

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.5 - phase: 0
(337 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
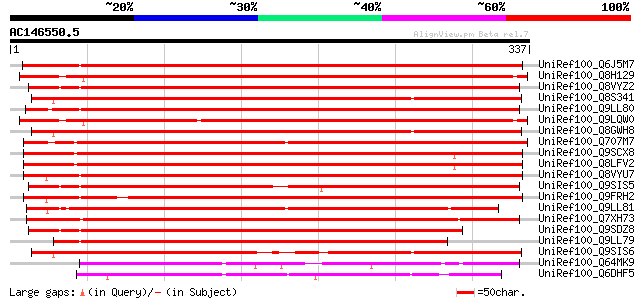
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6J5M7 Purple acid phosphatase 1 [Solanum tuberosum] 436 e-121
UniRef100_Q8H129 Putative purple acid phosphatase [Arabidopsis t... 411 e-113
UniRef100_Q8VYZ2 Purple acid phosphatase [Arabidopsis thaliana] 406 e-112
UniRef100_Q8S341 Purple acid phosphatase [Arabidopsis thaliana] 405 e-112
UniRef100_Q9LL80 Putative purple acid phosphatase [Glycine max] 405 e-111
UniRef100_Q9LQW0 F10B6.10 [Arabidopsis thaliana] 403 e-111
UniRef100_Q8GWH8 Putative purple acid phosphatase [Arabidopsis t... 401 e-110
UniRef100_Q707M7 Putative acid phosphatase [Lupinus luteus] 399 e-110
UniRef100_Q9SCX8 Acid phosphatase type 5 precursor [Arabidopsis ... 390 e-107
UniRef100_Q8LFV2 Acid phosphatase type 5 [Arabidopsis thaliana] 389 e-107
UniRef100_Q8VYU7 At1g25230/F4F7_8 [Arabidopsis thaliana] 389 e-107
UniRef100_Q9SIS5 Putative purple acid phosphatase [Arabidopsis t... 375 e-103
UniRef100_Q9FRH2 Hypothetical protein F4F7.38 [Arabidopsis thali... 375 e-103
UniRef100_Q9LL81 Putative purple acid phosphatase [Ipomoea batatas] 367 e-100
UniRef100_Q7XH73 Putative purple acid phosphatase [Oryza sativa] 364 2e-99
UniRef100_Q9SDZ8 Putative purple acid phosphatase precursor [Ara... 358 1e-97
UniRef100_Q9LL79 Putative purple acid phosphatase [Phaseolus vul... 354 2e-96
UniRef100_Q9SIS6 Putative purple acid phosphatase [Arabidopsis t... 332 9e-90
UniRef100_Q64MK9 Acid phosphatase [Bacteroides fragilis] 135 1e-30
UniRef100_Q6DHF5 Zgc:92339 [Brachydanio rerio] 135 2e-30
>UniRef100_Q6J5M7 Purple acid phosphatase 1 [Solanum tuberosum]
Length = 328
Score = 436 bits (1120), Expect = e-121
Identities = 201/325 (61%), Positives = 255/325 (77%), Gaps = 1/325 (0%)
Query: 9 MLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVA 68
M S I+ + + LL + +L R EH + S++FLVVGDWGR+G +NQS VA
Sbjct: 1 MASMKILNIFISFLLLLLFPAAMAELHRLEHPVNTDG-SISFLVVGDWGRRGTFNQSQVA 59
Query: 69 HQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHD 128
QMGI+G+ LNI+FVVSTGDNFYDDGL GVDDPAF ESF ++YTAPSLQ+ WYNVLGNHD
Sbjct: 60 QQMGIIGEKLNIDFVVSTGDNFYDDGLTGVDDPAFEESFTNVYTAPSLQKNWYNVLGNHD 119
Query: 129 YRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKG 188
YRGD AQLSPIL+QKDNRW+C+RS+I++TD EFFFVDTTPF + YFT P HTYDW+
Sbjct: 120 YRGDALAQLSPILKQKDNRWICMRSYIVNTDVAEFFFVDTTPFQDMYFTTPKDHTYDWRN 179
Query: 189 VLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNV 248
V+P + Y +++LK++DSAL +S+AKWKIVV HH IKSAG HG+++EL +LPIL++NNV
Sbjct: 180 VMPRKDYLSQVLKDLDSALRESSAKWKIVVGHHTIKSAGHHGSSEELGVHILPILQANNV 239
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQIS 308
D Y+NGHDHCLEHI +S + F TSGGGSK+W GD+ W+PKE+K Y+DGQGF++MQI+
Sbjct: 240 DFYLNGHDHCLEHISSSDSPLQFLTSGGGSKSWRGDMNWWNPKEMKFYYDGQGFMAMQIT 299
Query: 309 KTNASIVFYDVFGKVLHTWSMSKKL 333
+T I F+D+FG +LH WS SK L
Sbjct: 300 QTQVWIQFFDIFGNILHKWSASKNL 324
>UniRef100_Q8H129 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 366
Score = 411 bits (1057), Expect = e-113
Identities = 197/333 (59%), Positives = 252/333 (75%), Gaps = 9/333 (2%)
Query: 7 QHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQ 64
+H +++ V L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQ
Sbjct: 35 KHKPVNLVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQ 90
Query: 65 SLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVL 124
S VA QMG +G+ L+I+FV+STGDNFYD+GL + DP F +SF +IYTAPSLQ+ WY+VL
Sbjct: 91 SQVALQMGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYSVL 150
Query: 125 GNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTY 184
GNHDYRGDV AQLSP+LR DNRWVC+RSFI++ + V+ FFVDTTPFV++YF P KH Y
Sbjct: 151 GNHDYRGDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVY 210
Query: 185 DWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILK 244
DW GVLP ++Y LLKE+D AL +S AKWKIV+ HH IKSAG HGNT ELE+ LLPIL+
Sbjct: 211 DWSGVLPRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQ 270
Query: 245 SNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAW-GGDIKPWDPKELKLYHDGQGFV 303
+N VD Y+NGHDHCLEHI +S I F TSGGGSKAW GGD+ +P+E++ Y+DGQGF+
Sbjct: 271 ANEVDLYVNGHDHCLEHISSVDSNIQFMTSGGGSKAWKGGDVNYVEPEEMRFYYDGQGFM 330
Query: 304 SMQISKTNASIVFYDVFGKVLHTWSMSKKLKEA 336
S+ +S+ +VFYDVFG VLH W K KEA
Sbjct: 331 SVHVSEAELRVVFYDVFGHVLHHW--KKTYKEA 361
>UniRef100_Q8VYZ2 Purple acid phosphatase [Arabidopsis thaliana]
Length = 335
Score = 406 bits (1044), Expect = e-112
Identities = 196/319 (61%), Positives = 241/319 (75%), Gaps = 2/319 (0%)
Query: 13 VIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+I ++ L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGD 132
+GK+LNI+F++STGDNFYDDG+ D F +SF +IYTA SLQ+ WYNVLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPL 192
V AQLSPILR D RW+CLRS++++ + V+ FFVDTTPFV+ YF +P H YDW+GVLP
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYI 252
Y LL +VD AL +S AKWKIVV HH IKSAG HGNT ELE+QLLPIL++N VD YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNA 312
NGHDHCLEHI SGI F TSGGGSKAW GD+ W+P+E++ Y+DGQGF+S+ S+
Sbjct: 250 NGHDHCLEHISSINSGIQFMTSGGGSKAWKGDVNDWNPQEMRFYYDGQGFMSVYTSEAEL 309
Query: 313 SIVFYDVFGKVLHTWSMSK 331
+VFYD G VLH WS K
Sbjct: 310 RVVFYDGLGHVLHRWSTLK 328
>UniRef100_Q8S341 Purple acid phosphatase [Arabidopsis thaliana]
Length = 328
Score = 405 bits (1041), Expect = e-112
Identities = 194/321 (60%), Positives = 240/321 (74%), Gaps = 4/321 (1%)
Query: 15 IAVSVLLCLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQM 71
+ SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQM
Sbjct: 5 VCFSVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQM 64
Query: 72 GIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRG 131
G+VG+ L+I+FV+S GDNFYDDGLKGV+DP+F SF IYT PSLQ+ WY+VLGNHDYRG
Sbjct: 65 GVVGEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRG 124
Query: 132 DVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLP 191
+VEAQLS +L QKD RW C RSF+L + V+FFF DT PFVE+YFT+P HTYDW+ VLP
Sbjct: 125 NVEAQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLP 184
Query: 192 LESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAY 251
Y + LL ++D + +S A WK VV HH IK+AG HG TQEL +QLLPIL+ N VD Y
Sbjct: 185 RNKYISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLY 244
Query: 252 INGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTN 311
INGHDHCL+H I F TSGGGSKAW G ++PWDPKELKLY+DGQGF+S+ I+ +
Sbjct: 245 INGHDHCLQH-IGSHGKTQFLTSGGGSKAWRGHVQPWDPKELKLYYDGQGFMSLHITHSK 303
Query: 312 ASIVFYDVFGKVLHTWSMSKK 332
A ++YDV G VLH S+SK+
Sbjct: 304 AKFIYYDVSGNVLHRSSLSKR 324
>UniRef100_Q9LL80 Putative purple acid phosphatase [Glycine max]
Length = 332
Score = 405 bits (1040), Expect = e-111
Identities = 189/321 (58%), Positives = 241/321 (74%), Gaps = 3/321 (0%)
Query: 11 SSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQ 70
S I+A+ + C SS KL +H K SL+FLVVGDWGRKG YNQSLVA Q
Sbjct: 9 SCTIVAIFLAFCCFV--SSSKAKLESLQHAPKADG-SLSFLVVGDWGRKGAYNQSLVAFQ 65
Query: 71 MGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYR 130
MG++G+ L+++FV+STGDNFYD+GL GV DP+F ESF IYTAPSLQ+ WYNVLGNHDYR
Sbjct: 66 MGVIGEKLDVDFVISTGDNFYDNGLTGVFDPSFEESFTKIYTAPSLQKKWYNVLGNHDYR 125
Query: 131 GDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVL 190
G+ +AQ+S +LR +DNRWVC RS+ L+++NV+FFFVDTTP+V++YF + H YDW+G+L
Sbjct: 126 GNAKAQISHVLRYRDNRWVCFRSYTLNSENVDFFFVDTTPYVDKYFIEDKGHNYDWRGIL 185
Query: 191 PLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDA 250
P + Y + LLK+VD AL QSTA WK+V+ HH IK+ G HG+TQEL LP+LK+NNVD
Sbjct: 186 PRKRYTSNLLKDVDLALRQSTATWKVVIGHHTIKNIGHHGDTQELLIHFLPLLKANNVDL 245
Query: 251 YINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKT 310
Y+NGHDHCLEHI +S + F TSGGGSKAW GD K + E+K Y+DGQGF+S+ IS+T
Sbjct: 246 YMNGHDHCLEHISSLDSSVQFLTSGGGSKAWRGDTKQSEGDEMKFYYDGQGFMSVHISQT 305
Query: 311 NASIVFYDVFGKVLHTWSMSK 331
I F+DVFG +H W+ K
Sbjct: 306 QLRISFFDVFGNAIHKWNTCK 326
>UniRef100_Q9LQW0 F10B6.10 [Arabidopsis thaliana]
Length = 364
Score = 403 bits (1036), Expect = e-111
Identities = 195/333 (58%), Positives = 250/333 (74%), Gaps = 11/333 (3%)
Query: 7 QHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQ 64
+H +++ V L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQ
Sbjct: 35 KHKPVNLVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQ 90
Query: 65 SLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVL 124
S VA QMG +G+ L+I+FV+STGDNFYD+GL + DP F +SF +IYTAPSLQ+ WY+
Sbjct: 91 SQVALQMGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYS-- 148
Query: 125 GNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTY 184
GNHDYRGDV AQLSP+LR DNRWVC+RSFI++ + V+ FFVDTTPFV++YF P KH Y
Sbjct: 149 GNHDYRGDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVY 208
Query: 185 DWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILK 244
DW GVLP ++Y LLKE+D AL +S AKWKIV+ HH IKSAG HGNT ELE+ LLPIL+
Sbjct: 209 DWSGVLPRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQ 268
Query: 245 SNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAW-GGDIKPWDPKELKLYHDGQGFV 303
+N VD Y+NGHDHCLEHI +S I F TSGGGSKAW GGD+ +P+E++ Y+DGQGF+
Sbjct: 269 ANEVDLYVNGHDHCLEHISSVDSNIQFMTSGGGSKAWKGGDVNYVEPEEMRFYYDGQGFM 328
Query: 304 SMQISKTNASIVFYDVFGKVLHTWSMSKKLKEA 336
S+ +S+ +VFYDVFG VLH W K KEA
Sbjct: 329 SVHVSEAELRVVFYDVFGHVLHHW--KKTYKEA 359
>UniRef100_Q8GWH8 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 328
Score = 401 bits (1030), Expect = e-110
Identities = 193/321 (60%), Positives = 239/321 (74%), Gaps = 4/321 (1%)
Query: 15 IAVSVLLCLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQM 71
+ SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQM
Sbjct: 5 VCFSVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQM 64
Query: 72 GIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRG 131
G+VG+ L+I+FV+S GDNFYDDGLKGV+DP+F SF IYT PSLQ+ WY+VLGNHDYRG
Sbjct: 65 GVVGEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRG 124
Query: 132 DVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLP 191
+VEAQLS +L QKD RW C RSF+L + V+FFF DT PFVE+YFT+P HTYDW+ VLP
Sbjct: 125 NVEAQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLP 184
Query: 192 LESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAY 251
Y + LL ++D + +S A WK VV HH IK+AG HG TQEL +QLLPIL+ N VD Y
Sbjct: 185 RNKYISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLY 244
Query: 252 INGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTN 311
INGHDHCL+H I F TSGGGSKAW G ++PWDPKELKLY+DGQGF+S+ I+ +
Sbjct: 245 INGHDHCLQH-IGSHGKTQFLTSGGGSKAWRGHVQPWDPKELKLYYDGQGFMSLHITHSK 303
Query: 312 ASIVFYDVFGKVLHTWSMSKK 332
A ++YDV G VL S+SK+
Sbjct: 304 AKFIYYDVSGNVLLRSSLSKR 324
>UniRef100_Q707M7 Putative acid phosphatase [Lupinus luteus]
Length = 330
Score = 399 bits (1025), Expect = e-110
Identities = 194/327 (59%), Positives = 251/327 (76%), Gaps = 6/327 (1%)
Query: 10 LSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAH 69
+ S++ +S LC+L + +L +F H K SLNFLV+GDWGR+G YNQS +A
Sbjct: 9 MKSILFTISFGLCVLY----ASAELHKFAHSSK-HDGSLNFLVLGDWGRRGAYNQSEIAF 63
Query: 70 QMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDY 129
QMG VG+ L+I+FVVSTGDNFYD+GL D AF ESF ++YTA SLQ+ WY+VLGNHDY
Sbjct: 64 QMGKVGEKLDIDFVVSTGDNFYDNGLTSDQDTAFEESFTNVYTAKSLQKQWYSVLGNHDY 123
Query: 130 RGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGV 189
RGDVEAQLSP L+Q DNRW+CLRSFI+D++ VE FFVDTTPFVE+YFT+ KH YDW+G+
Sbjct: 124 RGDVEAQLSPFLKQIDNRWLCLRSFIVDSELVEIFFVDTTPFVEKYFTET-KHKYDWQGI 182
Query: 190 LPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVD 249
+P +SY LLK+++ A+ +STA+WKIVV HH I+S G HG+TQEL +QLLPIL+ N+VD
Sbjct: 183 IPQKSYITNLLKDLELAIKESTAQWKIVVGHHAIRSVGHHGDTQELIKQLLPILQENDVD 242
Query: 250 AYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISK 309
Y+NGHDHCLEHI D ES I F TSG GSKAW GDI+ + L ++DGQGF+S+++++
Sbjct: 243 FYMNGHDHCLEHISDTESSIQFLTSGAGSKAWRGDIQETNQGGLHFFYDGQGFMSVKLTQ 302
Query: 310 TNASIVFYDVFGKVLHTWSMSKKLKEA 336
T+A+I FYDV G VLH + SK L+ +
Sbjct: 303 TDATIEFYDVSGNVLHRLTSSKNLRSS 329
>UniRef100_Q9SCX8 Acid phosphatase type 5 precursor [Arabidopsis thaliana]
Length = 338
Score = 390 bits (1003), Expect = e-107
Identities = 184/327 (56%), Positives = 249/327 (75%), Gaps = 4/327 (1%)
Query: 10 LSSVIIAVSVLLCLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVA 68
L S ++S+LLC+ + ++ +L RF K S++F+V+GDWGR+G++NQSLVA
Sbjct: 8 LMSATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQSLVA 66
Query: 69 HQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHD 128
+QMG +G+ ++++FVVSTGDNFYD+GL DP F +SF +IYTAPSLQ+ WY+VLGNHD
Sbjct: 67 YQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLQKQWYSVLGNHD 126
Query: 129 YRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKG 188
YRGD EAQLS +LR+ D+RW+CLRSF++D + VE FFVDTTPFV+EY+T+ H+YDW+
Sbjct: 127 YRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYDWRA 186
Query: 189 VLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNV 248
V SY LL++++ +L S A+WKIVV HH ++S G HG+T+EL E+LLPILK N V
Sbjct: 187 VPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKENGV 246
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKP--WDPKELKLYHDGQGFVSMQ 306
D Y+NGHDHCL+H+ D++S I F TSG GSKAW GDI P +PK LK Y+DGQGF+S +
Sbjct: 247 DLYMNGHDHCLQHMSDEDSPIQFLTSGAGSKAWRGDINPVTINPKLLKFYYDGQGFMSAR 306
Query: 307 ISKTNASIVFYDVFGKVLHTWSMSKKL 333
+ ++A IVFYDVFG++LH W SK+L
Sbjct: 307 FTHSDAEIVFYDVFGEILHKWVTSKQL 333
>UniRef100_Q8LFV2 Acid phosphatase type 5 [Arabidopsis thaliana]
Length = 338
Score = 389 bits (999), Expect = e-107
Identities = 183/327 (55%), Positives = 250/327 (75%), Gaps = 4/327 (1%)
Query: 10 LSSVIIAVSVLLCLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVA 68
L S ++S+LLC+ + ++ +L RF K S++F+V+GDWGR+G++NQSLVA
Sbjct: 8 LISATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQSLVA 66
Query: 69 HQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHD 128
+QMG +G+ ++++FVVSTGDNFYD+GL DP F +SF +IYTAPSL++ WY+VLGNHD
Sbjct: 67 YQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLRKQWYSVLGNHD 126
Query: 129 YRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKG 188
YRGD EAQLS +LR+ D+RW+CLRSF++D + VE FFVDTTPFV+EY+T+ H+YDW+
Sbjct: 127 YRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYDWRA 186
Query: 189 VLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNV 248
V SY LL++++ +L S A+WKIVV HH ++S G HG+T+EL E+LLPILK N+V
Sbjct: 187 VPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKENSV 246
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKP--WDPKELKLYHDGQGFVSMQ 306
D Y+NGHDHCL+H+ D++S I F TSG GSKAW GDI P +PK LK Y+DGQGF+S +
Sbjct: 247 DLYMNGHDHCLQHMSDEDSPIQFLTSGAGSKAWRGDINPVTINPKLLKFYYDGQGFMSAR 306
Query: 307 ISKTNASIVFYDVFGKVLHTWSMSKKL 333
+ ++A IVFYDVFG++LH W SK+L
Sbjct: 307 FTHSDAEIVFYDVFGEILHKWVTSKQL 333
>UniRef100_Q8VYU7 At1g25230/F4F7_8 [Arabidopsis thaliana]
Length = 339
Score = 389 bits (998), Expect = e-107
Identities = 187/326 (57%), Positives = 239/326 (72%), Gaps = 3/326 (0%)
Query: 10 LSSVIIAVSVLLC--LLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLV 67
+ S+ I +++L+C LLS + +L +H P S++FLV+GDWGR G YNQS V
Sbjct: 7 IGSLSIVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQV 65
Query: 68 AHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNH 127
A QMG +G+ ++I FVVSTGDN YD+G+K +DDPAF SF +IYT+PSLQ+ WY VLGNH
Sbjct: 66 ALQMGRIGEEMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNH 125
Query: 128 DYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWK 187
DYRGDVEAQLSPILR D+RW+C+RSFI+D + E FFVDTTPFV+ YF P TYDW
Sbjct: 126 DYRGDVEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWS 185
Query: 188 GVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNN 247
GV P +SY +L E++ L +S+AKWKIVV HH IKSA HGNT+ELE LLPIL++N
Sbjct: 186 GVSPRKSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANK 245
Query: 248 VDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQI 307
VD Y+NGHDHCL+HI +S I F TSGGGSKAW G P+++K ++DGQGF+S++I
Sbjct: 246 VDLYMNGHDHCLQHISTSQSPIQFLTSGGGSKAWRGYYNWTTPEDMKFFYDGQGFMSVKI 305
Query: 308 SKTNASIVFYDVFGKVLHTWSMSKKL 333
+++ S+VFYDV G LH W SK L
Sbjct: 306 TRSELSVVFYDVSGNSLHKWDTSKML 331
>UniRef100_Q9SIS5 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 351
Score = 375 bits (963), Expect = e-103
Identities = 192/344 (55%), Positives = 235/344 (67%), Gaps = 36/344 (10%)
Query: 13 VIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+I ++ L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGD 132
+GK+LNI+F++STGDNFYDDG+ D F +SF +IYTA SLQ+ WYNVLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPL 192
V AQLSPILR D RW+CLRS++++ + V+ FFVDTTPF H YDW+GVLP
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPF---------DHVYDWRGVLPR 180
Query: 193 ESYRAELLK-------------------------EVDSALVQSTAKWKIVVAHHPIKSAG 227
Y LL +VD AL +S AKWKIVV HH IKSAG
Sbjct: 181 NKYLNSLLTVTFFFLLYHKLVISSNSLKIRIIKLDVDVALQESMAKWKIVVGHHTIKSAG 240
Query: 228 PHGNTQELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKP 287
HGNT ELE+QLLPIL++N VD YINGHDHCLEHI SGI F TSGGGSKAW GD+
Sbjct: 241 HHGNTIELEKQLLPILEANEVDLYINGHDHCLEHISSINSGIQFMTSGGGSKAWKGDVND 300
Query: 288 WDPKELKLYHDGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSK 331
W+P+E++ Y+DGQGF+S+ S+ +VFYD G VLH WS K
Sbjct: 301 WNPQEMRFYYDGQGFMSVYTSEAELRVVFYDGLGHVLHRWSTLK 344
>UniRef100_Q9FRH2 Hypothetical protein F4F7.38 [Arabidopsis thaliana]
Length = 333
Score = 375 bits (963), Expect = e-103
Identities = 184/326 (56%), Positives = 234/326 (71%), Gaps = 9/326 (2%)
Query: 10 LSSVIIAVSVLLC--LLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLV 67
+ S+ I +++L+C LLS + +L +H P S++FLV+GDWGR G YNQS V
Sbjct: 7 IGSLSIVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQV 65
Query: 68 AHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNH 127
A Q ++I FVVSTGDN YD+G+K +DDPAF SF +IYT+PSLQ+ WY VLGNH
Sbjct: 66 ALQ------EMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNH 119
Query: 128 DYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWK 187
DYRGDVEAQLSPILR D+RW+C+RSFI+D + E FFVDTTPFV+ YF P TYDW
Sbjct: 120 DYRGDVEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWS 179
Query: 188 GVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNN 247
GV P +SY +L E++ L +S+AKWKIVV HH IKSA HGNT+ELE LLPIL++N
Sbjct: 180 GVSPRKSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANK 239
Query: 248 VDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQI 307
VD Y+NGHDHCL+HI +S I F TSGGGSKAW G P+++K ++DGQGF+S++I
Sbjct: 240 VDLYMNGHDHCLQHISTSQSPIQFLTSGGGSKAWRGYYNWTTPEDMKFFYDGQGFMSVKI 299
Query: 308 SKTNASIVFYDVFGKVLHTWSMSKKL 333
+++ S+VFYDV G LH W SK L
Sbjct: 300 TRSELSVVFYDVSGNSLHKWDTSKML 325
>UniRef100_Q9LL81 Putative purple acid phosphatase [Ipomoea batatas]
Length = 312
Score = 367 bits (941), Expect = e-100
Identities = 183/313 (58%), Positives = 232/313 (73%), Gaps = 11/313 (3%)
Query: 12 SVIIAVSVLLCL-------LSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQ 64
+V +S++LCL + S+I E LP F HH SL+FLV+GDWGRKG+YNQ
Sbjct: 2 AVYSGISMVLCLWVGVVFGVCLASAIVE-LPTF-HHPTKGDGSLSFLVIGDWGRKGDYNQ 59
Query: 65 SLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVL 124
S VA QMG +G L I+FVVSTGDNFYD+GL G D AF ESF D+YTA SLQ+ WY+VL
Sbjct: 60 SQVAFQMGEIGDQLAIDFVVSTGDNFYDNGLTGEHDDAFTESFTDVYTAESLQKQWYSVL 119
Query: 125 GNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTY 184
GNHDYRGD EAQLS LR+ D+RW CLRSF+++T+ V+ FFVDTTPFVEEYF P +H Y
Sbjct: 120 GNHDYRGDAEAQLSSHLRKLDSRWPCLRSFVVNTETVDLFFVDTTPFVEEYFNSP-EHVY 178
Query: 185 DWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILK 244
DW+GV P ++Y +L ++ AL++ST KW+IV+ HH I+SAG HG+T+EL E+LLPIL+
Sbjct: 179 DWRGVFPQQTYTKNVLNGLEYALMKSTTKWRIVIGHHAIRSAGHHGDTKELVERLLPILR 238
Query: 245 SNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVS 304
+ NVD Y+NGHDH LEHI D ES I F TSG GSKAW GD+ D K + ++DGQGF+S
Sbjct: 239 TYNVDLYMNGHDHSLEHISDDESPIQFMTSGAGSKAWRGDV-TMDRKGVSFFYDGQGFMS 297
Query: 305 MQISKTNASIVFY 317
+Q+ +T+ IVFY
Sbjct: 298 VQLVETDIGIVFY 310
>UniRef100_Q7XH73 Putative purple acid phosphatase [Oryza sativa]
Length = 335
Score = 364 bits (935), Expect = 2e-99
Identities = 179/321 (55%), Positives = 235/321 (72%), Gaps = 3/321 (0%)
Query: 12 SVIIAVSVLLCLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQ 70
+V +A+ + LS+ +S A +L R EH + L LVVGDWGRKG YNQ+ VA Q
Sbjct: 2 AVALALLAAMSALSSCTSPATAELTRHEHPVAAGAP-LRLLVVGDWGRKGGYNQTRVAEQ 60
Query: 71 MGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYR 130
MG V + I+FVVSTGDNF ++GL GVDD AF++SF+D+YTA SL + WY VLGNHDYR
Sbjct: 61 MGKVAEETEIDFVVSTGDNFLENGLAGVDDMAFHDSFMDVYTAQSLHKPWYLVLGNHDYR 120
Query: 131 GDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVL 190
G+V AQ+ P LR+ D+R++C+RSFI+ V+FFFVDTTPF +Y+TDPG+ YDW+GV
Sbjct: 121 GNVLAQIDPALRKIDSRFICMRSFIVSAGIVDFFFVDTTPFQLQYWTDPGEDHYDWRGVA 180
Query: 191 PLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDA 250
P ++Y A LL++VD+A+ +STA WKI V HH ++S HG+TQEL E LLP+LK N VD
Sbjct: 181 PRDAYIANLLEDVDAAMKKSTATWKIAVGHHTMRSVSAHGDTQELLELLLPVLKENGVDF 240
Query: 251 YINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKT 310
YINGHDHCLEHI + S I +FTSGGGSKAW G + + K L+ ++DGQGF+S+++S+
Sbjct: 241 YINGHDHCLEHISSRNSPIQYFTSGGGSKAWRGIFQQNEDK-LQFFYDGQGFLSLELSEN 299
Query: 311 NASIVFYDVFGKVLHTWSMSK 331
A FYDVFG+ L+ WS SK
Sbjct: 300 RARFAFYDVFGEALYHWSFSK 320
>UniRef100_Q9SDZ8 Putative purple acid phosphatase precursor [Arabidopsis thaliana]
Length = 314
Score = 358 bits (919), Expect = 1e-97
Identities = 174/282 (61%), Positives = 215/282 (75%), Gaps = 2/282 (0%)
Query: 13 VIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+I ++ L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGD 132
+GK+LNI+F++STGDNFYDDG+ D F +SF +IYTA SLQ+ WYNVLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPL 192
V AQLSPILR D RW+CLRS++++ + V+ FFVDTTPFV+ YF +P H YDW+GVLP
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYI 252
Y LL +VD AL +S AKWKIVV HH IKSAG HGNT ELE+QLLPIL++N VD YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELK 294
NGHDHCLEHI SG+ F TSGGG KAW GD+ W+ +E++
Sbjct: 250 NGHDHCLEHISGINSGMQFMTSGGGFKAWKGDVNDWNLQEMR 291
>UniRef100_Q9LL79 Putative purple acid phosphatase [Phaseolus vulgaris]
Length = 331
Score = 354 bits (908), Expect = 2e-96
Identities = 169/256 (66%), Positives = 201/256 (78%), Gaps = 1/256 (0%)
Query: 29 SIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGD 88
+++ L R EH +K SL+ +V+GDWGRKG YNQS V+ QMG VG LNI+FV+STGD
Sbjct: 17 NVSALLQRLEHPVKADG-SLSLVVIGDWGRKGTYNQSEVSAQMGRVGAKLNIDFVISTGD 75
Query: 89 NFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDVEAQLSPILRQKDNRW 148
NFYDDGL GVDDPAF SF IYTA SLQ+ WY+VLGNHDYRGDVEAQL+ IL++ D RW
Sbjct: 76 NFYDDGLSGVDDPAFELSFSKIYTAKSLQKQWYSVLGNHDYRGDVEAQLNTILQKIDPRW 135
Query: 149 VCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPLESYRAELLKEVDSALV 208
+C RSFI+DT+ EFFFVDTTPFV++YF P HTYDW GVLP + Y ++LLK+++ AL
Sbjct: 136 ICQRSFIVDTEIAEFFFVDTTPFVDKYFLKPKDHTYDWTGVLPRDKYLSKLLKDLEIALK 195
Query: 209 QSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESG 268
STAKWKIVV HHP++S G HG+TQEL LLPIL++N+VD YINGHDHCLEHI S
Sbjct: 196 DSTAKWKIVVGHHPVRSIGHHGDTQELIRHLLPILEANDVDMYINGHDHCLEHISSTSSQ 255
Query: 269 IHFFTSGGGSKAWGGD 284
I F TSGGGSKAW GD
Sbjct: 256 IQFLTSGGGSKAWKGD 271
>UniRef100_Q9SIS6 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 304
Score = 332 bits (851), Expect = 9e-90
Identities = 174/321 (54%), Positives = 218/321 (67%), Gaps = 28/321 (8%)
Query: 15 IAVSVLLCLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQM 71
+ SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQM
Sbjct: 5 VCFSVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQM 64
Query: 72 GIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRG 131
G+VG+ L+I+FV+S GDNFYDDGLKGV+DP+F SF IYT PSLQ+ WY+VLGNHDYRG
Sbjct: 65 GVVGEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRG 124
Query: 132 DVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLP 191
+VEAQLS +L QKD RW C RSF+L + N+ F Y G
Sbjct: 125 NVEAQLSKVLTQKDWRWFCRRSFVLSSVNM---------FCCGY----------RNGGFF 165
Query: 192 LESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAY 251
L +++ L+ + +S A WK VV HH IK+AG HG TQEL +QLLPIL+ N VD Y
Sbjct: 166 LRGHKSFYLE-----IKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLY 220
Query: 252 INGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTN 311
INGHDHCL+H I F TSGGGSKAW G ++PWDPKELKLY+DGQGF+S+ I+ +
Sbjct: 221 INGHDHCLQH-IGSHGKTQFLTSGGGSKAWRGHVQPWDPKELKLYYDGQGFMSLHITHSK 279
Query: 312 ASIVFYDVFGKVLHTWSMSKK 332
A ++YDV G VLH S+SK+
Sbjct: 280 AKFIYYDVSGNVLHRSSLSKR 300
>UniRef100_Q64MK9 Acid phosphatase [Bacteroides fragilis]
Length = 310
Score = 135 bits (341), Expect = 1e-30
Identities = 92/297 (30%), Positives = 139/297 (45%), Gaps = 27/297 (9%)
Query: 46 QSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYE 105
+ NF + D GR G Y+Q +A MG +G+ + EFV++ GD + +G++ V+DP +
Sbjct: 30 KKFNFYIANDLGRNGYYDQKPIAELMGTMGEEIGPEFVLAAGDVHHFEGVRSVNDPLWMT 89
Query: 106 SFVDIYTAPSLQQIWYNVLGNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDT-----DN 160
+F IY+ P L WY VLGNH+YRG+ +A L RW + T
Sbjct: 90 NFELIYSHPELMIDWYPVLGNHEYRGNTQAVLD--YSGVSRRWTMPARYYTKTFEEKGAT 147
Query: 161 VEFFFVDTTPFVEEY----FTDPGKHTYDWKGVLPLESYRAELLKEVDSALVQSTAKWKI 216
V ++DT P +++Y T P D G L +DS L + W I
Sbjct: 148 VRIVWIDTAPLIDKYRNESATYPDACHQDMNGQLAW----------LDSVLTVAKEDWVI 197
Query: 217 VVAHHPIKSAGPHGNTQ--ELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFFTS 274
V HHPI + P ++ +L+ +L PIL+ + VD YI GH H +HI S I + +
Sbjct: 198 VAGHHPIYAETPKDQSERSDLQSRLDPILRKHKVDMYICGHIHNFQHIRVPGSDIDYIVN 257
Query: 275 GGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSK 331
GS A +KP + + GF I K ++ D G +L+T + K
Sbjct: 258 SAGSLA--RKVKPIEGTQ--FCSPEPGFSVCSIDKQELNLRMIDKKGNILYTVTRKK 310
>UniRef100_Q6DHF5 Zgc:92339 [Brachydanio rerio]
Length = 327
Score = 135 bits (340), Expect = 2e-30
Identities = 90/284 (31%), Positives = 146/284 (50%), Gaps = 18/284 (6%)
Query: 44 QQQSLNFLVVGDWGRKGNY-----NQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGV 98
Q SL F+ +GDWG + +Y +++ A ++ V ++ ++FV+S GDNFY DG+K V
Sbjct: 25 QASSLRFVGIGDWGGRPSYPFYTPHEADTAKELARVAQSSGLDFVLSLGDNFYYDGVKDV 84
Query: 99 DDPAFYESFVDIYTAPSLQQI-WYNVLGNHDYRGDVEAQLSPILRQKDNRWVCLRSFILD 157
DD F S+ +++ P+L I WY + GNHD+RG+V AQ++ + RW+ +
Sbjct: 85 DDTRFKFSYEQVFSHPALMTIPWYLIAGNHDHRGNVSAQIA--YSSRSERWIYPDLYYEL 142
Query: 158 TDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPLESYRA--ELLKEVDSALVQSTAKWK 215
V T + + G +TYD + E A + L ++ L + A +
Sbjct: 143 NFKVPHSNTSLTILMTDTVVVCG-NTYDGLDPVGPEDLAAANKQLAWIEQRLQSTKADFV 201
Query: 216 IVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFFTSG 275
IVV H+PI S G HG T+ L +L P+LK NV Y++GHDH L+ I ++ G F SG
Sbjct: 202 IVVGHYPIWSIGHHGPTKCLISKLRPLLKKYNVSLYLSGHDHSLQ-FIREDDGSSFVVSG 260
Query: 276 GGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNASIVFYDV 319
G + + K + S +++T+ S V+++V
Sbjct: 261 AGVE------EDSSTDHRKSFPSAWQLFSSPVNQTSGSFVYFEV 298
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 627,371,264
Number of Sequences: 2790947
Number of extensions: 28482614
Number of successful extensions: 61495
Number of sequences better than 10.0: 177
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 66
Number of HSP's that attempted gapping in prelim test: 61205
Number of HSP's gapped (non-prelim): 210
length of query: 337
length of database: 848,049,833
effective HSP length: 128
effective length of query: 209
effective length of database: 490,808,617
effective search space: 102579000953
effective search space used: 102579000953
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146550.5


