1 MADSQSGSNVFSLRIVSIDYYMASPIPGYNICYSSFQGSEVNEVPVIRIYGSTPAGQKTC 60
Query: 61 LHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCSLVR 120
LHIH ALPYLY+ CS+IPL+ + D T ++ LEKALKLKG+A S RQH+H C +VR
Sbjct: 61 LHIHRALPYLYIPCSEIPLEHHKGVDGSTLALSLELEKALKLKGNAASKRQHIHDCEIVR 120
Query: 121 AKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDY 180
AKKFYGY S EE FVKIYL Y+P DV+RAA+LLLAGAVL KSLQP+ESHIPFILQFLVDY
Sbjct: 121 AKKFYGYHSTEEAFVKIYLSYHPPDVARAASLLLAGAVLGKSLQPYESHIPFILQFLVDY 180
Query: 181 NLYGMGHLHLSKMRFRHPMPDSIHKK----SDINSQH------RKADPGAEACLESKIWM 230
NLYGMGH+H+SKM+FR P+P + D Q KA+ A A + +W
Sbjct: 181 NLYGMGHVHISKMKFRSPVPHHFRPRRFDLDDCPGQRIDEVAITKANSSAAASVSFPVWS 240
Query: 231 SSTISFDWMWSLPSES----GGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLS 286
STI WMW+L ES S + + H +RQS+CELEGD + +ILNQQFKM++SLS
Sbjct: 241 LSTIPGQWMWNLSEESDTPLSQSQHRHQHHYRRQSLCELEGDATSSDILNQQFKMYNSLS 300
Query: 287 QTSSNVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKLIQLC 346
Q S+ NMVQSLV+IWEE+ +R+G+H+A + D GKP DV++ +S + F L ++
Sbjct: 301 QAQSDTNMVQSLVAIWEEEYERTGVHDAPIPPDPGKPSAADVLQTMSDYVGFGNMLKEML 360
Query: 347 SE 348
++
Sbjct: 361 NK 362
Score = 120 bits (302), Expect = 3e-25
Identities = 92/269 (34%), Positives = 137/269 (50%), Gaps = 14/269 (5%)
Query: 430 QLKRLKSQNLLKWLATSQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQ 489
Q++ ++ L KW A+SQAAEDINSDDE+ ETILSPL P A+I+K+LE A+ Y S+SQ
Sbjct: 439 QIQDAEALGLFKWFASSQAAEDINSDDEILRETILSPLLPLASINKVLEMASTDYVSQSQ 498
Query: 490 QECQDILDSIDDMLALDLPKKKPSRSFDHNCPIEASSDNMIPQVDGSNDDEFSSPCASLA 549
+ECQDILDS +++ K+ S + + SSD +++ ++D SS +
Sbjct: 499 KECQDILDSQENLPDFGSSTKRALPSNPDSQNLRTSSDKQSLEIEVASDVPDSSTSNGAS 558
Query: 550 EISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQV 609
E S S+ SE N S N N WG LPF++T + D + +
Sbjct: 559 ENSFRRYRKSDL-HTSEVMEYKNRSFSKSNKPS-NSVWGPLPFTLTKNLQKDFDSTNA-- 614
Query: 610 AHLFERETEDSSLSDYLTRNEIKNNKYIKRNVGEGASD---SKEVHSLVNCSLRDLMRRK 666
+ + +S Y NE+ +N + V E +D + + + L CSLRDLMR+K
Sbjct: 615 ----SDKLGLTKISSY-PMNEMTDNYIVP--VKEHQADVCNTIDRNVLAGCSLRDLMRKK 667
Query: 667 RSYRVEHDERESGTAKKLNLDRHGGTKTC 695
R E + ++K+ RHG C
Sbjct: 668 RLCHGESPVSQHMKSRKVRDSRHGEKNEC 696
>UniRef100_O64795 T1F15.3 protein [Arabidopsis thaliana]
Length = 1894
Score = 1325 bits (3428), Expect = 0.0
Identities = 682/1064 (64%), Positives = 824/1064 (77%), Gaps = 42/1064 (3%)
Query: 970 EDDEMPGNALDV-FLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLL 1028
+ DE GN+ D+ F P+ +++++KK + + ++ G+ ++ NDGS+LYLL
Sbjct: 857 QHDEYEGNSNDIPFFPLE--DNKEEKKHFFQGTSL---------GIPLHHLNDGSNLYLL 905
Query: 1029 TPNILPPSAGSVQRWLFCDERDQEPDAEDQDVPKCTSGPLRHTPDQMRQEPGAKDKDISK 1088
TP PPS SV +W+ D+ D D+E Q PLR + GA D++
Sbjct: 906 TPAFSPPSVDSVLQWISNDKGDSNIDSEKQ--------PLRDN----HNDRGASFTDLAS 953
Query: 1089 CASGPTLRPELHQD------------TEKKLPCINEGQTERIKAHMDHSQDISQISGPGE 1136
++ ++ + Q TE ++ +G + SQ++SQISGP
Sbjct: 954 ASNVVSVSEHVEQHNNLFVNSESNAYTESEIDLKPKGTFLNLNLQASVSQELSQISGPDG 1013
Query: 1137 KSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
KS TPLSQ+GF+DPAS G GQQL +LSIEV AESRGDL PDP+FD+VN++AL Q D
Sbjct: 1014 KSGPTPLSQMGFRDPASMGAGQQLTILSIEVHAESRGDLRPDPRFDSVNVIALVVQNDDS 1073
Query: 1197 ATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQG 1256
EV VLL S QR++DGLS CK+ F +E+ L + F + + DPD+L+GW+IQG
Sbjct: 1074 FVAEVFVLLFSPDSIDQRNVDGLSGCKLSVFLEERQLFRYFIETLCKWDPDVLLGWDIQG 1133
Query: 1257 SSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARES 1316
S+GFLAERA+ LG LN++SRTPS + N+ D K + L +PD + E
Sbjct: 1134 GSIGFLAERAAQLGIRFLNNISRTPSPTTTNNSDNKRKLGNNL---LPDPLVANPAQVEE 1190
Query: 1317 SIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVL 1376
+IEDEWGRTHASGVHVGGRIVLN WRLIRGEVKLN+Y++EAV+EAVLR+KVPS+ + VL
Sbjct: 1191 VVIEDEWGRTHASGVHVGGRIVLNAWRLIRGEVKLNMYTIEAVSEAVLRQKVPSIPYKVL 1250
Query: 1377 TKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQY 1436
T+WFS GP ARY+CI+Y++ RA LNLEI++QLDM+NRTSELARVFGIDFFSVLSRGSQY
Sbjct: 1251 TEWFSSGPAGARYRCIEYVIRRANLNLEIMSQLDMINRTSELARVFGIDFFSVLSRGSQY 1310
Query: 1437 RVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPS 1496
RVESM LRLAHTQNY+AISPGNQQVASQPAMEC+PLVMEPES FY DPV+VLDFQSLYPS
Sbjct: 1311 RVESMLLRLAHTQNYLAISPGNQQVASQPAMECVPLVMEPESAFYDDPVIVLDFQSLYPS 1370
Query: 1497 MIIAYNLCFCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRG 1556
MIIAYNLCF TCLGK+ K NTLGVS + + +LQDL +QIL TPN VM+VP ++RRG
Sbjct: 1371 MIIAYNLCFSTCLGKLAHLKMNTLGVSSYSLDLDVLQDL-NQILQTPNSVMYVPPEVRRG 1429
Query: 1557 VLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRM 1616
+LPRLLEEILSTRIMVK+AMKKL+PSE VL RIFNARQLALKLI+NVTYGYTAAGFSGRM
Sbjct: 1430 ILPRLLEEILSTRIMVKKAMKKLTPSEAVLHRIFNARQLALKLIANVTYGYTAAGFSGRM 1489
Query: 1617 PCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSE 1676
PCAELADSIVQCGR TLEKAISFVN ++ WNA+V+YGDTDSMFVLLKGRT KEAF +G E
Sbjct: 1490 PCAELADSIVQCGRSTLEKAISFVNANDNWNARVVYGDTDSMFVLLKGRTVKEAFVVGQE 1549
Query: 1677 IASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTC 1736
IASAIT MNP PVTLKMEKVYHPCFL+TKKRYVGYSYE+PNQ EP+FDAKGIETVRRDTC
Sbjct: 1550 IASAITEMNPHPVTLKMEKVYHPCFLLTKKRYVGYSYESPNQREPIFDAKGIETVRRDTC 1609
Query: 1737 EAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARIS 1796
EAVAK MEQSLRLFFE +++ +VK+YL RQWKRILSGRVSL+DFIFAKEVRLGTYS R S
Sbjct: 1610 EAVAKTMEQSLRLFFEQKNISKVKSYLYRQWKRILSGRVSLQDFIFAKEVRLGTYSTRDS 1669
Query: 1797 S-LPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRI 1855
S LPPAAIVATK+M+ D R EPRYAER+PYVVIHGEPGARLVDMVVDPL +L +D+P+R+
Sbjct: 1670 SLLPPAAIVATKSMKADPRTEPRYAERVPYVVIHGEPGARLVDMVVDPLVLLDVDTPYRL 1729
Query: 1856 NDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKHAF-TPNFQRTRIDYYYL 1914
NDLYYINKQIIPALQRVFGLVGADLNQWF EMPR R + + + N +TRIDY+YL
Sbjct: 1730 NDLYYINKQIIPALQRVFGLVGADLNQWFLEMPRLTRSSLGQRPLNSKNSHKTRIDYFYL 1789
Query: 1915 SKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLL 1974
SKHC+LCG +VQ SA+LC++C +N+ AAA ++ KT+KLE+EMQHL +IC+HCGGGD ++
Sbjct: 1790 SKHCILCGEVVQESAQLCNRCLQNKSAAAATIVWKTSKLEREMQHLATICRHCGGGDWVV 1849
Query: 1975 ESGVKCTSISCLVFYERRKVQKELLAATHVAADKGFYPRCTVEW 2018
+SGVKC S++C VFYERRKVQKEL + +A + YP+C EW
Sbjct: 1850 QSGVKCNSLACSVFYERRKVQKELRGLSSIATESELYPKCMAEW 1893
Score = 399 bits (1026), Expect = e-109
Identities = 206/366 (56%), Positives = 263/366 (71%), Gaps = 18/366 (4%)
Query: 1 MSDSESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTC 60
M+DS+S S+VFS+RIVS+D+YMASPIP +I +S+F +VNEVPVIR+YGSTPAGQKTC
Sbjct: 1 MADSQSGSNVFSLRIVSIDYYMASPIPGYNICYSSFQGSEVNEVPVIRIYGSTPAGQKTC 60
Query: 61 LHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCSLVR 120
LHIH ALPYLY+ CS+IPL+ + D T ++ LEKALKLKG+A S RQH+H C +VR
Sbjct: 61 LHIHRALPYLYIPCSEIPLEHHKGVDGSTLALSLELEKALKLKGNAASKRQHIHDCEIVR 120
Query: 121 AKKFYGYCSFEEFFVKIYL----RYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQF 176
AKKFYGY S EE FVKIYL Y+P DV+RAA+LLLAGAVL KSLQP+ESHIPFILQF
Sbjct: 121 AKKFYGYHSTEEAFVKIYLYPYSSYHPPDVARAASLLLAGAVLGKSLQPYESHIPFILQF 180
Query: 177 LVDYNLYGMGHLHLSKMRFRHPMPDSIHKK----SDINSQH------RKADPGAEACLES 226
LVDYNLYGMGH+H+SKM+FR P+P + D Q KA+ A A +
Sbjct: 181 LVDYNLYGMGHVHISKMKFRSPVPHHFRPRRFDLDDCPGQRIDEVAITKANSSAAASVSF 240
Query: 227 KIWMSSTISFDWMWSLPSES----GGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMF 282
+W STI WMW+L ES S + + H +RQS+CELEGD + +ILNQQFKM+
Sbjct: 241 PVWSLSTIPGQWMWNLSEESDTPLSQSQHRHQHHYRRQSLCELEGDATSSDILNQQFKMY 300
Query: 283 SSLSQTSSNVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKL 342
+SLSQ S+ NMVQSLV+IWEE+ +R+G+H+A + D GKP DV++ +S + F L
Sbjct: 301 NSLSQAQSDTNMVQSLVAIWEEEYERTGVHDAPIPPDPGKPSAADVLQTMSDYVGFGNML 360
Query: 343 IQLCSE 348
++ ++
Sbjct: 361 KEMLNK 366
Score = 120 bits (302), Expect = 3e-25
Identities = 92/269 (34%), Positives = 137/269 (50%), Gaps = 14/269 (5%)
Query: 430 QLKRLKSQNLLKWLATSQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQ 489
Q++ ++ L KW A+SQAAEDINSDDE+ ETILSPL P A+I+K+LE A+ Y S+SQ
Sbjct: 443 QIQDAEALGLFKWFASSQAAEDINSDDEILRETILSPLLPLASINKVLEMASTDYVSQSQ 502
Query: 490 QECQDILDSIDDMLALDLPKKKPSRSFDHNCPIEASSDNMIPQVDGSNDDEFSSPCASLA 549
+ECQDILDS +++ K+ S + + SSD +++ ++D SS +
Sbjct: 503 KECQDILDSQENLPDFGSSTKRALPSNPDSQNLRTSSDKQSLEIEVASDVPDSSTSNGAS 562
Query: 550 EISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQV 609
E S S+ SE N S N N WG LPF++T + D + +
Sbjct: 563 ENSFRRYRKSDL-HTSEVMEYKNRSFSKSNKPS-NSVWGPLPFTLTKNLQKDFDSTNA-- 618
Query: 610 AHLFERETEDSSLSDYLTRNEIKNNKYIKRNVGEGASD---SKEVHSLVNCSLRDLMRRK 666
+ + +S Y NE+ +N + V E +D + + + L CSLRDLMR+K
Sbjct: 619 ----SDKLGLTKISSY-PMNEMTDNYIVP--VKEHQADVCNTIDRNVLAGCSLRDLMRKK 671
Query: 667 RSYRVEHDERESGTAKKLNLDRHGGTKTC 695
R E + ++K+ RHG C
Sbjct: 672 RLCHGESPVSQHMKSRKVRDSRHGEKNEC 700
>UniRef100_Q9CAG6 Putative DNA polymerase zeta catalytic subunit [Arabidopsis thaliana]
Length = 1871
Score = 1252 bits (3239), Expect = 0.0
Identities = 656/1038 (63%), Positives = 793/1038 (76%), Gaps = 42/1038 (4%)
Query: 970 EDDEMPGNALDV-FLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLL 1028
+ DE GN+ D+ F P+ +++++KK + + ++ G+ ++ NDGS+LYLL
Sbjct: 857 QHDEYEGNSNDIPFFPLE--DNKEEKKHFFQGTSL---------GIPLHHLNDGSNLYLL 905
Query: 1029 TPNILPPSAGSVQRWLFCDERDQEPDAEDQDVPKCTSGPLRHTPDQMRQEPGAKDKDISK 1088
TP PPS SV +W+ D+ D D+E Q PLR + GA D++
Sbjct: 906 TPAFSPPSVDSVLQWISNDKGDSNIDSEKQ--------PLRDN----HNDRGASFTDLAS 953
Query: 1089 CASGPTLRPELHQD------------TEKKLPCINEGQTERIKAHMDHSQDISQISGPGE 1136
++ ++ + Q TE ++ +G + SQ++SQISGP
Sbjct: 954 ASNVVSVSEHVEQHNNLFVNSESNAYTESEIDLKPKGTFLNLNLQASVSQELSQISGPDG 1013
Query: 1137 KSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
KS TPLSQ+GF+DPAS G GQQL +LSIEV AESRGDL PDP+FD+VN++AL Q D
Sbjct: 1014 KSGPTPLSQMGFRDPASMGAGQQLTILSIEVHAESRGDLRPDPRFDSVNVIALVVQNDDS 1073
Query: 1197 ATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQG 1256
EV VLL S QR++DGLS CK+ F +E+ L + F + + DPD+L+GW+IQG
Sbjct: 1074 FVAEVFVLLFSPDSIDQRNVDGLSGCKLSVFLEERQLFRYFIETLCKWDPDVLLGWDIQG 1133
Query: 1257 SSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARES 1316
S+GFLAERA+ LG LN++SRTPS + N+ D K + L +PD + E
Sbjct: 1134 GSIGFLAERAAQLGIRFLNNISRTPSPTTTNNSDNKRKLGNNL---LPDPLVANPAQVEE 1190
Query: 1317 SIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVL 1376
+IEDEWGRTHASGVHVGGRIVLN WRLIRGEVKLN+Y++EAV+EAVLR+KVPS+ + VL
Sbjct: 1191 VVIEDEWGRTHASGVHVGGRIVLNAWRLIRGEVKLNMYTIEAVSEAVLRQKVPSIPYKVL 1250
Query: 1377 TKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQY 1436
T+WFS GP ARY+CI+Y++ RA LNLEI++QLDM+NRTSELARVFGIDFFSVLSRGSQY
Sbjct: 1251 TEWFSSGPAGARYRCIEYVIRRANLNLEIMSQLDMINRTSELARVFGIDFFSVLSRGSQY 1310
Query: 1437 RVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPS 1496
RVESM LRLAHTQNY+AISPGNQQVASQPAMEC+PLVMEPES FY DPV+VLDFQSLYPS
Sbjct: 1311 RVESMLLRLAHTQNYLAISPGNQQVASQPAMECVPLVMEPESAFYDDPVIVLDFQSLYPS 1370
Query: 1497 MIIAYNLCFCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRG 1556
MIIAYNLCF TCLGK+ K NTLGVS + + +LQDL +QIL TPN VM+VP ++RRG
Sbjct: 1371 MIIAYNLCFSTCLGKLAHLKMNTLGVSSYSLDLDVLQDL-NQILQTPNSVMYVPPEVRRG 1429
Query: 1557 VLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRM 1616
+LPRLLEEILSTRIMVK+AMKKL+PSE VL RIFNARQLALKLI+NVTYGYTAAGFSGRM
Sbjct: 1430 ILPRLLEEILSTRIMVKKAMKKLTPSEAVLHRIFNARQLALKLIANVTYGYTAAGFSGRM 1489
Query: 1617 PCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSE 1676
PCAELADSIVQCGR TLEKAISFVN ++ WNA+V+YGDTDSMFVLLKGRT KEAF +G E
Sbjct: 1490 PCAELADSIVQCGRSTLEKAISFVNANDNWNARVVYGDTDSMFVLLKGRTVKEAFVVGQE 1549
Query: 1677 IASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTC 1736
IASAIT MNP PVTLKMEKVYHPCFL+TKKRYVGYSYE+PNQ EP+FDAKGIETVRRDTC
Sbjct: 1550 IASAITEMNPHPVTLKMEKVYHPCFLLTKKRYVGYSYESPNQREPIFDAKGIETVRRDTC 1609
Query: 1737 EAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARIS 1796
EAVAK MEQSLRLFFE +++ +VK+YL RQWKRILSGRVSL+DFIFAKEVRLGTYS R S
Sbjct: 1610 EAVAKTMEQSLRLFFEQKNISKVKSYLYRQWKRILSGRVSLQDFIFAKEVRLGTYSTRDS 1669
Query: 1797 S-LPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRI 1855
S LPPAAIVATK+M+ D R EPRYAER+PYVVIHGEPGARLVDMVVDPL +L +D+P+R+
Sbjct: 1670 SLLPPAAIVATKSMKADPRTEPRYAERVPYVVIHGEPGARLVDMVVDPLVLLDVDTPYRL 1729
Query: 1856 NDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKHAF-TPNFQRTRIDYYYL 1914
NDLYYINKQIIPALQRVFGLVGADLNQWF EMPR R + + + N +TRIDY+YL
Sbjct: 1730 NDLYYINKQIIPALQRVFGLVGADLNQWFLEMPRLTRSSLGQRPLNSKNSHKTRIDYFYL 1789
Query: 1915 SKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLL 1974
SKHC+LCG +VQ SA+LC++C +N+ AAA ++ KT+KLE+EMQHL ++ LL
Sbjct: 1790 SKHCILCGEVVQESAQLCNRCLQNKSAAAATIVWKTSKLEREMQHLATVSTPKSNVYELL 1849
Query: 1975 ESGVKCTSISCLVFYERR 1992
++ V T+I L +R
Sbjct: 1850 QNLVTFTTIILLSLLSKR 1867
Score = 399 bits (1026), Expect = e-109
Identities = 206/366 (56%), Positives = 263/366 (71%), Gaps = 18/366 (4%)
Query: 1 MSDSESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTC 60
M+DS+S S+VFS+RIVS+D+YMASPIP +I +S+F +VNEVPVIR+YGSTPAGQKTC
Sbjct: 1 MADSQSGSNVFSLRIVSIDYYMASPIPGYNICYSSFQGSEVNEVPVIRIYGSTPAGQKTC 60
Query: 61 LHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCSLVR 120
LHIH ALPYLY+ CS+IPL+ + D T ++ LEKALKLKG+A S RQH+H C +VR
Sbjct: 61 LHIHRALPYLYIPCSEIPLEHHKGVDGSTLALSLELEKALKLKGNAASKRQHIHDCEIVR 120
Query: 121 AKKFYGYCSFEEFFVKIYL----RYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQF 176
AKKFYGY S EE FVKIYL Y+P DV+RAA+LLLAGAVL KSLQP+ESHIPFILQF
Sbjct: 121 AKKFYGYHSTEEAFVKIYLYPYSSYHPPDVARAASLLLAGAVLGKSLQPYESHIPFILQF 180
Query: 177 LVDYNLYGMGHLHLSKMRFRHPMPDSIHKK----SDINSQH------RKADPGAEACLES 226
LVDYNLYGMGH+H+SKM+FR P+P + D Q KA+ A A +
Sbjct: 181 LVDYNLYGMGHVHISKMKFRSPVPHHFRPRRFDLDDCPGQRIDEVAITKANSSAAASVSF 240
Query: 227 KIWMSSTISFDWMWSLPSES----GGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMF 282
+W STI WMW+L ES S + + H +RQS+CELEGD + +ILNQQFKM+
Sbjct: 241 PVWSLSTIPGQWMWNLSEESDTPLSQSQHRHQHHYRRQSLCELEGDATSSDILNQQFKMY 300
Query: 283 SSLSQTSSNVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKL 342
+SLSQ S+ NMVQSLV+IWEE+ +R+G+H+A + D GKP DV++ +S + F L
Sbjct: 301 NSLSQAQSDTNMVQSLVAIWEEEYERTGVHDAPIPPDPGKPSAADVLQTMSDYVGFGNML 360
Query: 343 IQLCSE 348
++ ++
Sbjct: 361 KEMLNK 366
Score = 120 bits (302), Expect = 3e-25
Identities = 92/269 (34%), Positives = 137/269 (50%), Gaps = 14/269 (5%)
Query: 430 QLKRLKSQNLLKWLATSQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQ 489
Q++ ++ L KW A+SQAAEDINSDDE+ ETILSPL P A+I+K+LE A+ Y S+SQ
Sbjct: 443 QIQDAEALGLFKWFASSQAAEDINSDDEILRETILSPLLPLASINKVLEMASTDYVSQSQ 502
Query: 490 QECQDILDSIDDMLALDLPKKKPSRSFDHNCPIEASSDNMIPQVDGSNDDEFSSPCASLA 549
+ECQDILDS +++ K+ S + + SSD +++ ++D SS +
Sbjct: 503 KECQDILDSQENLPDFGSSTKRALPSNPDSQNLRTSSDKQSLEIEVASDVPDSSTSNGAS 562
Query: 550 EISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQV 609
E S S+ SE N S N N WG LPF++T + D + +
Sbjct: 563 ENSFRRYRKSDL-HTSEVMEYKNRSFSKSNKPS-NSVWGPLPFTLTKNLQKDFDSTNA-- 618
Query: 610 AHLFERETEDSSLSDYLTRNEIKNNKYIKRNVGEGASD---SKEVHSLVNCSLRDLMRRK 666
+ + +S Y NE+ +N + V E +D + + + L CSLRDLMR+K
Sbjct: 619 ----SDKLGLTKISSY-PMNEMTDNYIVP--VKEHQADVCNTIDRNVLAGCSLRDLMRKK 671
Query: 667 RSYRVEHDERESGTAKKLNLDRHGGTKTC 695
R E + ++K+ RHG C
Sbjct: 672 RLCHGESPVSQHMKSRKVRDSRHGEKNEC 700
>UniRef100_Q5TC33 REV3-like, catalytic subunit of DNA polymerase zeta [Homo sapiens]
Length = 3130
Score = 692 bits (1785), Expect = 0.0
Identities = 398/946 (42%), Positives = 558/946 (58%), Gaps = 66/946 (6%)
Query: 1078 EPGAKDKDISKCASGPTLRPELHQDTEKKLPCINEGQTERIKAHMDHSQDISQISGPGEK 1137
EP + K ++ TLR L + + +N Q E SQI GP
Sbjct: 2225 EPKTQKLSNKKGSNTDTLRRVLLTQAKNQFAAVNTPQKET-----------SQIDGPSLN 2273
Query: 1138 SSFTPLSQIGFQDPASAGHG-QQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
+++ I A A H Q L L+S+E+ A +R DL PDP+FD + + D
Sbjct: 2274 NTYGFKVSIQNLQEAKALHEIQNLTLISVELHARTRRDLEPDPEFDPICALFYCISSDTP 2333
Query: 1197 AT-------VEVLVL-----LHSKFVPCQRSL---DGLSDCKVLNFTDEKHLLKEFTKIV 1241
V+V+ + S+ + Q L G++ +V DEK L E I+
Sbjct: 2334 LPDTEKTELTGVIVIDKDKTVFSQDIRYQTPLLIRSGITGLEVTYAADEKALFHEIANII 2393
Query: 1242 SSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEP 1301
DPDIL+G+EIQ S G+L +RA+ L L +SR P D K + E
Sbjct: 2394 KRYDPDILLGYEIQMHSWGYLLQRAAALSIDLCRMISRVP--------DDKIENRFAAE- 2444
Query: 1302 DIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAE 1361
DE+G S +++ GRI LNLWR++R EV L Y+ E V+
Sbjct: 2445 ------------------RDEYGSYTMSEINIVGRITLNLWRIMRNEVALTNYTFENVSF 2486
Query: 1362 AVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARV 1421
VL ++ P VL+ WF R++ + + V R + NL++L QLD++ +TSE+AR+
Sbjct: 2487 HVLHQRFPLFTFRVLSDWFDNKTDLYRWKMVDHYVSRVRGNLQMLEQLDLIGKTSEMARL 2546
Query: 1422 FGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFY 1481
FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ + A +C+PL+MEPES FY
Sbjct: 2547 FGIQFLHVLTRGSQYRVESMMLRIAKPMNYIPVTPSVQQRSQMRAPQCVPLIMEPESRFY 2606
Query: 1482 SDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSPFVPEQHILQDLKDQ 1538
S+ V+VLDFQSLYPS++IAYN CF TCLG V G + +L ++
Sbjct: 2607 SNSVLVLDFQSLYPSIVIAYNYCFSTCLGHVENLGKYDEFKFGCTSLRVPPDLLYQVRHD 2666
Query: 1539 ILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALK 1598
I ++PNGV FV +R+GVLPR+LEEIL TR MVKQ+MK ++ L R+ +ARQL LK
Sbjct: 2667 ITVSPNGVAFVKPSVRKGVLPRMLEEILKTRFMVKQSMKAYK-QDRALSRMLDARQLGLK 2725
Query: 1599 LISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSM 1658
LI+NVT+GYT+A FSGRMPC E+ DSIV R TLE+AI VN +KW A+V+YGDTDSM
Sbjct: 2726 LIANVTFGYTSANFSGRMPCIEVGDSIVHKARETLERAIKLVNDTKKWGARVVYGDTDSM 2785
Query: 1659 FVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQ 1718
FVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L TKKRYVGY YE +Q
Sbjct: 2786 FVLLKGATKEQSFKIGQEIAEAVTATNPKPVKLKFEKVYLPCVLQTKKRYVGYMYETLDQ 2845
Query: 1719 IEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLK 1778
+PVFDAKGIETVRRD+C AV+KI+E+SL+L FE + + +K Y+QRQ ++L G+ S++
Sbjct: 2846 KDPVFDAKGIETVRRDSCPAVSKILERSLKLLFETRDISLIKQYVQRQCMKLLEGKASIQ 2905
Query: 1779 DFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVD 1838
DFIFAKE R G++S + + PA + K + DRR+EP+ ER+PYV+I+G PG L+
Sbjct: 2906 DFIFAKEYR-GSFSYKPGACVPALELTRKMLTYDRRSEPQVGERVPYVIIYGTPGVPLIQ 2964
Query: 1839 MVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKH 1898
+V P+EVL D R+N YYI KQI+P L R+F L+G D+ W+ E+PR + +A+
Sbjct: 2965 LVRRPVEVLQ-DPTLRLNATYYITKQILPPLARIFSLIGIDVFSWYHELPR-IHKATSSS 3022
Query: 1899 AFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQ 1958
P ++ I Y+ + HC +C L Q +CS+C A + + +LE++ +
Sbjct: 3023 RSEPEGRKGTISQYFTTLHCPVCDDLTQHG--ICSKCRSQPQHVAVILNQEIRELERQQE 3080
Query: 1959 HLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
LV IC++C G + + C S++C V ++ +V +EL A ++
Sbjct: 3081 QLVKICKNCTG---CFDRHIPCVSLNCPVLFKLSRVNRELSKAPYL 3123
Score = 189 bits (480), Expect = 8e-46
Identities = 120/346 (34%), Positives = 184/346 (52%), Gaps = 30/346 (8%)
Query: 10 VFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPY 69
+FSVRIV+ D+YMASP+ +D S + V +VPV+RV+G+TPAGQKTCLH+HG PY
Sbjct: 1 MFSVRIVTADYYMASPLQGLDTCQSPLTQAPVKKVPVVRVFGATPAGQKTCLHLHGIFPY 60
Query: 70 LYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVHGCSLVRAKKFYGYC 128
LYV Q+ ++Y +A S+++AL + G+ S+ QHV SLV FYGY
Sbjct: 61 LYVPYDG----YGQQPESYLSQMAFSIDRALNVALGNPSSTAQHVFKVSLVSGMPFYGYH 116
Query: 129 SFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGHL 188
E F+KIYL Y P V R LL +GA++NK QPHE+HIP++LQ +DYNLYGM +
Sbjct: 117 EKERHFMKIYL-YNPTMVKRICELLQSGAIMNKFYQPHEAHIPYLLQLFIDYNLYGMNLI 175
Query: 189 HLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMWSLPSESGG 248
+L+ ++FR +KS N+ H A ++ + +S + W +
Sbjct: 176 NLAAVKFR-----KARRKS--NTLH------ATGSCKNHLSGNSLADTLFRW---EQDEI 219
Query: 249 SSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMVQSLVSIWEEQRKR 308
S+ + QS CELE D +ILN+ + + L +IWE++++R
Sbjct: 220 PSSLILEGVEPQSTCELEVDAVAADILNR--------LDIEAQIGGNPGLQAIWEDEKQR 271
Query: 309 SGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKLIQLCSESDTTLS 354
T + + S F+K+L ++ ++D +++
Sbjct: 272 RRNRNETSQMSQPESQDHRFVPATESEKKFQKRLQEILKQNDFSVT 317
>UniRef100_Q5T0X9 REV3-like, catalytic subunit of DNA polymerase zeta [Homo sapiens]
Length = 3052
Score = 692 bits (1785), Expect = 0.0
Identities = 398/946 (42%), Positives = 558/946 (58%), Gaps = 66/946 (6%)
Query: 1078 EPGAKDKDISKCASGPTLRPELHQDTEKKLPCINEGQTERIKAHMDHSQDISQISGPGEK 1137
EP + K ++ TLR L + + +N Q E SQI GP
Sbjct: 2147 EPKTQKLSNKKGSNTDTLRRVLLTQAKNQFAAVNTPQKET-----------SQIDGPSLN 2195
Query: 1138 SSFTPLSQIGFQDPASAGHG-QQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
+++ I A A H Q L L+S+E+ A +R DL PDP+FD + + D
Sbjct: 2196 NTYGFKVSIQNLQEAKALHEIQNLTLISVELHARTRRDLEPDPEFDPICALFYCISSDTP 2255
Query: 1197 AT-------VEVLVL-----LHSKFVPCQRSL---DGLSDCKVLNFTDEKHLLKEFTKIV 1241
V+V+ + S+ + Q L G++ +V DEK L E I+
Sbjct: 2256 LPDTEKTELTGVIVIDKDKTVFSQDIRYQTPLLIRSGITGLEVTYAADEKALFHEIANII 2315
Query: 1242 SSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEP 1301
DPDIL+G+EIQ S G+L +RA+ L L +SR P D K + E
Sbjct: 2316 KRYDPDILLGYEIQMHSWGYLLQRAAALSIDLCRMISRVP--------DDKIENRFAAE- 2366
Query: 1302 DIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAE 1361
DE+G S +++ GRI LNLWR++R EV L Y+ E V+
Sbjct: 2367 ------------------RDEYGSYTMSEINIVGRITLNLWRIMRNEVALTNYTFENVSF 2408
Query: 1362 AVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARV 1421
VL ++ P VL+ WF R++ + + V R + NL++L QLD++ +TSE+AR+
Sbjct: 2409 HVLHQRFPLFTFRVLSDWFDNKTDLYRWKMVDHYVSRVRGNLQMLEQLDLIGKTSEMARL 2468
Query: 1422 FGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFY 1481
FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ + A +C+PL+MEPES FY
Sbjct: 2469 FGIQFLHVLTRGSQYRVESMMLRIAKPMNYIPVTPSVQQRSQMRAPQCVPLIMEPESRFY 2528
Query: 1482 SDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSPFVPEQHILQDLKDQ 1538
S+ V+VLDFQSLYPS++IAYN CF TCLG V G + +L ++
Sbjct: 2529 SNSVLVLDFQSLYPSIVIAYNYCFSTCLGHVENLGKYDEFKFGCTSLRVPPDLLYQVRHD 2588
Query: 1539 ILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALK 1598
I ++PNGV FV +R+GVLPR+LEEIL TR MVKQ+MK ++ L R+ +ARQL LK
Sbjct: 2589 ITVSPNGVAFVKPSVRKGVLPRMLEEILKTRFMVKQSMKAYK-QDRALSRMLDARQLGLK 2647
Query: 1599 LISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSM 1658
LI+NVT+GYT+A FSGRMPC E+ DSIV R TLE+AI VN +KW A+V+YGDTDSM
Sbjct: 2648 LIANVTFGYTSANFSGRMPCIEVGDSIVHKARETLERAIKLVNDTKKWGARVVYGDTDSM 2707
Query: 1659 FVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQ 1718
FVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L TKKRYVGY YE +Q
Sbjct: 2708 FVLLKGATKEQSFKIGQEIAEAVTATNPKPVKLKFEKVYLPCVLQTKKRYVGYMYETLDQ 2767
Query: 1719 IEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLK 1778
+PVFDAKGIETVRRD+C AV+KI+E+SL+L FE + + +K Y+QRQ ++L G+ S++
Sbjct: 2768 KDPVFDAKGIETVRRDSCPAVSKILERSLKLLFETRDISLIKQYVQRQCMKLLEGKASIQ 2827
Query: 1779 DFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVD 1838
DFIFAKE R G++S + + PA + K + DRR+EP+ ER+PYV+I+G PG L+
Sbjct: 2828 DFIFAKEYR-GSFSYKPGACVPALELTRKMLTYDRRSEPQVGERVPYVIIYGTPGVPLIQ 2886
Query: 1839 MVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKH 1898
+V P+EVL D R+N YYI KQI+P L R+F L+G D+ W+ E+PR + +A+
Sbjct: 2887 LVRRPVEVLQ-DPTLRLNATYYITKQILPPLARIFSLIGIDVFSWYHELPR-IHKATSSS 2944
Query: 1899 AFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQ 1958
P ++ I Y+ + HC +C L Q +CS+C A + + +LE++ +
Sbjct: 2945 RSEPEGRKGTISQYFTTLHCPVCDDLTQHG--ICSKCRSQPQHVAVILNQEIRELERQQE 3002
Query: 1959 HLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
LV IC++C G + + C S++C V ++ +V +EL A ++
Sbjct: 3003 QLVKICKNCTG---CFDRHIPCVSLNCPVLFKLSRVNRELSKAPYL 3045
Score = 109 bits (273), Expect = 8e-22
Identities = 80/264 (30%), Positives = 130/264 (48%), Gaps = 26/264 (9%)
Query: 92 VASSLEKALKLK-GSADSSRQHVHGCSLVRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAA 150
+A S+++AL + G+ S+ QHV SLV FYGY E F+KIYL Y P V R
Sbjct: 1 MAFSIDRALNVALGNPSSTAQHVFKVSLVSGMPFYGYHEKERHFMKIYL-YNPTMVKRIC 59
Query: 151 NLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGHLHLSKMRFRHPMPDSIHKKSDIN 210
LL +GA++NK QPHE+HIP++LQ +DYNLYGM ++L+ ++FR +KS N
Sbjct: 60 ELLQSGAIMNKFYQPHEAHIPYLLQLFIDYNLYGMNLINLAAVKFR-----KARRKS--N 112
Query: 211 SQHRKADPGAEACLESKIWMSSTISFDWMWSLPSESGGSSNDNTHSPKRQSICELEGDIS 270
+ H A ++ + +S + W + S+ + QS CELE D
Sbjct: 113 TLH------ATGSCKNHLSGNSLADTLFRW---EQDEIPSSLILEGVEPQSTCELEVDAV 163
Query: 271 VDEILNQQFKMFSSLSQTSSNVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMK 330
+ILN+ + + L +IWE++++R T + +
Sbjct: 164 AADILNR--------LDIEAQIGGNPGLQAIWEDEKQRRRNRNETSQMSQPESQDHRFVP 215
Query: 331 LLSSGLDFEKKLIQLCSESDTTLS 354
S F+K+L ++ ++D +++
Sbjct: 216 ATESEKKFQKRLQEILKQNDFSVT 239
>UniRef100_O60673 DNA polymerase zeta catalytic subunit [Homo sapiens]
Length = 3130
Score = 692 bits (1785), Expect = 0.0
Identities = 398/946 (42%), Positives = 558/946 (58%), Gaps = 66/946 (6%)
Query: 1078 EPGAKDKDISKCASGPTLRPELHQDTEKKLPCINEGQTERIKAHMDHSQDISQISGPGEK 1137
EP + K ++ TLR L + + +N Q E SQI GP
Sbjct: 2225 EPKTQKLSNKKGSNTDTLRRVLLTQAKNQFAAVNTPQKET-----------SQIDGPSLN 2273
Query: 1138 SSFTPLSQIGFQDPASAGHG-QQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
+++ I A A H Q L L+S+E+ A +R DL PDP+FD + + D
Sbjct: 2274 NTYGFKVSIQNLQEAKALHEIQNLTLISVELHARTRRDLEPDPEFDPICALFYCISSDTP 2333
Query: 1197 AT-------VEVLVL-----LHSKFVPCQRSL---DGLSDCKVLNFTDEKHLLKEFTKIV 1241
V+V+ + S+ + Q L G++ +V DEK L E I+
Sbjct: 2334 LPDTEKTELTGVIVIDKDKTVFSQDIRYQTPLLIRSGITGLEVTYAADEKALFHEIANII 2393
Query: 1242 SSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEP 1301
DPDIL+G+EIQ S G+L +RA+ L L +SR P D K + E
Sbjct: 2394 KRYDPDILLGYEIQMHSWGYLLQRAAALSIDLCRMISRVP--------DDKIENRFAAE- 2444
Query: 1302 DIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAE 1361
DE+G S +++ GRI LNLWR++R EV L Y+ E V+
Sbjct: 2445 ------------------RDEYGSYTMSEINIVGRITLNLWRIMRNEVALTNYTFENVSF 2486
Query: 1362 AVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARV 1421
VL ++ P VL+ WF R++ + + V R + NL++L QLD++ +TSE+AR+
Sbjct: 2487 HVLHQRFPLFTFRVLSDWFDNKTDLYRWKMVDHYVSRVRGNLQMLEQLDLIGKTSEMARL 2546
Query: 1422 FGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFY 1481
FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ + A +C+PL+MEPES FY
Sbjct: 2547 FGIQFLHVLTRGSQYRVESMMLRIAKPMNYIPVTPSVQQRSQMRAPQCVPLIMEPESRFY 2606
Query: 1482 SDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSPFVPEQHILQDLKDQ 1538
S+ V+VLDFQSLYPS++IAYN CF TCLG V G + +L ++
Sbjct: 2607 SNSVLVLDFQSLYPSIVIAYNYCFSTCLGHVENLGKYDEFKFGCTSLRVPPDLLYQVRHD 2666
Query: 1539 ILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALK 1598
I ++PNGV FV +R+GVLPR+LEEIL TR MVKQ+MK ++ L R+ +ARQL LK
Sbjct: 2667 ITVSPNGVAFVKPSVRKGVLPRMLEEILKTRFMVKQSMKAYK-QDRALSRMLDARQLGLK 2725
Query: 1599 LISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSM 1658
LI+NVT+GYT+A FSGRMPC E+ DSIV R TLE+AI VN +KW A+V+YGDTDSM
Sbjct: 2726 LIANVTFGYTSANFSGRMPCIEVGDSIVHKARETLERAIKLVNDTKKWGARVVYGDTDSM 2785
Query: 1659 FVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQ 1718
FVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L TKKRYVGY YE +Q
Sbjct: 2786 FVLLKGATKEQSFKIGQEIAEAVTATNPKPVKLKFEKVYLPCVLQTKKRYVGYMYETLDQ 2845
Query: 1719 IEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLK 1778
+PVFDAKGIETVRRD+C AV+KI+E+SL+L FE + + +K Y+QRQ ++L G+ S++
Sbjct: 2846 KDPVFDAKGIETVRRDSCPAVSKILERSLKLLFETRDISLIKQYVQRQCMKLLEGKASIQ 2905
Query: 1779 DFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVD 1838
DFIFAKE R G++S + + PA + K + DRR+EP+ ER+PYV+I+G PG L+
Sbjct: 2906 DFIFAKEYR-GSFSYKPGACVPALELTRKMLTYDRRSEPQVGERVPYVIIYGTPGVPLIQ 2964
Query: 1839 MVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKH 1898
+V P+EVL D R+N YYI KQI+P L R+F L+G D+ W+ E+PR + +A+
Sbjct: 2965 LVRRPVEVLQ-DPTLRLNATYYITKQILPPLARIFSLIGIDVFSWYHELPR-IHKATSSS 3022
Query: 1899 AFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQ 1958
P ++ I Y+ + HC +C L Q +CS+C A + + +LE++ +
Sbjct: 3023 RSEPEGRKGTISQYFTTLHCPVCDDLTQHG--ICSKCRSQPQHVAVILNQEIRELERQQE 3080
Query: 1959 HLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
LV IC++C G + + C S++C V ++ +V +EL A ++
Sbjct: 3081 QLVKICKNCTG---CFDRHIPCVSLNCPVLFKLSRVNRELSKAPYL 3123
Score = 189 bits (480), Expect = 8e-46
Identities = 120/346 (34%), Positives = 184/346 (52%), Gaps = 30/346 (8%)
Query: 10 VFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPY 69
+FSVRIV+ D+YMASP+ +D S + V +VPV+RV+G+TPAGQKTCLH+HG PY
Sbjct: 1 MFSVRIVTADYYMASPLQGLDTCQSPLTQAPVKKVPVVRVFGATPAGQKTCLHLHGIFPY 60
Query: 70 LYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVHGCSLVRAKKFYGYC 128
LYV Q+ ++Y +A S+++AL + G+ S+ QHV SLV FYGY
Sbjct: 61 LYVPYDG----YGQQPESYLSQMAFSIDRALNVALGNPSSTAQHVFKVSLVSGMPFYGYH 116
Query: 129 SFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGHL 188
E F+KIYL Y P V R LL +GA++NK QPHE+HIP++LQ +DYNLYGM +
Sbjct: 117 EKERHFMKIYL-YNPTMVKRICELLQSGAIMNKFYQPHEAHIPYLLQLFIDYNLYGMNLI 175
Query: 189 HLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMWSLPSESGG 248
+L+ ++FR +KS N+ H A ++ + +S + W +
Sbjct: 176 NLAAVKFR-----KARRKS--NTLH------ATGSCKNHLSGNSLADTLFRW---EQDEI 219
Query: 249 SSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMVQSLVSIWEEQRKR 308
S+ + QS CELE D +ILN+ + + L +IWE++++R
Sbjct: 220 PSSLILEGVEPQSTCELEVDAVAADILNR--------LDIEAQIGGNPGLQAIWEDEKQR 271
Query: 309 SGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKLIQLCSESDTTLS 354
T + + S F+K+L ++ ++D +++
Sbjct: 272 RRNRNETSQMSQPESQDHRFVPATESEKKFQKRLQEILKQNDFSVT 317
>UniRef100_UPI000023CD58 UPI000023CD58 UniRef100 entry
Length = 1681
Score = 691 bits (1784), Expect = 0.0
Identities = 385/888 (43%), Positives = 556/888 (62%), Gaps = 61/888 (6%)
Query: 1124 HSQDISQISGPGEKSSFTPLSQIGFQ-----DPASAGHG-QQLALLSIEVLAESRGDLLP 1177
H SQI GP TP ++ GF+ S H Q ++ +S+EV +RG+ +P
Sbjct: 807 HRNCHSQIEGP------TPKNRHGFKYTPGKKATSVKHEVQYMSTMSLEVHVNTRGNFVP 860
Query: 1178 DPQFDAVNIVALGFQKDCDAT--------VEVLVLLHSKFVPCQRSLDGLSDCKVLNFTD 1229
+P+ D V + D +T V+ +LL S + + + ++ T
Sbjct: 861 NPEEDEVQCAFWAIKSDGTSTDSQSSAGTVQTGILLLSNDAEFTQRVQRQTSADIIEETS 920
Query: 1230 EKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERAS-HLGFGLLNDLSRTPSNSWINS 1288
E L+ +IV + DPDIL G+E+ GSS G+L ERA + L ++ SR
Sbjct: 921 ELDLMVRMVEIVRNHDPDILTGYEVHGSSWGYLIERARLKYDYNLCDEFSR--------- 971
Query: 1289 QDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGE 1348
+KT D D WG S + V GR ++N+WR +RGE
Sbjct: 972 --MKTESHGRFGKD-----------------NDRWGFNTTSTIRVTGRHMVNVWRAMRGE 1012
Query: 1349 VKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQ 1408
+ L Y++E VA +L +++P + LT W+ G + + ++Y +R KL++EIL+
Sbjct: 1013 LNLLQYTMENVAWHLLHKRIPHYSWKSLTSWYKSGKHRELNRMLRYYQNRTKLDIEILDA 1072
Query: 1409 LDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAME 1468
+++ RTSE AR+ G+DFFSV SRGSQ++VES+ R+A +N++ +SP +QV Q A+E
Sbjct: 1073 NELIARTSEQARLLGVDFFSVFSRGSQFKVESIMFRIAKPENFLLVSPSRKQVGGQNALE 1132
Query: 1469 CLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSK-TNTLGVSPFVP 1527
CLPLVMEP+S FY+ P++VLDFQSLYPS++IAYN C+ T LG+V + T+ +G + +
Sbjct: 1133 CLPLVMEPQSAFYNSPLLVLDFQSLYPSVMIAYNYCYSTFLGRVTNWRGTSKMGFTEYKR 1192
Query: 1528 EQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQ 1587
+Q +L LKD I + PNG+M+ ++IR+ +L ++L EIL TR+MVK MK+ ++ LQ
Sbjct: 1193 QQGLLTLLKDYINIAPNGMMYAKTEIRKSLLAKMLTEILETRVMVKSGMKQ-DKDDKTLQ 1251
Query: 1588 RIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWN 1647
++ N RQLALKL++NVTYGYT+A FSGRMPC+E+ADSIVQ GR TLE+AI++++ EKW
Sbjct: 1252 QLLNNRQLALKLLANVTYGYTSASFSGRMPCSEIADSIVQTGRETLERAIAYIHSVEKWG 1311
Query: 1648 AKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKR 1707
A+V+YGDTDS+F+ LKGRT EAF IG+EI+ AIT MNP P+ LK EKVYHPC L+ KKR
Sbjct: 1312 AEVVYGDTDSLFISLKGRTKDEAFDIGNEISKAITEMNPQPIKLKFEKVYHPCVLLAKKR 1371
Query: 1708 YVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQW 1767
YVGY YE+ +Q++P FDAKGIETVRRD A I E++LRL FE L +VK Y Q+Q
Sbjct: 1372 YVGYKYESKDQVKPEFDAKGIETVRRDGTPAEQMIEEKALRLLFETADLSQVKEYFQKQC 1431
Query: 1768 KRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVV 1827
++I+ VS++DF FAKEVRLGTYS + + P A+++TK M D RAEP+Y ER+PYVV
Sbjct: 1432 QKIMRNNVSVQDFCFAKEVRLGTYSDK-GAPPAGALISTKRMLQDARAEPQYGERVPYVV 1490
Query: 1828 IHGEPGARLVDMVVDPLEVLAIDSP-FRINDLYYINKQIIPALQRVFGLVGADLNQWFAE 1886
I G PGARL+D V P E+L D+P ++++ YYI+K +IP L+R+F LVGA++ QW+ E
Sbjct: 1491 ITGAPGARLIDRCVAPEELL--DNPHWQLDAEYYISKNLIPPLERIFNLVGANVRQWYDE 1548
Query: 1887 MPRPVREASVKHAFTPNFQRTRIDYYYLSKHCVLCG-GLVQASARLCSQCSENEVAAATA 1945
MP+ R V HA T + ++T ++ Y S HC++C Q LC C N +A +
Sbjct: 1549 MPKVQR---VHHATTLSNKKTTLESYMKSTHCLVCNIKFSQEGNPLCPTCRTNIPSALLS 1605
Query: 1946 VIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRK 1993
+ + E+ +Q ++S+C+ C G + V+C S C VF+ R K
Sbjct: 1606 LQTRLTTEERCLQEILSLCRSCSGVGPV--EDVQCDSKDCPVFWTRMK 1651
Score = 155 bits (392), Expect = 1e-35
Identities = 112/342 (32%), Positives = 172/342 (49%), Gaps = 51/342 (14%)
Query: 9 DVFSVRIVSVDHYMAS-----PIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHI 63
D+F +R+ +DHY A+ P DI S +G +VP++R++G+T GQK C HI
Sbjct: 2 DLFRIRLNCIDHYQATATQYDPQLRNDIRPSQISKGP--KVPIVRIFGATETGQKVCAHI 59
Query: 64 HGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQH---VHGCSLVR 120
HGA PYLYV L D+ G AY Y + S++ AL + D + V +LV+
Sbjct: 60 HGAFPYLYVEYEG-GLSPDEVG-AYIYRLHLSIDHALAVSYRRDQKNDNARFVARITLVK 117
Query: 121 AKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDY 180
FYG+ FF+KIY+ + P ++R A+LL G ++ QP+E+H+ F+LQF+ D+
Sbjct: 118 GIPFYGFHVGYRFFLKIYM-FNPVVMTRLADLLQQGVIMKHKFQPYEAHLQFLLQFMTDF 176
Query: 181 NLYGMGHLHLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMW 240
NLYG +L S FR P+P+ + NS H +W S
Sbjct: 177 NLYGCDYLESSSTGFRSPVPE---YGEETNSSH--------------LWHSE-------- 211
Query: 241 SLPSESGGSSNDNTHSPKRQSICELEGDISVDEILN----QQFKMFSSLSQTS----SNV 292
S+P E D T P R S C LE DI V ILN Q+ + ++ + S++
Sbjct: 212 SIPQE---DVTDETTLP-RSSHCSLEVDICVQNILNRHRVQERPLHHDFTERTNPLPSDM 267
Query: 293 NMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLSS 334
+V S+ S+W+++ KR + S+ P P +V+ +S+
Sbjct: 268 KLVYSMASLWKDETKRRK-QKMQDPSEGSSPFPPEVLVSMSA 308
>UniRef100_Q61493 DNA polymerase zeta catalytic subunit [Mus musculus]
Length = 3122
Score = 691 bits (1782), Expect = 0.0
Identities = 389/898 (43%), Positives = 544/898 (60%), Gaps = 55/898 (6%)
Query: 1126 QDISQISGPGEKSSFTPLSQIGFQDPASAGHG-QQLALLSIEVLAESRGDLLPDPQFDAV 1184
++ SQI GP +++ I A A H Q L L+S+E+ A +R DL PDP+FD +
Sbjct: 2254 KETSQIDGPSLNNTYGFKVSIQNLQEAKALHEIQNLTLISVELHARTRRDLQPDPEFDPI 2313
Query: 1185 NIVALGFQKDCDAT-------VEVLVLLHSKFVPCQ--RSL------DGLSDCKVLNFTD 1229
+ D V+V+ K V Q RS G++ +V D
Sbjct: 2314 CALFYCISSDTPLPDTEKTELTGVIVIDKDKTVTHQDIRSQTPLLIRSGITGLEVTYAAD 2373
Query: 1230 EKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWINSQ 1289
EK L +E T I+ DPDIL+G+EIQ S G+L +RA+ L L +SR P
Sbjct: 2374 EKALFQEITNIIKRYDPDILLGYEIQMHSWGYLLQRAAALSVDLCQMISRVP-------- 2425
Query: 1290 DIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEV 1349
D K + E D++G S +++ GRI LNLWR++R EV
Sbjct: 2426 DDKIENRFAAE-------------------RDDYGSDTMSEINIVGRITLNLWRIMRNEV 2466
Query: 1350 KLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQL 1409
L Y+ E V+ VL ++ P VL+ WF R++ + + V R + NL++L QL
Sbjct: 2467 GLTNYTFENVSFHVLHQRFPLFTFRVLSDWFDNKTDLYRWKMVDHYVSRVRGNLQMLEQL 2526
Query: 1410 DMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMEC 1469
D++ +TSE+AR+FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ + A +C
Sbjct: 2527 DLIGKTSEMARLFGIQFLHVLTRGSQYRVESMMLRIAKPMNYIPVTPSIQQRSQMRAPQC 2586
Query: 1470 LPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSPFV 1526
+PL+MEPES FYS+ V+VLDFQSLYPS++IAYN CF TCLG V G +
Sbjct: 2587 VPLIMEPESRFYSNSVLVLDFQSLYPSIVIAYNYCFSTCLGHVENLGKYDEFKFGCTSLR 2646
Query: 1527 PEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVL 1586
+L ++ + ++PNGV FV +R+GVLPR+LEEIL TR+MVKQ+MK ++ L
Sbjct: 2647 VPPDLLYQIRHDVTVSPNGVAFVKPSVRKGVLPRMLEEILKTRLMVKQSMKSYK-QDRAL 2705
Query: 1587 QRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKW 1646
R+ NARQL LKLI+NVT+GYTAA FSGRMPC E+ DSIV R TLE+AI VN +KW
Sbjct: 2706 SRMLNARQLGLKLIANVTFGYTAANFSGRMPCIEVGDSIVHKARETLERAIKLVNDTKKW 2765
Query: 1647 NAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKK 1706
A+V+YGDTDSMFVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L TKK
Sbjct: 2766 GARVVYGDTDSMFVLLKGATKEQSFKIGQEIAEAVTATNPRPVKLKFEKVYLPCVLQTKK 2825
Query: 1707 RYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQ 1766
RYVGY YE +Q EPVFDAKGIETVRRD+C AV+KI+E+SL+L FE + + +K Y+ RQ
Sbjct: 2826 RYVGYMYETLDQKEPVFDAKGIETVRRDSCPAVSKILERSLKLLFETRDISLIKQYVHRQ 2885
Query: 1767 WKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYV 1826
+++ G+ S++DFIFAKE R G++S R + PA + K + DRR+EPR ER+PYV
Sbjct: 2886 CMKLVEGKASIQDFIFAKEYR-GSFSYRPGACVPALELTRKMLAYDRRSEPRVGERVPYV 2944
Query: 1827 VIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAE 1886
+I+G PG L+ ++ P EVL D R+N YYI KQI+P L R+F L+G D+ W+ E
Sbjct: 2945 IIYGTPGLPLIQLIRRPAEVLQ-DPTLRLNATYYITKQILPPLARIFSLIGIDVFSWYQE 3003
Query: 1887 MPRPVREASVKHAFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAV 1946
+PR +++A+ ++ I Y+ + HC +C L Q +CS+C A +
Sbjct: 3004 LPR-IQKATSSSRSELEGRKGTISQYFTTLHCPVCDDLTQHG--ICSKCRSQPQHVAIIL 3060
Query: 1947 IGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
+ +LE++ + L+ IC++C G + + C S++C V ++ +V +EL A ++
Sbjct: 3061 NQEIRELERKQEQLIKICRNCTGS---FDRHIPCVSLNCPVLFKLSRVNRELSKAPYL 3115
Score = 188 bits (478), Expect = 1e-45
Identities = 124/349 (35%), Positives = 191/349 (54%), Gaps = 36/349 (10%)
Query: 10 VFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPY 69
+FSVRIV+ D+YMASP+P +D S + V +VPV+RV+G+TPAGQKTCLH+HG PY
Sbjct: 1 MFSVRIVTADYYMASPLPGLDTCQSPLTQLPVKKVPVVRVFGATPAGQKTCLHLHGIFPY 60
Query: 70 LYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVHGCSLVRAKKFYGYC 128
LYV Q+ ++Y +A S+++AL + G+ S+ QHV SLV FYGY
Sbjct: 61 LYVPYDG----YGQQPESYLSQMAFSIDRALNVALGNPSSTAQHVFKVSLVSGMPFYGYH 116
Query: 129 SFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGHL 188
E F+KIYL Y P V R LL +GA++NK QPHE+HIP++LQ +DYNLYGM +
Sbjct: 117 EKERHFMKIYL-YNPAMVKRICELLQSGAIMNKCYQPHEAHIPYLLQLFIDYNLYGMNLI 175
Query: 189 HLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMWSLPSESGG 248
+L+ ++FR +K N+ H A + ++ +S + W E
Sbjct: 176 NLAAVKFR-----KARRKG--NASH------ATGLFKHQLSGNSPAGTLFRW---EEDEI 219
Query: 249 SSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMVQSLVSIWE-EQRK 307
S+ + S CELE D +ILN+ + + L +IWE E+++
Sbjct: 220 PSSLLLEGVEPLSTCELEVDAVAADILNR--------LDIEAQIGGNPGLQAIWEDEKQR 271
Query: 308 RSGIHEATMLSDSGKPLPED--VMKLLSSGLDFEKKLIQLCSESDTTLS 354
R +E++ +S +P +D + S F+K+L ++ ++D +++
Sbjct: 272 RRNRNESSQIS---QPESQDCRFVPATESEKQFQKRLQEVLKQNDFSVT 317
>UniRef100_UPI00003ACFBF UPI00003ACFBF UniRef100 entry
Length = 3114
Score = 682 bits (1760), Expect = 0.0
Identities = 387/900 (43%), Positives = 543/900 (60%), Gaps = 59/900 (6%)
Query: 1126 QDISQISGPGEKSSFT-PLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAV 1184
++ SQI GP +++ +S Q+ S Q L L+S+E+ A +R DL PDP+FD
Sbjct: 2246 RETSQIEGPSLNNTYGFKVSVQNLQEAKSLHEVQHLTLISMELHARTRRDLEPDPEFDP- 2304
Query: 1185 NIVALGFQKDCDATVE---------VLVLLHSKFVPCQRSLD--------GLSDCKVLNF 1227
I AL + D + +V+ + + Q S D G++ +V
Sbjct: 2305 -ICALFYCVSSDTALPNTDKKEITGAIVIDSDRTLSSQGSRDQAPLLTRSGITGLEVSYA 2363
Query: 1228 TDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWIN 1287
TDE+ L +E IV DPDIL+G+E+Q S G+L +RA+ L L +SR P + N
Sbjct: 2364 TDERTLFQEVVNIVKRYDPDILLGYEVQMHSWGYLLQRAAALNVDLCQMISRVPDD---N 2420
Query: 1288 SQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRG 1347
++ +E DE+G S +++ GRI+LN+WR++R
Sbjct: 2421 KENRFAAEL------------------------DEYGSDTMSEINIVGRIILNIWRMMRN 2456
Query: 1348 EVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILN 1407
EV L Y+ E V VL ++ P VL+ WF R++ + + V R NL++L
Sbjct: 2457 EVNLMNYTFENVGFHVLHQRFPLFTFRVLSDWFDNKADVYRWKMVDHYVSRVHGNLQLLQ 2516
Query: 1408 QLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAM 1467
+LD+++RTSE+AR+FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ A A
Sbjct: 2517 KLDLIDRTSEMARLFGIQFLHVLTRGSQYRVESMMLRVAKPMNYIPVTPSIQQRAQMKAP 2576
Query: 1468 ECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSP 1524
+C+PL+MEPES FYS+ V+VLDFQSLYPS++IAYN CF TCLG V G +
Sbjct: 2577 QCVPLIMEPESRFYSNAVLVLDFQSLYPSIVIAYNYCFSTCLGPVENLGKYDAFKFGCTS 2636
Query: 1525 FVPEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQ 1584
+L ++ I ++P+GV FV +R+GVLPR+LEEIL TRIMVKQ+MK ++
Sbjct: 2637 LRVPPDLLYQIRHDITVSPSGVAFVKPSVRKGVLPRMLEEILKTRIMVKQSMKAYK-HDK 2695
Query: 1585 VLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHE 1644
+ R+ ARQL LK I+N TYGYTAA FSGRMPC EL DSIV R TLE+AI VN +
Sbjct: 2696 AITRMLEARQLGLKYIANFTYGYTAANFSGRMPCIELGDSIVHKARETLERAIKLVNDTK 2755
Query: 1645 KWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLIT 1704
KW A V+YGDTDSMFVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L T
Sbjct: 2756 KWGAHVVYGDTDSMFVLLKGATKEQSFKIGQEIAEAVTATNPKPVKLKFEKVYLPCVLQT 2815
Query: 1705 KKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQ 1764
KKRYVGY YE +Q +PVFDAKGIETVRRD+C AV+KI+E S++L FE + + ++K Y+Q
Sbjct: 2816 KKRYVGYMYETLDQKDPVFDAKGIETVRRDSCPAVSKILECSIKLLFETRDISQIKQYVQ 2875
Query: 1765 RQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIP 1824
Q ++L G+ S++DFIFAKE R G+ + + S PA + + + DRR+EPR ER+P
Sbjct: 2876 NQCMKLLEGKASIQDFIFAKEYR-GSSAYKPGSCVPALEITRRMLAYDRRSEPRVGERVP 2934
Query: 1825 YVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWF 1884
YV+++G PG L+ +V P EVL D R+N YYI KQI+P L RVF L+G D+ W+
Sbjct: 2935 YVIVYGMPGLPLIQLVRRPTEVLQ-DPSLRLNATYYITKQILPPLARVFSLIGIDVFSWY 2993
Query: 1885 AEMPRPVREASVKHAFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAAT 1944
E+PR + AS + + T I Y+ + HC +C L Q +C++C A
Sbjct: 2994 HELPRIQKTASTARSELEGRKGT-ISQYFTTLHCPVCDELTQYG--ICTKCRSQPQHVAV 3050
Query: 1945 AVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
+ + +LE++ LV IC++C G + + C S++C V ++ +V +ELL A ++
Sbjct: 3051 ILNQEIRELERKQDQLVKICKNCTG---CFDRQIPCVSLNCPVLFKLSRVSRELLKAPYL 3107
Score = 146 bits (368), Expect = 7e-33
Identities = 104/304 (34%), Positives = 153/304 (50%), Gaps = 38/304 (12%)
Query: 56 GQKTCLHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVH 114
GQKTCLH+HG PYLYV Q + Y VA S+++AL + G+ S+ QHV
Sbjct: 1 GQKTCLHLHGVFPYLYVPYDGF----GQHPEHYLRQVAFSIDRALNVALGNPSSTIQHVF 56
Query: 115 GCSLVRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFIL 174
SLV FYGY E F+KIYL Y P V R LL GA++NKS QPHE+HIP++L
Sbjct: 57 KVSLVSGMPFYGYHEKERQFMKIYL-YNPAMVKRVCELLQGGAIMNKSYQPHEAHIPYLL 115
Query: 175 QFLVDYNLYGMGHLHLSKMRFRHPMPDSIHKKSDIN--SQHRKADPGAEACLESKIWMSS 232
Q +DYNLYGM ++L+ ++FR +KSD + S+H + + A + +S
Sbjct: 116 QLFIDYNLYGMNLINLAAVKFR-----KARRKSDTSGGSKHDRINSAANS--------NS 162
Query: 233 TISFDWMWSLPSESGGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNV 292
+W E S+ + QS CELE D +ILNQ + +
Sbjct: 163 ASFIEW-----EEDEIPSSLMLKGVEPQSTCELEVDAVAADILNQ--------LDIEAQI 209
Query: 293 NMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPED--VMKLLSSGLDFEKKLIQLCSESD 350
L +IWE++++R E S P +D + S F+K+L ++ ++D
Sbjct: 210 GRNPGLQAIWEDEKQRR--REKCESSQIKTPESQDRGFVPATESEKYFQKRLKEILKQND 267
Query: 351 TTLS 354
+++
Sbjct: 268 FSVT 271
>UniRef100_UPI00003ACFBE UPI00003ACFBE UniRef100 entry
Length = 3076
Score = 682 bits (1760), Expect = 0.0
Identities = 387/900 (43%), Positives = 543/900 (60%), Gaps = 59/900 (6%)
Query: 1126 QDISQISGPGEKSSFT-PLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAV 1184
++ SQI GP +++ +S Q+ S Q L L+S+E+ A +R DL PDP+FD
Sbjct: 2208 RETSQIEGPSLNNTYGFKVSVQNLQEAKSLHEVQHLTLISMELHARTRRDLEPDPEFDP- 2266
Query: 1185 NIVALGFQKDCDATVE---------VLVLLHSKFVPCQRSLD--------GLSDCKVLNF 1227
I AL + D + +V+ + + Q S D G++ +V
Sbjct: 2267 -ICALFYCVSSDTALPNTDKKEITGAIVIDSDRTLSSQGSRDQAPLLTRSGITGLEVSYA 2325
Query: 1228 TDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWIN 1287
TDE+ L +E IV DPDIL+G+E+Q S G+L +RA+ L L +SR P + N
Sbjct: 2326 TDERTLFQEVVNIVKRYDPDILLGYEVQMHSWGYLLQRAAALNVDLCQMISRVPDD---N 2382
Query: 1288 SQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRG 1347
++ +E DE+G S +++ GRI+LN+WR++R
Sbjct: 2383 KENRFAAEL------------------------DEYGSDTMSEINIVGRIILNIWRMMRN 2418
Query: 1348 EVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILN 1407
EV L Y+ E V VL ++ P VL+ WF R++ + + V R NL++L
Sbjct: 2419 EVNLMNYTFENVGFHVLHQRFPLFTFRVLSDWFDNKADVYRWKMVDHYVSRVHGNLQLLQ 2478
Query: 1408 QLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAM 1467
+LD+++RTSE+AR+FGI F VL+RGSQYRVESM LR+A NY+ ++P QQ A A
Sbjct: 2479 KLDLIDRTSEMARLFGIQFLHVLTRGSQYRVESMMLRVAKPMNYIPVTPSIQQRAQMKAP 2538
Query: 1468 ECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVV---PSKTNTLGVSP 1524
+C+PL+MEPES FYS+ V+VLDFQSLYPS++IAYN CF TCLG V G +
Sbjct: 2539 QCVPLIMEPESRFYSNAVLVLDFQSLYPSIVIAYNYCFSTCLGPVENLGKYDAFKFGCTS 2598
Query: 1525 FVPEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQ 1584
+L ++ I ++P+GV FV +R+GVLPR+LEEIL TRIMVKQ+MK ++
Sbjct: 2599 LRVPPDLLYQIRHDITVSPSGVAFVKPSVRKGVLPRMLEEILKTRIMVKQSMKAYK-HDK 2657
Query: 1585 VLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHE 1644
+ R+ ARQL LK I+N TYGYTAA FSGRMPC EL DSIV R TLE+AI VN +
Sbjct: 2658 AITRMLEARQLGLKYIANFTYGYTAANFSGRMPCIELGDSIVHKARETLERAIKLVNDTK 2717
Query: 1645 KWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLIT 1704
KW A V+YGDTDSMFVLLKG T +++F+IG EIA A+TA NP PV LK EKVY PC L T
Sbjct: 2718 KWGAHVVYGDTDSMFVLLKGATKEQSFKIGQEIAEAVTATNPKPVKLKFEKVYLPCVLQT 2777
Query: 1705 KKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQ 1764
KKRYVGY YE +Q +PVFDAKGIETVRRD+C AV+KI+E S++L FE + + ++K Y+Q
Sbjct: 2778 KKRYVGYMYETLDQKDPVFDAKGIETVRRDSCPAVSKILECSIKLLFETRDISQIKQYVQ 2837
Query: 1765 RQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIP 1824
Q ++L G+ S++DFIFAKE R G+ + + S PA + + + DRR+EPR ER+P
Sbjct: 2838 NQCMKLLEGKASIQDFIFAKEYR-GSSAYKPGSCVPALEITRRMLAYDRRSEPRVGERVP 2896
Query: 1825 YVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWF 1884
YV+++G PG L+ +V P EVL D R+N YYI KQI+P L RVF L+G D+ W+
Sbjct: 2897 YVIVYGMPGLPLIQLVRRPTEVLQ-DPSLRLNATYYITKQILPPLARVFSLIGIDVFSWY 2955
Query: 1885 AEMPRPVREASVKHAFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAAT 1944
E+PR + AS + + T I Y+ + HC +C L Q +C++C A
Sbjct: 2956 HELPRIQKTASTARSELEGRKGT-ISQYFTTLHCPVCDELTQYG--ICTKCRSQPQHVAV 3012
Query: 1945 AVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHV 2004
+ + +LE++ LV IC++C G + + C S++C V ++ +V +ELL A ++
Sbjct: 3013 ILNQEIRELERKQDQLVKICKNCTG---CFDRQIPCVSLNCPVLFKLSRVSRELLKAPYL 3069
Score = 146 bits (368), Expect = 7e-33
Identities = 104/304 (34%), Positives = 153/304 (50%), Gaps = 38/304 (12%)
Query: 56 GQKTCLHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVH 114
GQKTCLH+HG PYLYV Q + Y VA S+++AL + G+ S+ QHV
Sbjct: 1 GQKTCLHLHGVFPYLYVPYDGF----GQHPEHYLRQVAFSIDRALNVALGNPSSTIQHVF 56
Query: 115 GCSLVRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFIL 174
SLV FYGY E F+KIYL Y P V R LL GA++NKS QPHE+HIP++L
Sbjct: 57 KVSLVSGMPFYGYHEKERQFMKIYL-YNPAMVKRVCELLQGGAIMNKSYQPHEAHIPYLL 115
Query: 175 QFLVDYNLYGMGHLHLSKMRFRHPMPDSIHKKSDIN--SQHRKADPGAEACLESKIWMSS 232
Q +DYNLYGM ++L+ ++FR +KSD + S+H + + A + +S
Sbjct: 116 QLFIDYNLYGMNLINLAAVKFR-----KARRKSDTSGGSKHDRINSAANS--------NS 162
Query: 233 TISFDWMWSLPSESGGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNV 292
+W E S+ + QS CELE D +ILNQ + +
Sbjct: 163 ASFIEW-----EEDEIPSSLMLKGVEPQSTCELEVDAVAADILNQ--------LDIEAQI 209
Query: 293 NMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPED--VMKLLSSGLDFEKKLIQLCSESD 350
L +IWE++++R E S P +D + S F+K+L ++ ++D
Sbjct: 210 GRNPGLQAIWEDEKQRR--REKCESSQIKTPESQDRGFVPATESEKYFQKRLKEILKQND 267
Query: 351 TTLS 354
+++
Sbjct: 268 FSVT 271
>UniRef100_UPI00003C1736 UPI00003C1736 UniRef100 entry
Length = 1678
Score = 676 bits (1743), Expect = 0.0
Identities = 371/881 (42%), Positives = 526/881 (59%), Gaps = 77/881 (8%)
Query: 1158 QQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEV---------LVLLHSK 1208
Q + L++EV+ +R L PDP DA+ + FQ + + + L+++
Sbjct: 796 QHMTTLAVEVMTTTRDSLFPDPALDAIQSIVYSFQNEDENLQDTGSRPDLRTGLIIVSGD 855
Query: 1209 FVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERAS- 1267
+ R GLS V DE L +V + DP+IL+GWEI SS G++ ERAS
Sbjct: 856 EINFDRL--GLSHLAVEVVEDELELFNTLIDLVRAFDPEILVGWEIHSSSWGYIVERASK 913
Query: 1268 HLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTH 1327
+ L+ + R I + KS D + T
Sbjct: 914 EYDYDLVPQIGRA-----IIHNTGRAGGKS-----------------------DNYAYTQ 945
Query: 1328 ASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQA 1387
+S + GR LNLWRL++GE+ LNLY++E V +L R++P +H LT+W+ G +
Sbjct: 946 SSALRFTGRHTLNLWRLMKGELTLNLYTLENVCYHLLHRRIPKYSHQTLTQWYKSGRAEL 1005
Query: 1388 RYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAH 1447
+ + Y VD +L LEI+ + + V RT+E AR++GIDFFSV+SRGSQ++VES+ LR+A
Sbjct: 1006 IRRALLYYVDMVELELEIIAESEFVLRTAEFARIYGIDFFSVISRGSQFKVESIMLRIAK 1065
Query: 1448 TQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCT 1507
+N+V ISP QV Q A ECLPL+MEP+S FY P++VLDFQSLYPS++IAYNLC+ T
Sbjct: 1066 PENFVLISPSRAQVGKQNAAECLPLIMEPQSAFYKGPLIVLDFQSLYPSIMIAYNLCYST 1125
Query: 1508 CLGKVVPSK-TNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEIL 1566
CLG+V K T+ GV+ + P + +L L+D + ++ NG++FV +R+ +L ++L EIL
Sbjct: 1126 CLGRVAGFKGTSKFGVTEYAPPKGLLSLLQDDVNISSNGLLFVKPSVRKSLLAKMLSEIL 1185
Query: 1567 STRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIV 1626
TR+MVK +MK + S++ QRI NARQ++LKL++NVTYGYT+A FSGRMPC E+AD+IV
Sbjct: 1186 DTRVMVKSSMKA-TKSDKSFQRIQNARQISLKLLANVTYGYTSASFSGRMPCVEIADAIV 1244
Query: 1627 QCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNP 1686
Q GR TLEKA+ +N E+W A+V+YGDTDS+FV L GRT +EAF +G+ IA +T++NP
Sbjct: 1245 QTGRETLEKAMDLINGTEEWGAQVVYGDTDSLFVYLPGRTKEEAFTLGNVIADKVTSLNP 1304
Query: 1687 SPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQS 1746
PV LK EKVY P L+ KKRYVG+ YE ++ EP FDAKGIETVRRD A+ +++E
Sbjct: 1305 RPVKLKFEKVYLPSVLLAKKRYVGFKYETLDEREPGFDAKGIETVRRDFHPAIQRMLEAC 1364
Query: 1747 LRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVAT 1806
+R+ F + L VK+YLQRQW++IL GRVS +DFIFAKEVRLG+YS +++ PP A VA+
Sbjct: 1365 IRILFRSRDLSLVKSYLQRQWRKILEGRVSAQDFIFAKEVRLGSYSDKVAP-PPGAAVAS 1423
Query: 1807 KAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQII 1866
+ M D R+EP+Y ER+PY++ GEP A+L V P E D +IN LYYI + II
Sbjct: 1424 RRMLADPRSEPQYGERVPYIISQGEPRAKLNQQAVSP-EAFLKDPQLQINALYYITRTII 1482
Query: 1867 PALQRVFGLVGADLNQWFAEMPRP-----------------------------VREASVK 1897
P L RVF L+GAD+ WF MP+P + S
Sbjct: 1483 PPLARVFNLLGADVEAWFHHMPKPNNAYWNKRPTAALLSALLAQQDPSSSTHEIGSDSAA 1542
Query: 1898 HAFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEM 1957
T + ++ +Y ++ C+LC + A +C C ++ + + A K KLE ++
Sbjct: 1543 SKATSSTAIATLNSHYATETCLLCSS--RTEALVCLDCRQSPLESIHATSSKLHKLEAQV 1600
Query: 1958 QHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKEL 1998
+ SIC C +R V C S C Y++ KV K L
Sbjct: 1601 AAIDSICSTCAHMER--PGPVDCVSFDCPYTYQKTKVHKSL 1639
Score = 128 bits (322), Expect = 2e-27
Identities = 93/323 (28%), Positives = 157/323 (47%), Gaps = 44/323 (13%)
Query: 2 SDSESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGK-VNEVPVIRVYGSTPAGQKTC 60
S S S+ F VR++++DH + P P S + G+ + +VPV+R++G+TPAGQ+ C
Sbjct: 26 SASASSDPFFRVRLINIDHILTVPTPFDRTSCAFNAEGQPLRKVPVLRLFGATPAGQRVC 85
Query: 61 LHIHGALPYLYVSCS------DIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVH 114
L+IH PY Y+ ++ + + G +A+SL + L + Q +
Sbjct: 86 LYIHNVYPYCYIQYKGSLDPDNVLRYIHRLGRGLNAAMAASLRRNLH----DTDANQFIA 141
Query: 115 GCSLVRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFIL 174
L + FYGY +++KI P R A +L +G V+ QP E HI + L
Sbjct: 142 AIHLCKGVNFYGYHVGYSYYLKISF-VDPAHNYRIAAILESGGVMKTVFQPFEIHIRYQL 200
Query: 175 QFLVDYNLYGMGHLHLSKMRFRHPMPD-SIHKKSDINSQHRKADPGAEACLESKIWMSST 233
QF++DYN++G ++ L +RFR P+P+ ++ D++ + SKIW ++
Sbjct: 201 QFMLDYNIFGCDYVDLDDIRFRLPIPEGNLVANDDLSPLDSR----------SKIWNRNS 250
Query: 234 ISFDWMWSLPSESGGSSNDNTHSPKRQSICELEGDISVDEILN----QQFKMFSSLSQTS 289
I + ++ R S CELE D S I+N +Q + S+ +TS
Sbjct: 251 IPY-------------AHVQAPDVHRGSYCELEADASAPWIINRRRFKQRDIHSNFDETS 297
Query: 290 SNVN----MVQSLVSIWEEQRKR 308
+ + +V SL +WE++ KR
Sbjct: 298 TTPSDQDILVPSLSGLWEDEMKR 320
>UniRef100_Q8X142 Polymerase zeta subunit [Emericella nidulans]
Length = 1681
Score = 660 bits (1704), Expect = 0.0
Identities = 375/881 (42%), Positives = 528/881 (59%), Gaps = 49/881 (5%)
Query: 1128 ISQISGPGEKSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIV 1187
+SQI GP +K+ + + Q ++ +S+EV +RG LLP+P+ D +
Sbjct: 806 LSQIEGPTQKNEHGFKYSQHKKSTSVEHQTQYMSTMSLEVHVNTRGVLLPNPEEDEIAAF 865
Query: 1188 ALGFQKDCDATVE---VLVLLHSKFVPCQRSLDGLSDCKVLNF-----TDEKHLLKEFTK 1239
+ + E + V+ + K L F + E L+
Sbjct: 866 FWCVNSEDEPQEEDDSLSVVNMGVVFQGEEEYPESKISKALRFECEHESTELDLINRLVD 925
Query: 1240 IVSSSDPDILMGWEIQGSSLGFLAERA-SHLGFGLLNDLSRTPSNSWINSQDIKTSEKSI 1298
IV DPDI+ G+E+ SS GF+ ERA F L ++LSR S++
Sbjct: 926 IVRLHDPDIITGYEVHNSSWGFIIERARKKYDFDLCDELSRVKSHAHGRF---------- 975
Query: 1299 LEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEA 1358
+E+ D+WG H S + + GR ++N+WR +R E+ L Y++E
Sbjct: 976 --------------GKEA----DKWGFDHTSSIRITGRHIINIWRAMRSEMNLLQYTMEN 1017
Query: 1359 VAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSEL 1418
V +L R++P + LT W+ + + I Y R ++N+EILN +++ RTSE
Sbjct: 1018 VVFHLLHRRIPHYSFRDLTAWYQSDKPRDFMKVINYYTSRVQMNIEILNANELIPRTSEQ 1077
Query: 1419 ARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPES 1478
AR+ GIDF+SV SRGSQ++VES+ R+A +N++ +SP +QV Q A+ECLPLVMEP+S
Sbjct: 1078 ARLLGIDFYSVFSRGSQFKVESLMFRIAKPENFILVSPSRKQVGQQNALECLPLVMEPQS 1137
Query: 1479 GFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSK-TNTLGVSPFVPEQHILQDLKD 1537
FY+ P++VLDFQSLYPS++IAYN C+ T LG+V + N +G + +L+ LKD
Sbjct: 1138 DFYTSPLIVLDFQSLYPSIMIAYNYCYSTFLGRVQSWRGRNKMGFLDYNRPPRLLELLKD 1197
Query: 1538 QILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLAL 1597
I + PNG+++ S++R+ +L ++L EIL TR+MVK MK + ++ LQR+ N RQLAL
Sbjct: 1198 HINIAPNGMIYARSEVRKSLLAKMLTEILETRVMVKTGMK-VDKDDRALQRLLNNRQLAL 1256
Query: 1598 KLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDS 1657
KLI+NVTYGYT+A FSGRMPC+E+ADSIVQ GR TLEKAI+F++ E+W A+V+YGDTDS
Sbjct: 1257 KLIANVTYGYTSASFSGRMPCSEIADSIVQSGRETLEKAIAFIHSVERWGAEVVYGDTDS 1316
Query: 1658 MFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPN 1717
+FV LKGRT +AF IG EIA A+T MNP PV LK EKVYHPC L+ KKRYVG+ YE+ N
Sbjct: 1317 LFVYLKGRTRDQAFDIGEEIAQAVTNMNPHPVKLKFEKVYHPCVLLAKKRYVGFKYEHRN 1376
Query: 1718 QIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSL 1777
Q EP FDAKGIETVRRD A KI E++L+L F L +VK+Y Q Q +I+ GRVS+
Sbjct: 1377 QKEPEFDAKGIETVRRDGTPAEQKIEEKALKLLFRTADLSQVKSYFQSQCTKIMQGRVSV 1436
Query: 1778 KDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLV 1837
+DF FA+ V+LGTYS +LP A+++TK M D RAEP+Y ER+PYVV+ G PG+RL+
Sbjct: 1437 QDFCFARAVKLGTYSEN-GTLPAGALISTKKMLEDPRAEPQYGERVPYVVVTGAPGSRLI 1495
Query: 1838 DMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVR----E 1893
D V P E L D+ ++ YYI K IIP L+R+F LVGA++ QW+ EMP+ R E
Sbjct: 1496 DRCVPP-ETLLQDAQLDLDAEYYITKNIIPPLERIFNLVGANVRQWYDEMPKVHRIRRVE 1554
Query: 1894 ASVKHAFTPNFQRTRIDYYYLSKHCVLC-GGLVQASARLCSQCSENEVAAATAVIGKTAK 1952
+ A + RT ++ Y + CV+C L A +CS C + ++ +
Sbjct: 1555 GTSSLAGGKSSIRT-LESYMKASSCVVCKAKLYDAEIPVCSDCVRQPHVSLLELVSRQRH 1613
Query: 1953 LEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRK 1993
EQ + L IC+ C D VKC S+ C VFY R +
Sbjct: 1614 AEQRVATLERICRSC--MDVPFGDEVKCDSLDCPVFYARTR 1652
Score = 152 bits (383), Expect = 1e-34
Identities = 105/314 (33%), Positives = 159/314 (50%), Gaps = 50/314 (15%)
Query: 11 FSVRIVSVDHYMASPI---PSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGAL 67
F VR+ VDHY A+P P + +G + +VPVIR++G+T GQK C+H+HGA
Sbjct: 4 FQVRLNCVDHYQANPSEFDPPLPYRDGNNEKGYMPKVPVIRIFGTTETGQKICVHVHGAF 63
Query: 68 PYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKL---KGSADSSRQHVHGCSLVRAKKF 124
PYLYV D L D A + S++ AL + + + D V +LV+ F
Sbjct: 64 PYLYVQY-DGDLSPDSVRSA-ARSLHLSIDHALAVSYRRNAHDKKTVFVAHITLVKGVPF 121
Query: 125 YGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYG 184
YGY FF KIYL P ++R A+LLL GAV+ + +QP+ESH+ ++ Q++ DYNL+G
Sbjct: 122 YGYHVGYRFFFKIYL-LNPIYITRLADLLLQGAVMKRPIQPYESHLQYVPQWMCDYNLFG 180
Query: 185 MGHLHLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMWSLPS 244
+ K +FR P+P+ + + HR W +I W+
Sbjct: 181 CAFMKCGKAKFRSPIPEYLDLP---DLSHR--------------WHDRSIPPGWIL---- 219
Query: 245 ESGGSSNDNTHSPKRQSICELEGDISVDEILNQ----------QFKMFSSLSQTSSNVNM 294
D + PK QS C LE D+ V +ILN+ F+ F L+ +SN +
Sbjct: 220 -------DESVLPK-QSHCPLEADVCVQDILNRLEISERSIHHDFREF--LNPLTSNEKL 269
Query: 295 VQSLVSIWEEQRKR 308
V S+ +WE++++R
Sbjct: 270 VPSMAGLWEDEKRR 283
>UniRef100_Q8X1U8 Catalytic subunit of DNA polymerase zeta [Neurospora crassa]
Length = 1926
Score = 632 bits (1629), Expect = e-179
Identities = 348/780 (44%), Positives = 495/780 (62%), Gaps = 56/780 (7%)
Query: 1129 SQISGPGEKSSFTPLSQIGFQDP------ASAGHGQQLALLSIEVLAESRGDLLPDPQFD 1182
SQI+G TP ++ GF+ P + Q ++ +S+EV +RG L+PDP+ D
Sbjct: 898 SQIAG------VTPKNKHGFEHPEKTKSESKQDQAQYMSAMSLEVHVNTRGKLVPDPEED 951
Query: 1183 AVNIV---------ALGFQKDCDATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHL 1233
V V AL + D T +++ + R S V+ T E +
Sbjct: 952 EVQCVFWYLRSEVNALRGTQTPDDTARGIIVFSEDSLLADRIRKHTS-VPVVQETTELDM 1010
Query: 1234 LKEFTKIVSSSDPDILMGWEIQGSSLGFLAERAS-HLGFGLLNDLSRTPSNSWINSQDIK 1292
+ +IV + DPDI G+E+ GSS G+L ERA L ++ SR S S N + K
Sbjct: 1011 MVRMVEIVRNHDPDIFTGYEVHGSSWGYLIERARIKYELDLCDEFSRMKSQS--NGRIGK 1068
Query: 1293 TSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLN 1352
+ D WG S + + GR ++N+WR +RGE+ L
Sbjct: 1069 DA--------------------------DRWGFNTTSSIRITGRHMINIWRAMRGELNLL 1102
Query: 1353 LYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMV 1412
Y++E V +L R++P + L+ W+ + + ++Y + R +L++EIL + +++
Sbjct: 1103 QYTMENVVWHLLHRRIPHYSWKTLSDWYLSDRPKDLDKVLRYYLTRTRLDIEILEKNELI 1162
Query: 1413 NRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPL 1472
RTSE AR+ G+DFFSV SRGSQ++VES+ R+A +N++ SP +QV +Q A+ECLPL
Sbjct: 1163 PRTSEQARLLGVDFFSVFSRGSQFKVESIMFRIAKPENFLLPSPSRKQVGAQNALECLPL 1222
Query: 1473 VMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSK-TNTLGVSPFVPEQHI 1531
VMEP+S FYS P++VLDFQSLYPS++IAYN C+ T LG++V + N +G + ++ +
Sbjct: 1223 VMEPQSAFYSSPLLVLDFQSLYPSVMIAYNYCYSTFLGRIVSWRGRNKMGFMDYKRQEGL 1282
Query: 1532 LQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFN 1591
L LKD I + PNG+M+ IR+ +L ++L EIL TRIMVK MK+ ++ +Q++ N
Sbjct: 1283 LSLLKDYINIAPNGMMYTKPHIRKSLLAKMLTEILETRIMVKSGMKQ-DKDDRAIQQLLN 1341
Query: 1592 ARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVI 1651
RQLALKL++NVTYGYT+A FSGRMPC+E+ADSIVQ GR TLE+AI+F++ +KW+A V+
Sbjct: 1342 NRQLALKLLANVTYGYTSASFSGRMPCSEIADSIVQTGRETLERAIAFIHSVQKWDADVV 1401
Query: 1652 YGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGY 1711
YGDTDS+FV LKGRT ++AF+IG EIA A+T +NP PV LK EKVYHPC L+ KKRYVGY
Sbjct: 1402 YGDTDSLFVSLKGRTREQAFEIGQEIADAVTKLNPRPVKLKFEKVYHPCVLLAKKRYVGY 1461
Query: 1712 SYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRIL 1771
YE+ +Q PVFDAKGIETVRRD A +I E++L++ FE L +VK+Y Q Q +I+
Sbjct: 1462 KYESRDQTVPVFDAKGIETVRRDGTPAEQRIEEKALKILFETADLSQVKSYFQEQCHKIM 1521
Query: 1772 SGRVSLKDFIFAKEVRLGTY--SARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIH 1829
G VS++DF FA+EV+LGTY S R P A++ATK M+ D RAEP+Y ER+PYVV+
Sbjct: 1522 RGAVSVQDFCFAREVKLGTYSTSGRGGPAPAGALIATKKMKEDARAEPQYGERVPYVVMA 1581
Query: 1830 GEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPR 1889
G PG RLVD V+P E+L ++ ++ YYINK IIP L+R+F LVGA++ W+ EMP+
Sbjct: 1582 GAPGMRLVDRCVEPEELLN-NAHATLDADYYINKNIIPPLERIFNLVGANVRTWYEEMPK 1640
Score = 146 bits (368), Expect = 7e-33
Identities = 110/340 (32%), Positives = 163/340 (47%), Gaps = 64/340 (18%)
Query: 13 VRIVSVDHYMASPI---PSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPY 69
+R+ +DHY A+P P D K +VPVIRV+GST GQK C HIHGA PY
Sbjct: 6 LRLNCIDHYQATPTRYDPQFDQDVRFSRSRKAAKVPVIRVFGSTDKGQKVCAHIHGAFPY 65
Query: 70 LYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSR---QHVHGCSLVRAKKFYG 126
LYV Y + + + AL + D R ++V SL + FYG
Sbjct: 66 LYVE--------------YDGNLEPNKDHALAISYRKDPIRDRPKYVARISLTKGIPFYG 111
Query: 127 YCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMG 186
+ F++KIYL + P +SR +LL G ++++ QP+E+H+ ++LQF+ DYNLYG
Sbjct: 112 FHVGYRFYLKIYL-FNPVVMSRLVDLLQQGVIMSRKFQPYEAHLQYLLQFMADYNLYGCN 170
Query: 187 HLHLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDWMWSLPSES 246
+L + FR P+P K D N + R+A+ W +TI PSE
Sbjct: 171 YLDAAMATFRAPVP-----KHDSNIEGREAE---------HHWDDTTI--------PSE- 207
Query: 247 GGSSNDNTHSPKRQSICELEGDISVDEILNQQ--------FKMFSSLSQTSSNVNMVQSL 298
DN +S R S C LE DI V++ILN++ SS +V S+
Sbjct: 208 --LITDN-YSLPRASHCSLEVDICVEDILNRKQVKERRLHHDFIEKEQPVSSQEKLVHSM 264
Query: 299 VSIWEEQ----RKRSGIHEATMLSDSGKPLPEDVMKLLSS 334
+W ++ +KR GI + + P P +V+ +S+
Sbjct: 265 AGLWTDETNRRKKRMGITDPEV-----NPFPPEVLVSMSA 299
Score = 40.8 bits (94), Expect = 0.43
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 2/82 (2%)
Query: 1923 GLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTS 1982
G V LC +C+ + + K + E+ ++ +CQ C G L V C S
Sbjct: 1751 GAVDRGLPLCHRCASDPPTLMVDMQAKVNRAEKSYVEIMKVCQSCAG--FALSEEVPCDS 1808
Query: 1983 ISCLVFYERRKVQKELLAATHV 2004
C VFY R K + ++ A V
Sbjct: 1809 KDCPVFYSRVKQRTKVTAVKRV 1830
>UniRef100_UPI000021B964 UPI000021B964 UniRef100 entry
Length = 1782
Score = 625 bits (1612), Expect = e-177
Identities = 345/785 (43%), Positives = 498/785 (62%), Gaps = 57/785 (7%)
Query: 1126 QDISQISGPGEKSSFTPLSQIGF-----QDPASAGHG-QQLALLSIEVLAESRGDLLPDP 1179
+ +SQI G TP ++ GF Q S H ++ +SIE+ +RG L+PDP
Sbjct: 832 RQLSQIEGA------TPKNKHGFRYSQKQKTTSVQHEVAYMSTMSIEIHVNTRGKLVPDP 885
Query: 1180 QFDAVNIVALGFQKDC----------DATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTD 1229
+ D V + + D D +V+L + +R + ++L T
Sbjct: 886 EKDEVKCIFWCVRSDDKSNEASSQAPDGLQSGVVVLSEEGTLAERIKHQIKG-ELLEETS 944
Query: 1230 EKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHL-GFGLLNDLSRTPSNSWINS 1288
E L+ +IV + DPDIL G+E+ G S G+L ERA + + L ++ SR S S +
Sbjct: 945 ELDLMVRMVEIVRTHDPDILTGYEVHGGSWGYLIERARCMYDYNLCDEFSRMKSQS--HG 1002
Query: 1289 QDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGE 1348
+ K + D WG S + V GR ++N+WR +RGE
Sbjct: 1003 RYGKDA--------------------------DRWGFNTTSTIRVTGRHMINIWRAMRGE 1036
Query: 1349 VKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQ 1408
+ L Y++E V +L R++P + LT W+ G + ++Y R +L++EIL +
Sbjct: 1037 LNLLQYTMENVVWHLLHRRIPHYSWQTLTTWYGSGRHGDLDKLLRYYQTRTRLDIEILEE 1096
Query: 1409 LDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAME 1468
+++RTSE AR+ G+DFFSV SRGSQ++VES+ R+A +N++ +SP +QV Q A+E
Sbjct: 1097 NGLISRTSEQARLLGVDFFSVFSRGSQFKVESIMFRIAKPENFMLVSPSRKQVGGQNALE 1156
Query: 1469 CLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSK-TNTLGVSPFVP 1527
CLPLVMEP+S FY+ PV+VLDFQSLYPS++IAYN C+ T LG++V + TN +G + +
Sbjct: 1157 CLPLVMEPQSAFYNSPVLVLDFQSLYPSVMIAYNYCYSTFLGRIVDWRGTNKMGFAEYRR 1216
Query: 1528 EQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQ 1587
+ +L+ L++ I + PNGVM+ +IR+ +L ++L EIL TR+MVK MK+ ++ LQ
Sbjct: 1217 RKRLLELLQEHINIAPNGVMYTKPEIRKSLLAKMLTEILETRVMVKSGMKQ-DKDDKTLQ 1275
Query: 1588 RIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWN 1647
++ N RQLALKL++NVTYGYT+A FSGR+PC+E+ADSIVQ GR TLE+AI+F++ +W
Sbjct: 1276 QLLNNRQLALKLLANVTYGYTSASFSGRLPCSEIADSIVQTGRETLERAIAFIHSVPRWG 1335
Query: 1648 AKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKR 1707
A+V+YGDTDS+F+ LKGRT ++AF+IG+E+A AIT MNP P+ LK EKVY PC L+ KKR
Sbjct: 1336 AEVVYGDTDSLFIHLKGRTKEQAFEIGNEMAKAITDMNPRPMKLKFEKVYLPCVLLAKKR 1395
Query: 1708 YVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQW 1767
YVGY YE+ +Q P FDAKGIETVRRD A KI E++LRL FE L ++K Y QRQ
Sbjct: 1396 YVGYKYEHVDQQVPDFDAKGIETVRRDGTPAEQKIEEKALRLLFETADLSQIKEYFQRQC 1455
Query: 1768 KRILSGRVSLKDFIFAKEVRLGTYSAR--ISSLPPAAIVATKAMRVDRRAEPRYAERIPY 1825
+I+ G+VS++DF FAKEV+LGTYS R + PP A+++TK M D RAEP+Y ER+ Y
Sbjct: 1456 NKIMRGQVSIQDFCFAKEVKLGTYSDRNGVGPAPPGALISTKKMLADPRAEPQYGERVLY 1515
Query: 1826 VVIHGEPGARLVDMVVDPLEVLA-IDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWF 1884
VV+ G PGARL D V P ++L+ + R++ YYI+K +IP L+R+F LVGA + W+
Sbjct: 1516 VVVTGAPGARLADRCVPPEDMLSPAGAHLRLDADYYISKNLIPPLERIFNLVGAHIRGWY 1575
Query: 1885 AEMPR 1889
E+P+
Sbjct: 1576 DELPK 1580
Score = 161 bits (407), Expect = 2e-37
Identities = 107/343 (31%), Positives = 177/343 (51%), Gaps = 50/343 (14%)
Query: 9 DVFSVRIVSVDHYMASPI---PSI--DISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHI 63
DVF VR+ +DHY A+P P + D+ HS K +VPVIRV+GST GQK C HI
Sbjct: 2 DVFKVRLNCIDHYQATPTQYDPCLRNDVRHSQLF--KEPKVPVIRVFGSTQTGQKVCAHI 59
Query: 64 HGALPYLYVSCSDIPLQLDQEG-DAYTYMVASSLEKALKLKGSADS----SRQHVHGCSL 118
HGA PYL++ + +LDQE A++Y + S++ AL + D+ + ++V +L
Sbjct: 60 HGAFPYLFIEYNG---KLDQEEVGAFSYRLHLSIDHALAVSYRQDAYARDTPKYVARITL 116
Query: 119 VRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLV 178
V+ FYG+ ++++KIY+ P ++R A+LL G ++ + QP+E+H+ +ILQF+
Sbjct: 117 VKGVPFYGFHVGYKYYLKIYM-LNPVVMTRLADLLHQGVIMKRKFQPYEAHLQYILQFMT 175
Query: 179 DYNLYGMGHLHLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISFDW 238
DYNLYG G++ S ++FR P+P + D ++ + IW + +IS
Sbjct: 176 DYNLYGCGYIEASSVKFRAPVP-------------QIDDDDMDSVMTPHIWHNQSIS--- 219
Query: 239 MWSLPSESGGSSNDNTHSPKRQSICELEGDISVDEILNQQ--------FKMFSSLSQTSS 290
+ HS R S C +E DI V +ILN++ L+ +
Sbjct: 220 ---------QKQITDHHSLPRASHCPIEVDICVQDILNRKDVKERRLHHDFVERLNPLPT 270
Query: 291 NVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLS 333
++ +V S+ +W ++ KR + + P P + + +S
Sbjct: 271 DMKLVASMAGLWRDETKRRK-RQLGISDPKSSPFPAEALVSMS 312
>UniRef100_Q9GSR1 DNA polymerase zeta catalytic subunit [Drosophila melanogaster]
Length = 2130
Score = 612 bits (1578), Expect = e-173
Identities = 362/863 (41%), Positives = 504/863 (57%), Gaps = 60/863 (6%)
Query: 1160 LALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEVLVLLHSKFVPCQRSLDGL 1219
L ++++EV +RGDL PDP D + + + E L ++ D L
Sbjct: 1300 LTIITLEVFVSTRGDLQPDPMHDEIRCLFYAIEHSLPD--EKLPSKACGYIMVNTVQDLL 1357
Query: 1220 S---------DCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLG 1270
S D +V T E + + D DI G+EI+ SS G++ +RA HL
Sbjct: 1358 SEGPFHGIDRDIEVQVVTSEAEAFEALLALCERWDADIYAGYEIEMSSWGYVIDRAKHLC 1417
Query: 1271 FGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASG 1330
F + LSR P+ + ++ D T LD
Sbjct: 1418 FNIAPLLSRVPTQK----------VRDFVDEDREQFTDLDV------------------E 1449
Query: 1331 VHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQ 1390
+ + GRI+L++WRL+R E+ L Y+ E V +L ++ P LT+WF G R+
Sbjct: 1450 MKLCGRILLDVWRLMRSEIALTSYTFENVMYHILHKRCPWHTAKSLTEWF--GSPCTRWI 1507
Query: 1391 CIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQN 1450
++Y ++R + L +L+QLD++ RTSE+A++ GI F+ VLSRGSQ+RVESM LR+A +N
Sbjct: 1508 VMEYYLERVRGTLTLLDQLDLLGRTSEMAKLIGIQFYEVLSRGSQFRVESMMLRIAKPKN 1567
Query: 1451 YVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLG 1510
V +SP Q A A E L L+MEP+S FY+DP++VLDFQSLYPSMIIAYN CF TCLG
Sbjct: 1568 LVPLSPSVQARAHMRAPEYLALIMEPQSRFYADPLIVLDFQSLYPSMIIAYNYCFSTCLG 1627
Query: 1511 KVVP---SKTNTLGVSPFVPEQHILQDLKDQILLT--PNGVMFVPSKIRRGVLPRLLEEI 1565
+V S G S + +LQ L + L+T P GV+FV ++R G+LPR+L EI
Sbjct: 1628 RVEHLGGSSPFEFGASQLRVSRQMLQKLLEHDLVTVSPCGVVFVKREVREGILPRMLTEI 1687
Query: 1566 LSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSI 1625
L TR MVKQ+MK L LQRI ++RQL LKL++NVTYGYTAA FSGRMP E+ DS+
Sbjct: 1688 LDTRQMVKQSMK-LHKDSSALQRILHSRQLGLKLMANVTYGYTAANFSGRMPSVEVGDSV 1746
Query: 1626 VQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMN 1685
V GR TLE+AI V +E+W +V+YGDTDSMFVL+ GR EAF+IG EIA A+T MN
Sbjct: 1747 VSKGRETLERAIKLVENNEEWKVRVVYGDTDSMFVLVPGRNRAEAFRIGEEIAKAVTEMN 1806
Query: 1686 PSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQ 1745
P PV LK+EKVY PC L TKKRYVGY YE +Q +PV++AKGIETVRRD C AVAK++E+
Sbjct: 1807 PQPVKLKLEKVYQPCMLQTKKRYVGYMYETADQEQPVYEAKGIETVRRDGCPAVAKMLEK 1866
Query: 1746 SLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVA 1805
LR+ FE Q + ++K Y+ RQ+ ++LSGR +L+D IFAKE R G + ++ PA +
Sbjct: 1867 VLRILFETQDVSKIKAYVCRQFTKLLSGRANLQDLIFAKEFR-GLNGYKPTACVPALELT 1925
Query: 1806 TKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQI 1865
K M+ D R PR ER+P+++++G PG +L+ +V P ++LA + +IN +YYI K I
Sbjct: 1926 RKWMQKDPRHVPRRGERVPFIIVNGPPGMQLIRLVRSPHDILA-NEGHKINAIYYITKAI 1984
Query: 1866 IPALQRVFGLVGADLNQWFAEMPRPV----REASVKHAFTPNFQRTRIDYYYLSKHCVL- 1920
IP L R L+GA+++ WFA +PR + + ++ I Y+ + CV+
Sbjct: 1985 IPPLNRCLLLIGANVHDWFASLPRKLLMTPAVGTANELAGARGAKSTISQYFSTTSCVID 2044
Query: 1921 CGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKC 1980
CG Q A +C C +N + KTA+LE+ Q ICQ C G RL ++C
Sbjct: 2045 CGR--QTKAGICPDCLKNATTCVVVLSDKTARLERGYQLTRQICQACCG--RL--GSLQC 2098
Query: 1981 TSISCLVFYERRKVQKELLAATH 2003
S+ C V Y ++EL H
Sbjct: 2099 DSLDCPVLYVLEGKRRELQQIEH 2121
Score = 163 bits (413), Expect = 4e-38
Identities = 86/202 (42%), Positives = 122/202 (59%), Gaps = 7/202 (3%)
Query: 5 ESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIH 64
E+ V+SVR+V D YM P +D +S ++ VPVIRV+G GQKTC+H+H
Sbjct: 6 EAIDGVYSVRLVIADFYMEKPQFGMDPCYSELRGKEIKRVPVIRVFGGNSRGQKTCMHVH 65
Query: 65 GALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVHGCSLVRAKK 123
G PYLY+ + + G +A L+KA+ + G S+ QHV LV+
Sbjct: 66 GVFPYLYIPYDKKDFESLERG---ILQMAMHLDKAINISLGQGSSNAQHVFKIQLVKGIP 122
Query: 124 FYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLY 183
FYGY E F+KIY+ + P+ V RAANLL +GA+L+K+ PHESH+P+ILQF++DYNLY
Sbjct: 123 FYGYHRVEHQFLKIYM-FNPRFVRRAANLLQSGAILSKNFSPHESHVPYILQFMIDYNLY 181
Query: 184 GMGHLH--LSKMRFRHPMPDSI 203
GM ++H L ++FR D +
Sbjct: 182 GMSYVHVPLEVLKFRRNHDDDV 203
>UniRef100_Q95TM2 LD40801p [Drosophila melanogaster]
Length = 1325
Score = 612 bits (1578), Expect = e-173
Identities = 362/863 (41%), Positives = 504/863 (57%), Gaps = 60/863 (6%)
Query: 1160 LALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEVLVLLHSKFVPCQRSLDGL 1219
L ++++EV +RGDL PDP D + + + E L ++ D L
Sbjct: 495 LTIITLEVFVSTRGDLQPDPMHDEIRCLFYAIEHSLPD--EKLPSKACGYIMVNTVQDLL 552
Query: 1220 S---------DCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLG 1270
S D +V T E + + D DI G+EI+ SS G++ +RA HL
Sbjct: 553 SEGPFHGIDRDIEVQVVTSEAEAFEALLALCERWDADIYAGYEIEMSSWGYVIDRAKHLC 612
Query: 1271 FGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASG 1330
F + LSR P+ + ++ D T LD
Sbjct: 613 FNIAPLLSRVPTQK----------VRDFVDEDREQFTDLDV------------------E 644
Query: 1331 VHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQ 1390
+ + GRI+L++WRL+R E+ L Y+ E V +L ++ P LT+WF G R+
Sbjct: 645 MKLCGRILLDVWRLMRSEIALTSYTFENVMYHILHKRCPWHTAKSLTEWF--GSPCTRWI 702
Query: 1391 CIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQN 1450
++Y ++R + L +L+QLD++ RTSE+A++ GI F+ VLSRGSQ+RVESM LR+A +N
Sbjct: 703 VMEYYLERVRGTLTLLDQLDLLGRTSEMAKLIGIQFYEVLSRGSQFRVESMMLRIAKPKN 762
Query: 1451 YVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLG 1510
V +SP Q A A E L L+MEP+S FY+DP++VLDFQSLYPSMIIAYN CF TCLG
Sbjct: 763 LVPLSPSVQARAHMRAPEYLALIMEPQSRFYADPLIVLDFQSLYPSMIIAYNYCFSTCLG 822
Query: 1511 KVVP---SKTNTLGVSPFVPEQHILQDLKDQILLT--PNGVMFVPSKIRRGVLPRLLEEI 1565
+V S G S + +LQ L + L+T P GV+FV ++R G+LPR+L EI
Sbjct: 823 RVEHLGGSSPFEFGASQLRVSRQMLQKLLEHDLVTVSPCGVVFVKREVREGILPRMLTEI 882
Query: 1566 LSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSI 1625
L TR MVKQ+MK L LQRI ++RQL LKL++NVTYGYTAA FSGRMP E+ DS+
Sbjct: 883 LDTRQMVKQSMK-LHKDSSALQRILHSRQLGLKLMANVTYGYTAANFSGRMPSVEVGDSV 941
Query: 1626 VQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMN 1685
V GR TLE+AI V +E+W +V+YGDTDSMFVL+ GR EAF+IG EIA A+T MN
Sbjct: 942 VSKGRETLERAIKLVENNEEWKVRVVYGDTDSMFVLVPGRNRAEAFRIGEEIAKAVTEMN 1001
Query: 1686 PSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQ 1745
P PV LK+EKVY PC L TKKRYVGY YE +Q +PV++AKGIETVRRD C AVAK++E+
Sbjct: 1002 PQPVKLKLEKVYQPCMLQTKKRYVGYMYETADQEQPVYEAKGIETVRRDGCPAVAKMLEK 1061
Query: 1746 SLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVA 1805
LR+ FE Q + ++K Y+ RQ+ ++LSGR +L+D IFAKE R G + ++ PA +
Sbjct: 1062 VLRILFETQDVSKIKAYVCRQFTKLLSGRANLQDLIFAKEFR-GLNGYKPTACVPALELT 1120
Query: 1806 TKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQI 1865
K M+ D R PR ER+P+++++G PG +L+ +V P ++LA + +IN +YYI K I
Sbjct: 1121 RKWMQKDPRHVPRRGERVPFIIVNGPPGMQLIRLVRSPHDILA-NEGHKINAIYYITKAI 1179
Query: 1866 IPALQRVFGLVGADLNQWFAEMPRPV----REASVKHAFTPNFQRTRIDYYYLSKHCVL- 1920
IP L R L+GA+++ WFA +PR + + ++ I Y+ + CV+
Sbjct: 1180 IPPLNRCLLLIGANVHDWFASLPRKLLMTPAVGTANELAGARGAKSTISQYFSTTSCVID 1239
Query: 1921 CGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKC 1980
CG Q A +C C +N + KTA+LE+ Q ICQ C G RL ++C
Sbjct: 1240 CGR--QTKAGICPDCLKNATTCVVVLSDKTARLERGYQLTRQICQACCG--RL--GSLQC 1293
Query: 1981 TSISCLVFYERRKVQKELLAATH 2003
S+ C V Y ++EL H
Sbjct: 1294 DSLDCPVLYVLEGKRRELQQIEH 1316
>UniRef100_UPI0000243912 UPI0000243912 UniRef100 entry
Length = 2070
Score = 612 bits (1577), Expect = e-173
Identities = 457/1390 (32%), Positives = 697/1390 (49%), Gaps = 162/1390 (11%)
Query: 703 ETMQTDEVEKELQKNSDHEVRSRDNLVCRKQS---FPSDSDSFLNISKDECLVQHERHYL 759
E + D+ ++ +S + S +N V K + D +S ++++ + + +
Sbjct: 733 EMERIDDDASDIPASSLPDESSHENAVPDKHNGVTITIDDNSDISLNSESASKPPKEKSI 792
Query: 760 EAGMVLKNSANQSSSSMHERPVLLETSHLADSIDK-SVACREKNTKDRTTYEKHAASDAY 818
E+ ++ N+ P LL+ L+D K + A E +K +A+ A
Sbjct: 793 ESEPIISIPDNEEDDFRMITPSLLDAIPLSDLFQKDATAANETINISDDDSDKQSAAPAV 852
Query: 819 TPNPSLDTHLRTDDDHKVRAPERCQETDSVASGSRQNSLVDDEVSG---KSKC------- 868
T LD + D+ + E+ +T A + ++ D+E +S C
Sbjct: 853 TIESQLDGVVELMDEVEDAVGEK--QTHDTAEATNAEAVEDEEEDNANIQSFCEKTLLCE 910
Query: 869 IDKTSSGS----IFFVQHDQMKSYEH----AVGKSAASDTRVLLTDKVDNQKLDKNLLCK 920
+D+ +S S + D + E+ V +A + TR +D ++ +
Sbjct: 911 LDEINSDSDDSCVGAWNKDSTEKEENEQPVVVSLAAGAPTRAEAQQAIDKFEIPAVINPT 970
Query: 921 TVGSEPIVDDLKSNHMKLTKVVIGNSSLVD-KNLKSHLSLPTFPNNLHLDEDDEMPG-NA 978
S+P D + T + +G +SL D + S ++ +L D+ G N
Sbjct: 971 PFYSDP-ADVTGKKEVGHTVLHVGGNSLNDVEEFNSTIAGLGSLKHLRYDKLQRTFGENI 1029
Query: 979 LDVFLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLLTPNILPPSAG 1038
DV P+ A + E S G+ + TP+ LPPS
Sbjct: 1030 NDVLGPVGANGPSSDRMK-------ELLASEGSVAI--------------TPSQLPPSRT 1068
Query: 1039 SVQRWLF--CDERDQEPDAEDQDV----------------------PKCTSGPLRHTPDQ 1074
W+ C +R + P D + PK S +
Sbjct: 1069 EASEWIDNRCRKRSEGPADSDSPIKVKPVDTLMTVDDGATPARLPAPKIDSESSLNLSVL 1128
Query: 1075 MRQEPGAKDKDISKCASGPTL--RPELHQDTEKKLPCINEGQTERIKAHMDHSQD----I 1128
+ + A S + T+ R + QD + + E A + ++ +
Sbjct: 1129 VANDGNAAPSSSSTDKTNATMQERTKPSQDCDAIESSMMESTEAAPNAATNEVKETIAPV 1188
Query: 1129 SQISGPGEK---------SSFTPLSQ-IGF-------QDPASAGHGQQLALLSIEVLAES 1171
S I+ + SS TP GF QD S L ++S+E+ +
Sbjct: 1189 SSIASSNQSDLEQSNANISSATPSDNTFGFKVDYENLQDAKSKCEYNYLTVMSVEIHVNT 1248
Query: 1172 RGDLLPDPQFDAVNIVALGFQKDCD------ATVEVLVLLHS------KFVPCQRSLDGL 1219
RGDL P+P D + + D ++V ++L K+ C D
Sbjct: 1249 RGDLRPNPATDPIAAIFYRIHNDVPPNHPRASSVCGIILNRDHEGRDFKYNQCPYVAD-- 1306
Query: 1220 SDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSR 1279
V+ +DE L ++F ++S DPDI G+EI+ S G++ ER L L+ LSR
Sbjct: 1307 ----VVLVSDESELYEKFLLLISFWDPDIFTGYEIESVSWGYVIERGYALEMNLMKRLSR 1362
Query: 1280 TPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVL 1339
PS ++ + SE+ RE + D +++G+ + GRI+L
Sbjct: 1363 VPS-----AETVHVSEEE---------------QRELLEMHD-----YSAGLKIPGRILL 1397
Query: 1340 NLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRA 1399
++WRL+R E+ L Y+ E V +L R+VP+ ++ L++ +S+ +R+ ++Y ++R
Sbjct: 1398 DIWRLMRHEIALTSYTFENVVYHILHRRVPNHSYRQLSRLWSKP--YSRWIVLEYYLERV 1455
Query: 1400 KLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQ 1459
N EILNQLD++ RT+ELA++FGI F+ VLSRGSQ+RVESM +R+A +N+V++SP
Sbjct: 1456 NGNFEILNQLDLIGRTAELAKLFGIQFYEVLSRGSQFRVESMMMRIAKPRNFVSVSPSIN 1515
Query: 1460 QVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSKTNT 1519
Q A A E LPL++EP S FY+DP++VLDFQSLYPS+IIAYN CF TCLG++ +T
Sbjct: 1516 QRAHMRAPEYLPLILEPNSRFYADPIIVLDFQSLYPSVIIAYNYCFSTCLGRIEHLGKDT 1575
Query: 1520 ----LGVSPFVPEQHILQDLKDQILLT--PNGVMFVPSKIRRGVLPRLLEEILSTRIMVK 1573
G S +L+ L D+ L+T P GV FV ++R GVLPR+L EIL+TR+MVK
Sbjct: 1576 ESFEFGASRLRVPPKMLKALLDKNLVTFSPCGVAFVKKRVREGVLPRMLSEILNTRLMVK 1635
Query: 1574 QAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTL 1633
++MK L VLQR+ ++RQL LKLI+NVTYGYTAA FSGRMPC E+ DS+V GR TL
Sbjct: 1636 KSMK-LHKDNSVLQRVLHSRQLGLKLIANVTYGYTAANFSGRMPCVEIGDSVVAKGRETL 1694
Query: 1634 EKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKM 1693
E+AI V E+W AKV+YGDTDS+FVL GR+ + AF+IG EIA A+T NP PV LK+
Sbjct: 1695 ERAIKLVEGTERWGAKVVYGDTDSLFVLCPGRSREAAFKIGEEIAEAVTNANPPPVKLKL 1754
Query: 1694 EKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEH 1753
EKVY P L TKKRYVGY YE+ +Q +P+++AKGIETVRRD C VAK++E+ LR+ FE
Sbjct: 1755 EKVYQPAILQTKKRYVGYMYESADQEKPIYEAKGIETVRRDGCPVVAKMLEKVLRILFET 1814
Query: 1754 QSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDR 1813
+ +VK Y RQ+ +IL GRV+L+DFIFAKE R G + + PA + K D
Sbjct: 1815 CDVSKVKQYTCRQFTKILEGRVNLQDFIFAKEFR-GESGYKPGACVPALELMRKWKLTDP 1873
Query: 1814 RAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVF 1873
R EPR +R+PYV+I+G P L+ +V P E+L+ + +IN YYI+K IIP L R
Sbjct: 1874 RREPRQGQRVPYVIINGPPLVPLIRLVRSPDELLS-NEGLKINANYYISKAIIPPLNRCL 1932
Query: 1874 GLVGADLNQWFAEMPRPV-----REASVKHAFTPNFQRTRIDYYYLSKHCVL-CGGLVQA 1927
L+GAD +QW++E+PR + P+ + T I Y+ + +C++ CG Q
Sbjct: 1933 LLIGADAHQWYSELPRKTLMLHNTGGGGGGSGGPSVKIT-ISQYFSTTNCIVDCG--KQT 1989
Query: 1928 SARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLV 1987
+C C + A V+ K KLE+++ +C+ C R E+ C S+ C V
Sbjct: 1990 HHGVCKDCRQQPQRALVHVMNKVNKLERKLVLTEKMCRSC--CQRTFETA--CISLDCPV 2045
Query: 1988 FYERRKVQKE 1997
+ + + E
Sbjct: 2046 VFSLNRRELE 2055
Score = 145 bits (367), Expect = 1e-32
Identities = 82/194 (42%), Positives = 111/194 (56%), Gaps = 11/194 (5%)
Query: 13 VRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPYLYV 72
+RIVSVDHYM+ P P D +S F +V +VPVIR++GS G +C+HIHG PY YV
Sbjct: 1 IRIVSVDHYMSKPDPQFDTCYSEFRGAEVKQVPVIRLFGSNAEGTHSCVHIHGVFPYFYV 60
Query: 73 ----SCSDIPLQLDQEGDAYTYMVASSLEKALK-LKGSADSSRQHVHGCSLVRAKKFYGY 127
S +D L +D++ Y +A +L+K + + G A S +H+ LV+ YGY
Sbjct: 61 PYDGSVAD-RLAVDKK----IYQLAGALDKGINVMLGRASSQAKHIFKIVLVKGIPIYGY 115
Query: 128 CSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGH 187
+ KIY+ Y P V A LL+ G +L+ S Q HE+HIP+ILQF +DYNLYGM
Sbjct: 116 HRNPHHYFKIYM-YNPYLVRNATQLLMNGTILSGSYQVHETHIPYILQFFIDYNLYGMSF 174
Query: 188 LHLSKMRFRHPMPD 201
L L R D
Sbjct: 175 LELDSKGLRQRTGD 188
>UniRef100_Q9GSR2 DNA polymerase zeta catalytic subunit [Drosophila melanogaster]
Length = 2130
Score = 612 bits (1577), Expect = e-173
Identities = 386/992 (38%), Positives = 546/992 (54%), Gaps = 81/992 (8%)
Query: 1052 EPDAEDQDVPKCTSGPLRHTPDQMRQEPG---AKDKDISKCASGPTLRPELHQDTEKKLP 1108
+ DA D+ + PL D+ + P A+++ S+ G L QD E
Sbjct: 1171 DEDAGDELDCSLSLTPLSQAKDKCKATPTSSKARERGKSRLKPGTRLSFIGSQDEEPPSS 1230
Query: 1109 CINEGQ--TERIKAHMDHSQDISQISGPGEK---------SSFTPLSQIGF-------QD 1150
+E + +A +D S + Q+ G + S T + GF Q
Sbjct: 1231 QSSEQSVFSSAAQAELDRSSFLRQLEGSSQDRQHDLSFGLSHATLDNTFGFKVNLENLQQ 1290
Query: 1151 PASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEVLVLLHSKFV 1210
+ L ++++EV +RGDL PDP D + + + E L ++
Sbjct: 1291 AKADIDCNHLTIITLEVFVSTRGDLQPDPMHDEIRCLFYAIEHSLPD--EKLPSKACGYI 1348
Query: 1211 PCQRSLDGLS---------DCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGF 1261
D LS D +V T E + + D DI G+EI+ SS G+
Sbjct: 1349 MVNTVQDLLSEGPFHGIDRDIEVQVVTSEAEAFEALLALCERWDADIYAGYEIEMSSWGY 1408
Query: 1262 LAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIED 1321
+ +RA HL F + LSR P+ + ++ D T LD
Sbjct: 1409 VIDRAKHLCFNIAPLLSRVPTQK----------VRDFVDEDREQFTDLDV---------- 1448
Query: 1322 EWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFS 1381
+ + GRI+L++WRL+R E+ L Y+ E V +L ++ P LT+WF
Sbjct: 1449 --------EMKLCGRILLDVWRLMRSEIALTSYTFENVMYHILHKRCPWHTAKSLTEWF- 1499
Query: 1382 RGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESM 1441
G R+ ++Y ++R + L +L QLD++ RTSE+A++ GI F+ VLSRGSQ+RVESM
Sbjct: 1500 -GSPCTRWIVMEYYLERVRGTLTLLEQLDLLGRTSEMAKLIGIQFYEVLSRGSQFRVESM 1558
Query: 1442 FLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAY 1501
LR+A +N V +SP Q A A E L L+MEP+S FY+DP++VLDFQSL+PSMIIAY
Sbjct: 1559 MLRIAKPKNLVPLSPSVQARAHMRAPEYLALIMEPQSRFYADPLIVLDFQSLHPSMIIAY 1618
Query: 1502 NLCFCTCLGKVVP---SKTNTLGVSPFVPEQHILQDLKDQILLT--PNGVMFVPSKIRRG 1556
N CF TCLG+V S G S + +LQ L + L+T P GV+FV ++R G
Sbjct: 1619 NYCFSTCLGRVEHLGGSSPFEFGASQLRVSRQMLQKLLEHDLVTVSPCGVVFVKREVREG 1678
Query: 1557 VLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRM 1616
+LPR+L EIL TR MVKQ+MK L LQRI ++RQL LKL++NVTYGYTAA FSGRM
Sbjct: 1679 ILPRMLTEILDTRQMVKQSMK-LHKDSSALQRILHSRQLGLKLMANVTYGYTAANFSGRM 1737
Query: 1617 PCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSE 1676
P E+ DS+V GR TLE+AI V +E+W +V+YGDTDSMFVL+ GR EAF+IG E
Sbjct: 1738 PSVEVGDSVVSKGRETLERAIKLVENNEEWKVRVVYGDTDSMFVLVPGRNRAEAFRIGEE 1797
Query: 1677 IASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTC 1736
IA A+T MNP PV LK+EKVY PC L TKKRYVGY YE +Q +PV++AKGIET RRD C
Sbjct: 1798 IAKAVTEMNPQPVKLKLEKVYQPCMLQTKKRYVGYMYETADQEQPVYEAKGIETERRDGC 1857
Query: 1737 EAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARIS 1796
AVAK++E+ LR+ FE Q + ++K Y+ RQ+ ++LSGR +L+D IFAKE R G + +
Sbjct: 1858 PAVAKMLEKVLRILFETQDVSKIKAYVCRQFTKLLSGRANLQDLIFAKEFR-GLNGYKPT 1916
Query: 1797 SLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRIN 1856
+ PA + K M+ D R PR ER+P+++++G PG +L+ +V P ++LA + +IN
Sbjct: 1917 ACVPALELTRKWMQKDPRHVPRRGERVPFIIVNGPPGMQLIRLVRSPHDILA-NEGHKIN 1975
Query: 1857 DLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPV----REASVKHAFTPNFQRTRIDYY 1912
+YYI K IIP L R L+GA+++ WFA +PR + + P ++ I Y
Sbjct: 1976 AIYYITKAIIPPLNRCLLLIGANVHDWFASLPRKLLMTPAVGTANELAGPRGAKSTISQY 2035
Query: 1913 YLSKHCVL-CGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGD 1971
+ + CV+ CG Q A +C C +N + KTA+LE+ Q ICQ C G
Sbjct: 2036 FSTTSCVIDCGR--QTKAGICPDCLKNATTCVVVLSDKTARLERGYQLTRQICQACCG-- 2091
Query: 1972 RLLESGVKCTSISCLVFYERRKVQKELLAATH 2003
RL ++C S+ C V Y ++EL H
Sbjct: 2092 RL--GSLQCDSLDCPVLYVLEGKRRELQQIEH 2121
Score = 163 bits (412), Expect = 6e-38
Identities = 86/202 (42%), Positives = 122/202 (59%), Gaps = 7/202 (3%)
Query: 5 ESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIH 64
E+ V+SVR+V D YM P +D +S ++ VPVIRV+G GQKTC+H+H
Sbjct: 6 EAIDGVYSVRLVIADFYMEKPQFGMDPCYSELRGKEIKRVPVIRVFGGNSRGQKTCMHVH 65
Query: 65 GALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLK-GSADSSRQHVHGCSLVRAKK 123
G PYLY+ + + G +A L+KA+ + G S+ QHV LV+
Sbjct: 66 GVFPYLYIPYDKKDFESLERG---ILQMAVHLDKAINISLGQGSSNAQHVFKIQLVKGIP 122
Query: 124 FYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLY 183
FYGY E F+KIY+ + P+ V RAANLL +GA+L+K+ PHESH+P+ILQF++DYNLY
Sbjct: 123 FYGYHRVEHQFLKIYM-FNPRFVRRAANLLQSGAILSKNFSPHESHVPYILQFMIDYNLY 181
Query: 184 GMGHLH--LSKMRFRHPMPDSI 203
GM ++H L ++FR D +
Sbjct: 182 GMSYVHVPLEVLKFRRNHDDDV 203
>UniRef100_Q6CB61 Similar to sp|P14284 Saccharomyces cerevisiae DNA polymerase zeta
catalytic subunit [Yarrowia lipolytica]
Length = 1338
Score = 608 bits (1568), Expect = e-172
Identities = 363/873 (41%), Positives = 508/873 (57%), Gaps = 61/873 (6%)
Query: 1128 ISQISGPGEKSSFTPLSQIGFQDPASAGH-GQQLALLSIEVLAESRGDLLPDPQFDAVNI 1186
+SQI+ P +K ++ SQ + S H L ++S+EV +R L PDP D +
Sbjct: 499 LSQIAPPTQKPKYSFPSQ----NTLSQRHESAALTVMSMEVHVNTRKGLNPDPDLDPILF 554
Query: 1187 VALGFQKDCDATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDP 1246
V V+++++ P + + E ++ K+V + DP
Sbjct: 555 VVWHMLGGPSG-----VIINAEDCP---DFTNIIEAPSTVVKTECEMITHLQKMVENFDP 606
Query: 1247 DILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDT 1306
DIL G+E+Q SS G++ +R + + + R Q K S K
Sbjct: 607 DILTGFEVQASSWGYVQDRCKTMLYPC--EFGRV--------QHHKMSSKI--------- 647
Query: 1307 TSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRR 1366
D WG ASGV V GR VLNLWR+IRGEV L Y++E V VL
Sbjct: 648 --------------DTWGARKASGVKVVGRHVLNLWRMIRGEVSLLKYTLENVVFHVLHE 693
Query: 1367 KVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDF 1426
++P H+ LT+ G R ++Y R+ NL++L+ L++V+RT+E ARV GIDF
Sbjct: 694 RIPFYTHDTLTE-MCVGSLSDRKLLVEYKCTRSNYNLQLLSTLEIVSRTAEQARVVGIDF 752
Query: 1427 FSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVV 1486
SV +RGSQY+VES RL ++N++ SP +QV Q A+EC+ LVMEPES Y+ PV+
Sbjct: 753 SSVYTRGSQYKVESFLARLTKSENFMLASPSREQVGQQNALECIALVMEPESRLYTSPVL 812
Query: 1487 VLDFQSLYPSMIIAYNLCFCTCLGKVVPSKT--NTLGVSPFVPEQHILQDL--KDQILLT 1542
VLDFQSLYPS+I+A+N C+ TCLGK + NTLG + +++ L KD + ++
Sbjct: 813 VLDFQSLYPSVILAHNYCYSTCLGKWTDFEKDGNTLGTEKLFYKPSLVKRLLTKDDVTIS 872
Query: 1543 PNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISN 1602
PNG++FV IR +L R+L EIL TR MVK K L QR+ + RQLALKL++N
Sbjct: 873 PNGLVFVKPHIRVSLLARMLGEILETRFMVKDTAK-LDRDNVSFQRLNHNRQLALKLVAN 931
Query: 1603 VTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLL 1662
VTYGYT+A FSGRMPCAE+AD+IVQ GR TLEK I ++ +KW AKV+YGDTDS+FV L
Sbjct: 932 VTYGYTSASFSGRMPCAEIADAIVQSGRETLEKCIDVIHGSDKWAAKVVYGDTDSLFVCL 991
Query: 1663 KGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPV 1722
GRT EAF+IG+EIA IT+ NP+P+ L+ EKVY PC LI+KKRYVGY +E Q+ P+
Sbjct: 992 PGRTKDEAFKIGTEIADTITSANPAPMKLQFEKVYLPCMLISKKRYVGYKWEYSKQMYPI 1051
Query: 1723 FDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIF 1782
FDAKGIETVRRD A KI E++LR+ F+ L VK+YL QW +IL+G+VS++DF F
Sbjct: 1052 FDAKGIETVRRDGTPAAQKIEEKALRILFDTADLSLVKSYLYEQWTKILTGKVSIQDFCF 1111
Query: 1783 AKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVD 1842
AKEV+LG Y LP A ++ + M VD R EP+Y ERIPYVV+ G P +RL+D V
Sbjct: 1112 AKEVKLGQYKEG-GVLPAGAKISAEKMAVDMRFEPQYKERIPYVVVAGPPKSRLIDRCVS 1170
Query: 1843 PLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKHAFTP 1902
P E++ + ++ YYI+K +IP L R F VGA + +W+ EMP+ R +
Sbjct: 1171 PEELVRNANSLILDADYYIHKTLIPPLDRFFNCVGASVLKWYEEMPKKRRYEFYQTVSRA 1230
Query: 1903 NFQRTRIDYYYLSKHCVLC-GGLVQASARLCSQCSENEVAAATAVIGKTAKL-EQEMQHL 1960
+T + Y +S C++C ++LC+ C E++ AT V+ + E+ + L
Sbjct: 1231 KTNQTTLKSYAVSNSCIVCKTAQTSGLSKLCATC-ESDPGHATFVLNSDLRYREKRLDEL 1289
Query: 1961 VSICQHCGGGDRLLESGVKCTSISCLVFYERRK 1993
+IC C G + +C S C ++Y R K
Sbjct: 1290 NTICAQCSG-----DFETRCESQDCPIYYSRVK 1317
Score = 75.9 bits (185), Expect = 1e-11
Identities = 73/312 (23%), Positives = 122/312 (38%), Gaps = 36/312 (11%)
Query: 13 VRIVSVDHYMASPIP----------SIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLH 62
++I +D+ MA P P S + + HR VPV+R++G T G +C H
Sbjct: 6 IQINYIDYVMAPPGPLDWSKVPETVSCECDYKDLHR-----VPVLRIFGGTSDGLSSCAH 60
Query: 63 IHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCS----- 117
+H Y Y+ + L D+ ++ L+L+ + CS
Sbjct: 61 VHNVFQYFYIPYTGPSL---SPADSEPFIADMYRRINLQLRSKRTRGEKLPEECSFLANI 117
Query: 118 -LVRAKKFYGYCSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQF 176
L +A FYGY +++KI + P V ++ G + ESH+ +ILQF
Sbjct: 118 VLCKATPFYGYHEGWRYYLKIVV-VDPSHVGMLVDMFRNGVFGWDHSRVFESHLSYILQF 176
Query: 177 LVDYNLYGMGHLHLSKMRFRHPMPDSIHKKSDINSQHRKADPGAEACLESKIWMSSTISF 236
DYNL+G G + S+ R ++ I+ + D AE L + ++
Sbjct: 177 FCDYNLHGCGWMEASRYMMR--------REGMISKSEFEVDLQAEYILNRLVISTNKKPG 228
Query: 237 DWMWSLPSESGGSSNDNTHSPKRQSICELEGDISVDEILNQQF---KMFSSLSQTSSNVN 293
D + E S + G+ +V++ Q F MFS L + S +
Sbjct: 229 DKAFHSLKELWTSLQSIRDKFSKGEYKSDYGERTVNDPREQWFSAQSMFSELEEKVSKAD 288
Query: 294 MVQSLVSIWEEQ 305
S V W ++
Sbjct: 289 SAHSYVQFWPKR 300
>UniRef100_Q7PUB0 ENSANGP00000009889 [Anopheles gambiae str. PEST]
Length = 2126
Score = 602 bits (1551), Expect = e-170
Identities = 386/1042 (37%), Positives = 570/1042 (54%), Gaps = 118/1042 (11%)
Query: 1008 SSGTKGVSTYYQNDGSHLYLLTPNILPPSAGSVQRWLFCDER---DQEPDAEDQDVPKCT 1064
S + +S NDG N P S+ + + ER Q+ DA + + + T
Sbjct: 1136 SESSLNLSVLVANDG--------NAAPSSSSTDKTNATMQERTKPSQDCDAIESSMMEST 1187
Query: 1065 SGPLRHTPDQMRQEPGAKDKDISKCASGPTLRPELHQDTEKKLPCINEGQTERIKAHMDH 1124
++ + E A ++ I+ +S + N+ E+ A++
Sbjct: 1188 EAAPNAATNEEQAEKNAAEETIAPVSS---------------IASSNQSDLEQSNANISS 1232
Query: 1125 SQDISQISGPGEKSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAV 1184
+ P + + + QD S L ++S+E+ +RGDL P+P D +
Sbjct: 1233 AT-------PSDNTFGFKVDYENLQDAKSKCEYNYLTVMSVEIHVNTRGDLRPNPATDPI 1285
Query: 1185 NIVALGFQKDCD------ATVEVLVLLHSKFVPCQRSLDGLSDCKVLNF----------- 1227
+ D ++V ++L + G + K +F
Sbjct: 1286 AAIFYRIHNDVPPNHPRASSVCGIILNRDHEGREFERVQGQAGGKTASFKYNQCPYVADV 1345
Query: 1228 ---TDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRTPSNS 1284
+DE L ++F ++S DPDI G+EI+ S G++ ER L L+ LSR PS
Sbjct: 1346 VLVSDESELYEKFLLLISFWDPDIFTGYEIESVSWGYVIERGYALEMNLMKRLSRVPS-- 1403
Query: 1285 WINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRL 1344
++ + SE+ RE + D +++G+ + GRI+L++WRL
Sbjct: 1404 ---AETVHVSEEE---------------QRELLEMHD-----YSAGLKIPGRILLDIWRL 1440
Query: 1345 IRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAKLNLE 1404
+R E+ L Y+ E V +L R+VP+ ++ L++ +S+ +R+ ++Y ++R N E
Sbjct: 1441 MRHEIALTSYTFENVVYHILHRRVPNHSYRQLSRLWSKP--YSRWIVLEYYLERVNGNFE 1498
Query: 1405 ILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQVASQ 1464
ILNQLD++ RT+ELA++FGI F+ VLSRGSQ+RVESM +R+A +N+V++SP Q A
Sbjct: 1499 ILNQLDLIGRTAELAKLFGIQFYEVLSRGSQFRVESMMMRIAKPRNFVSVSPSINQRAHM 1558
Query: 1465 PAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSKTNT----L 1520
A E LPL++EP S FY+DP++VLDFQSLYPS+IIAYN CF TCLG++ +T
Sbjct: 1559 RAPEYLPLILEPNSRFYADPIIVLDFQSLYPSVIIAYNYCFSTCLGRIEHLGKDTESFEF 1618
Query: 1521 GVSPFVPEQHILQDLKDQILLT--PNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKK 1578
G S +L+ L D+ L+T P GV FV ++R GVLPR+L EIL+TR+MVK++MK
Sbjct: 1619 GASRLRVPPKMLKALLDKNLVTFSPCGVAFVKKRVREGVLPRMLSEILNTRLMVKKSMK- 1677
Query: 1579 LSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLEKAIS 1638
L VLQR+ ++RQL LKLI+NVTYGYTAA FSGRMPC E+ DS+V GR TLE+AI
Sbjct: 1678 LHKDNSVLQRVLHSRQLGLKLIANVTYGYTAANFSGRMPCVEIGDSVVAKGRETLERAIK 1737
Query: 1639 FVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSPVTLKMEKVYH 1698
V E+W AKV+YGDTDS+FVL GR+ + AF+IG EIA A+T NP PV LK+EKVY
Sbjct: 1738 LVEGTERWGAKVVYGDTDSLFVLCPGRSREAAFKIGEEIAEAVTNANPPPVKLKLEKVYQ 1797
Query: 1699 PCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLE 1758
P L TKKRYVGY YE+ +Q +P+++AKGIETVRRD C VAK++E+ LR+ FE + +
Sbjct: 1798 PAILQTKKRYVGYMYESADQEKPIYEAKGIETVRRDGCPVVAKMLEKVLRILFETCDVSK 1857
Query: 1759 VKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPR 1818
VK Y RQ+ +IL GRV+L+DFIFAKE R G + + PA + K D R EPR
Sbjct: 1858 VKQYTCRQFTKILEGRVNLQDFIFAKEFR-GESGYKPGACVPALELMRKWKLTDPRREPR 1916
Query: 1819 YAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGA 1878
+R+PYV+I+G P L+ +V P E+L+ + +IN YYI+K IIP L R L+GA
Sbjct: 1917 QGQRVPYVIINGPPLVPLIRLVRSPDELLS-NEGLKINANYYISKAIIPPLNRCLLLIGA 1975
Query: 1879 DLNQWFAEMPR------------------PVREASVKHAFTPNFQRTR----IDYYYLSK 1916
D +QW++E+PR V + A +R I Y+ +
Sbjct: 1976 DAHQWYSELPRKTLMLHNTGGGGGGSGGPSVVKMGAALATAGGDERLHKKITISQYFSTT 2035
Query: 1917 HCVL-CGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLLE 1975
+C++ CG Q +C C + A V+ K KLE+++ +C+ C R E
Sbjct: 2036 NCIVDCG--KQTHHGVCKDCRQQPQRALVHVMNKVNKLERKLVLTEKMCRSC--CQRTFE 2091
Query: 1976 SGVKCTSISCLVFYERRKVQKE 1997
+ C S+ C V + + + E
Sbjct: 2092 TA--CISLDCPVVFSLNRRELE 2111
Score = 145 bits (367), Expect = 1e-32
Identities = 82/194 (42%), Positives = 111/194 (56%), Gaps = 11/194 (5%)
Query: 13 VRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPYLYV 72
+RIVSVDHYM+ P P D +S F +V +VPVIR++GS G +C+HIHG PY YV
Sbjct: 1 IRIVSVDHYMSKPDPQFDTCYSEFRGAEVKQVPVIRLFGSNAEGTHSCVHIHGVFPYFYV 60
Query: 73 ----SCSDIPLQLDQEGDAYTYMVASSLEKALK-LKGSADSSRQHVHGCSLVRAKKFYGY 127
S +D L +D++ Y +A +L+K + + G A S +H+ LV+ YGY
Sbjct: 61 PYDGSVAD-RLAVDKK----IYQLAGALDKGINVMLGRASSQAKHIFKIVLVKGIPIYGY 115
Query: 128 CSFEEFFVKIYLRYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQFLVDYNLYGMGH 187
+ KIY+ Y P V A LL+ G +L+ S Q HE+HIP+ILQF +DYNLYGM
Sbjct: 116 HRNPHHYFKIYM-YNPYLVRNATQLLMNGTILSGSYQVHETHIPYILQFFIDYNLYGMSF 174
Query: 188 LHLSKMRFRHPMPD 201
L L R D
Sbjct: 175 LELDSKGLRQRTGD 188
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,316,897,671
Number of Sequences: 2790947
Number of extensions: 140612340
Number of successful extensions: 354853
Number of sequences better than 10.0: 973
Number of HSP's better than 10.0 without gapping: 491
Number of HSP's successfully gapped in prelim test: 484
Number of HSP's that attempted gapping in prelim test: 350520
Number of HSP's gapped (non-prelim): 2342
length of query: 2018
length of database: 848,049,833
effective HSP length: 143
effective length of query: 1875
effective length of database: 448,944,412
effective search space: 841770772500
effective search space used: 841770772500
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 83 (36.6 bits)
Medicago: description of AC146571.7


