

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.6 + phase: 0
(486 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
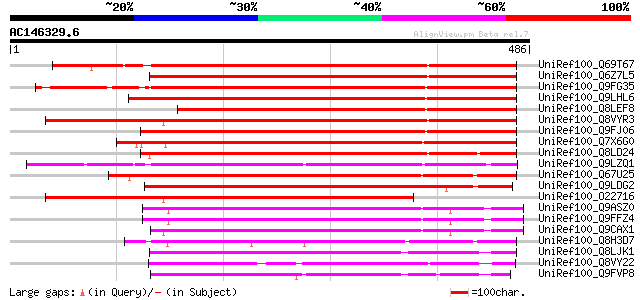
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q69T67 Lustrin A-like [Oryza sativa] 529 e-149
UniRef100_Q6Z7L5 Hypothetical protein OJ1448_G06.4 [Oryza sativa] 505 e-141
UniRef100_Q9FG35 Emb|CAB82953.1 [Arabidopsis thaliana] 496 e-139
UniRef100_Q9LHL6 Gb|AAF30301.1 [Arabidopsis thaliana] 490 e-137
UniRef100_Q8LEF8 Hypothetical protein [Arabidopsis thaliana] 450 e-125
UniRef100_Q8VYR3 Hypothetical protein At1g60790 [Arabidopsis tha... 431 e-119
UniRef100_Q9FJ06 Emb|CAB82953.1 [Arabidopsis thaliana] 419 e-115
UniRef100_Q7X6G0 OSJNBa0043L24.14 protein [Oryza sativa] 417 e-115
UniRef100_Q8LD24 Hypothetical protein [Arabidopsis thaliana] 383 e-105
UniRef100_Q9LZQ1 Hypothetical protein T12C14_90 [Arabidopsis tha... 371 e-101
UniRef100_Q67U25 Lustrin A-like [Oryza sativa] 362 2e-98
UniRef100_Q9LDG2 Hypothetical protein F24F17.6 [Arabidopsis thal... 319 1e-85
UniRef100_O22716 F8A5.30 protein [Arabidopsis thaliana] 306 9e-82
UniRef100_Q9ASZ0 AT5g06230/MBL20_11 [Arabidopsis thaliana] 284 4e-75
UniRef100_Q9FFZ4 Gb|AAF00670.1 [Arabidopsis thaliana] 284 4e-75
UniRef100_Q9CAX1 Hypothetical protein F24K9.24 [Arabidopsis thal... 284 5e-75
UniRef100_Q8H3D7 Leaf senescence related protein-like [Oryza sat... 278 3e-73
UniRef100_Q8LJK1 P0018C10.29 protein [Oryza sativa] 276 7e-73
UniRef100_Q8VY22 Hypothetical protein At1g29050 [Arabidopsis tha... 272 1e-71
UniRef100_Q9FVP8 Hypothetical protein F27K7.9 [Arabidopsis thali... 270 5e-71
>UniRef100_Q69T67 Lustrin A-like [Oryza sativa]
Length = 847
Score = 529 bits (1362), Expect = e-149
Identities = 253/443 (57%), Positives = 316/443 (71%), Gaps = 18/443 (4%)
Query: 41 STNETTTHDKTSHSSNKDTELVVSSRSSSQISRKE---------GSGLAPKQSNQNDGQL 91
S N+T+ S + N+ + V S SS S++E S P Q+ +
Sbjct: 405 SKNQTSVASTNSKNQNQTSAGVASGGSSGTTSKQEETTSQGSVGSSKDHPAQAINSKTSN 464
Query: 92 FPQIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDDSFPLYKS 151
+ ++ K A P+ +VD + S CD++ G+WV D+S+PLY
Sbjct: 465 YSEVLVKGNGSSTKQ-ASQKQPDKKVDWIKEMAS-------CDMFHGNWVRDESYPLYPE 516
Query: 152 GSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNR 211
GSCPHIDEPF+C+LNGR D Y+K RWQP +CN+PRLN ML+ LRGKRLVFVGDSLNR
Sbjct: 517 GSCPHIDEPFDCYLNGRPDRAYQKLRWQPSSCNIPRLNPTDMLERLRGKRLVFVGDSLNR 576
Query: 212 NMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEM 271
NMW+SLVCIL NSVK+K ++FEASGR EF+TE SYSF+FTDYNCS+EFFRSPFLVQEWEM
Sbjct: 577 NMWESLVCILRNSVKDKRKVFEASGRHEFKTEGSYSFLFTDYNCSVEFFRSPFLVQEWEM 636
Query: 272 PGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLN 331
G KKETLRLD+VE+S KYK+AD LIFNTGHWWTHEKT GK+YYQEGNH++ +LN
Sbjct: 637 KVSNGKKKETLRLDIVEQSSPKYKDADFLIFNTGHWWTHEKTSLGKDYYQEGNHVYSELN 696
Query: 332 VEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTEST 391
V +A KA +TWSRW+D N++PKKT V FRGYS SHF GG+W+SGG C+ ETEP++ E
Sbjct: 697 VVDAFHKALVTWSRWIDANVNPKKTTVLFRGYSASHFSGGQWNSGGSCDKETEPIRNEQY 756
Query: 392 DLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQ 451
LS PP M +E VI MKTPVVYLNI++MTD+R DAHPS+YR N++E+ ++ ++Q
Sbjct: 757 -LSTYPPKMSILEDVIHKMKTPVVYLNITRMTDYRKDAHPSIYRKRNLTEDERRSPERYQ 815
Query: 452 DCSHWCLPGVPDLWNEFVYAHLL 474
DCSHWCLPGVPD WNE +YA LL
Sbjct: 816 DCSHWCLPGVPDSWNELLYAQLL 838
>UniRef100_Q6Z7L5 Hypothetical protein OJ1448_G06.4 [Oryza sativa]
Length = 700
Score = 505 bits (1300), Expect = e-141
Identities = 221/343 (64%), Positives = 277/343 (80%), Gaps = 1/343 (0%)
Query: 132 NCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGK 191
+CD++ G+WV DDS+PLY GSCPHIDE FNC LNGR DN Y++ RWQP C++PRLN
Sbjct: 350 SCDMFYGNWVRDDSYPLYPEGSCPHIDESFNCPLNGRPDNAYQRLRWQPSGCSIPRLNPS 409
Query: 192 YMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFT 251
ML+ LRGKRLVFVGDSLNRNMW+SLVCIL NSVK+K ++FE SGRQ+F+ E SYSF+F
Sbjct: 410 DMLERLRGKRLVFVGDSLNRNMWESLVCILRNSVKDKRKVFEVSGRQQFRAEGSYSFLFQ 469
Query: 252 DYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHE 311
DYNCS+EFFRSPFLVQEWE P +KG KETLRLD++ S +YK+AD++IFNTGHWWTHE
Sbjct: 470 DYNCSVEFFRSPFLVQEWEFPVRKGLTKETLRLDMISNSFPRYKDADIIIFNTGHWWTHE 529
Query: 312 KTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGG 371
KT GK+YYQEGN ++ +LNV++A KA +TW++WVD++++PKKT VFFRGYS SHF GG
Sbjct: 530 KTSLGKDYYQEGNRVYSELNVDDAFQKALITWAKWVDSSVNPKKTTVFFRGYSSSHFSGG 589
Query: 372 EWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHP 431
+W+SGG C+ ETEP+ E L+ P M +E V+ GMKTPVVYLNI++MTD+R +AHP
Sbjct: 590 QWNSGGSCDKETEPITNEKF-LTPYPRKMSILEDVLSGMKTPVVYLNITRMTDYRKEAHP 648
Query: 432 SMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
S+YR ++EE KK +QDCSHWCLPGVPD WNE +YA ++
Sbjct: 649 SVYRKQKLTEEEKKSPQIYQDCSHWCLPGVPDSWNELLYAQIM 691
>UniRef100_Q9FG35 Emb|CAB82953.1 [Arabidopsis thaliana]
Length = 608
Score = 496 bits (1277), Expect = e-139
Identities = 233/450 (51%), Positives = 319/450 (70%), Gaps = 18/450 (4%)
Query: 25 NNSSSFSPPPSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKEGSGLAPKQS 84
NN+SS P S +TT + DTE V ++++S K + K +
Sbjct: 164 NNTSSH-------PLLSDKSSTTGSNNQSRTTADTETVNRNQTTSPAPSKAPVSVDLKTN 216
Query: 85 NQNDGQLFPQIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDD 144
+ ++ AS+ +Q K+ + S ++ + E L+N C+ ++G W+ DD
Sbjct: 217 SSSNSST----ASSTPKKQTKTVDLVSSVKQEIEKWS-----ESLKN-CEFFDGEWIKDD 266
Query: 145 SFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVF 204
S+PLYK GSC IDE FNC NGR D ++K +W+PK C++PRLNG +L+MLRG+RLVF
Sbjct: 267 SYPLYKPGSCNLIDEQFNCITNGRPDKDFQKLKWKPKKCSLPRLNGAILLEMLRGRRLVF 326
Query: 205 VGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPF 264
VGDSLNRNMW+SLVCIL SVK++++++EA GR F+ E YSF+F DYNC++EFF SPF
Sbjct: 327 VGDSLNRNMWESLVCILKGSVKDETKVYEARGRHHFRGEAEYSFVFQDYNCTVEFFVSPF 386
Query: 265 LVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGN 324
LVQEWE+ KKG+KKETLRLDLV +S ++YK ADV++FNTGHWWTHEKT +G++YYQEG+
Sbjct: 387 LVQEWEIVDKKGTKKETLRLDLVGKSSEQYKGADVIVFNTGHWWTHEKTSKGEDYYQEGS 446
Query: 325 HIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETE 384
+++ +L V EA KA TW RWV+ N++P K++VFFRGYS SHF GG+W+SGG C+SETE
Sbjct: 447 NVYHELAVLEAFRKALTTWGRWVEKNVNPAKSLVFFRGYSASHFSGGQWNSGGACDSETE 506
Query: 385 PMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETK 444
P+K + T L+ P M+ +E V++GMKTPV YLNI+++TD+R D HPS+YR ++SE+ K
Sbjct: 507 PIKND-TYLTPYPSKMKVLEKVLRGMKTPVTYLNITRLTDYRKDGHPSVYRKQSLSEKEK 565
Query: 445 KYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
K L +QDCSHWCLPGVPD WNE +YA L+
Sbjct: 566 KSPLLYQDCSHWCLPGVPDSWNEILYAELI 595
>UniRef100_Q9LHL6 Gb|AAF30301.1 [Arabidopsis thaliana]
Length = 556
Score = 490 bits (1262), Expect = e-137
Identities = 218/363 (60%), Positives = 281/363 (77%), Gaps = 1/363 (0%)
Query: 112 SPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDN 171
SP + K+ ++ +C+ +EG WV DDS+PLYK GSC IDE FNC NGR D
Sbjct: 175 SPKRKQRRKSSLRKVIESLKSCEFFEGDWVKDDSYPLYKPGSCNLIDEQFNCISNGRPDV 234
Query: 172 KYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRI 231
++K +W+PK C++PRLNG +L+M+RG+RLVFVGDSLNRNMW+SLVCIL SVK++S++
Sbjct: 235 DFQKLKWKPKQCSLPRLNGGKLLEMIRGRRLVFVGDSLNRNMWESLVCILKGSVKDESQV 294
Query: 232 FEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSC 291
FEA GR +F+ E YSF+F DYNC++EFF SPFLVQEWE+ K G+KKETLRLDLV +S
Sbjct: 295 FEAHGRHQFRWEAEYSFVFKDYNCTVEFFASPFLVQEWEVTEKNGTKKETLRLDLVGKSS 354
Query: 292 DKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNI 351
++YK AD+L+FNTGHWWTHEKT +G++YYQEG+ +H +L+V+EA KA TW RWVD N+
Sbjct: 355 EQYKGADILVFNTGHWWTHEKTSKGEDYYQEGSTVHPKLDVDEAFRKALTTWGRWVDKNV 414
Query: 352 DPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK 411
+PKK++VFFRGYSPSHF GG+W++GG C+ ETEP+K E T L+ M +E V++GMK
Sbjct: 415 NPKKSLVFFRGYSPSHFSGGQWNAGGACDDETEPIKNE-TYLTPYMLKMEILERVLRGMK 473
Query: 412 TPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYA 471
TPV YLNI+++TD+R DAHPS+YR +S E K L +QDCSHWCLPGVPD WNE YA
Sbjct: 474 TPVTYLNITRLTDYRKDAHPSIYRKQKLSAEESKSPLLYQDCSHWCLPGVPDSWNEIFYA 533
Query: 472 HLL 474
LL
Sbjct: 534 ELL 536
>UniRef100_Q8LEF8 Hypothetical protein [Arabidopsis thaliana]
Length = 329
Score = 450 bits (1157), Expect = e-125
Identities = 196/317 (61%), Positives = 256/317 (79%), Gaps = 1/317 (0%)
Query: 158 DEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSL 217
DE FNC NGR D ++K +W+PK C++PRLNG +L+MLRG+RLVFVGDSLNRNMW+SL
Sbjct: 1 DEQFNCITNGRPDKDFQKLKWKPKKCSLPRLNGAILLEMLRGRRLVFVGDSLNRNMWESL 60
Query: 218 VCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGS 277
VCIL SVK++++++EA GR F+ E YSF+F DYNC++EFF SPFLVQEWE+ KKG
Sbjct: 61 VCILKGSVKDETKVYEARGRHHFRGEAEYSFVFQDYNCTVEFFVSPFLVQEWEIVXKKGX 120
Query: 278 KKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAIS 337
KKETLRLDLV +S ++YK ADV++FNTGHWWTHEKT +G++YYQEG++++ +L V EA
Sbjct: 121 KKETLRLDLVGKSSEQYKGADVIVFNTGHWWTHEKTSKGEDYYQEGSNVYHELAVLEAFR 180
Query: 338 KAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENP 397
KA TW RWV+ N++P K++VFFRGYS SHF GG+W+SGG C+SETEP+K + T L+ P
Sbjct: 181 KALTTWGRWVEKNVNPAKSLVFFRGYSASHFSGGQWNSGGACDSETEPIKND-TYLTPYP 239
Query: 398 PMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWC 457
M+ +E V++GMKTPV YLNI+++TD+R D HPS+YR ++SE+ KK L +QDCSHWC
Sbjct: 240 SKMKVLEKVLRGMKTPVTYLNITRLTDYRKDGHPSVYRKQSLSEKEKKSPLLYQDCSHWC 299
Query: 458 LPGVPDLWNEFVYAHLL 474
LPGVPD WNE +YA L+
Sbjct: 300 LPGVPDSWNEILYAELI 316
>UniRef100_Q8VYR3 Hypothetical protein At1g60790 [Arabidopsis thaliana]
Length = 541
Score = 431 bits (1109), Expect = e-119
Identities = 203/445 (45%), Positives = 292/445 (65%), Gaps = 5/445 (1%)
Query: 34 PSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKEGSGLAPKQSNQNDGQLFP 93
PS+ + ++ TT + S + D+ + +EG+G N +
Sbjct: 90 PSFVVPDAGSKNTTLSEESKVPSFDSGQRSGETVKNSSLAEEGNGSVADDQNTLEANATT 149
Query: 94 QIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEH-LRNNCDIYEGSWVL--DDSFPLYK 150
+ ++ D + N ++ +V + +CDIY+GSWV D++ P Y
Sbjct: 150 SVGNSSSLVSDLGGRFVVPANTSKENGSVTEDRSRGSYEDCDIYDGSWVRADDETMPYYP 209
Query: 151 SGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLN 210
GSCP+ID FNC NGR D+ Y K+RWQP C++PRLNG L+ LRGK+LVFVGDS+N
Sbjct: 210 PGSCPYIDRDFNCHANGRPDDAYVKWRWQPNGCDIPRLNGTDFLEKLRGKKLVFVGDSIN 269
Query: 211 RNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWE 270
RNMW+SL+CIL +S+K+K R++E SGR+EF+ + Y+F F DYNC+++F SPF V+E
Sbjct: 270 RNMWESLICILRHSLKDKKRVYEISGRREFKKKGFYAFRFEDYNCTVDFVGSPFFVRESS 329
Query: 271 MPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQL 330
G G+ ETLRLD+++++ Y++AD+LIFNTGHWWTH+KT G+ YYQEGN ++ +L
Sbjct: 330 FKGVNGTTLETLRLDMMDKTTSMYRDADILIFNTGHWWTHDKTKLGENYYQEGNVVYPRL 389
Query: 331 NVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTES 390
V EA +A +TW++WVD NID +T + FRGYS +HFRGG W+SGG+C ETEP+ S
Sbjct: 390 KVLEAYKRALITWAKWVDKNIDRSQTHIVFRGYSVTHFRGGPWNSGGQCHKETEPIFNTS 449
Query: 391 TDLSENPPMMRTIESVIKG-MKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLK 449
L++ P M+ +E +++ MKTPV+Y+NIS++TDFR D HPS+YR + +E+ K+ +
Sbjct: 450 Y-LAKYPSKMKALEYILRDTMKTPVIYMNISRLTDFRKDGHPSIYRMVYRTEKEKREAVS 508
Query: 450 HQDCSHWCLPGVPDLWNEFVYAHLL 474
HQDCSHWCLPGVPD WN+ +Y LL
Sbjct: 509 HQDCSHWCLPGVPDTWNQLLYVSLL 533
>UniRef100_Q9FJ06 Emb|CAB82953.1 [Arabidopsis thaliana]
Length = 457
Score = 419 bits (1076), Expect = e-115
Identities = 197/354 (55%), Positives = 257/354 (71%), Gaps = 3/354 (0%)
Query: 123 IQSDEHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKN 182
I S+ +CDI++G+WV DDS P+Y G CP +++ FNCF NGR D+ + + RWQP
Sbjct: 90 IDSETKELASCDIFDGTWVFDDSEPVYLPGYCPFVEDKFNCFKNGRPDSGFLRHRWQPHG 149
Query: 183 CNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQ-EFQ 241
C++PR +GK MLKMLRGKR+VFVGDSLNRNMW+SLVC L +++++K+R+ + G+Q
Sbjct: 150 CSIPRFDGKKMLKMLRGKRVVFVGDSLNRNMWESLVCSLRSTLEDKNRVSKIIGKQSNLP 209
Query: 242 TEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDK-YKNADVL 300
E Y F F D+ CSI+F +SPFLVQE E+ G ++ETLRLD+++RS K YKNAD++
Sbjct: 210 NEGFYGFRFNDFECSIDFIKSPFLVQESEVVDVYGKRRETLRLDMIQRSMTKIYKNADIV 269
Query: 301 IFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFF 360
IFNTGHWWTH+KT EGK YYQEGN ++ +L V+EA +KA TW+ WVD+NI+ KT VFF
Sbjct: 270 IFNTGHWWTHQKTYEGKGYYQEGNRVYERLEVKEAYTKAIHTWADWVDSNINSTKTRVFF 329
Query: 361 RGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNIS 420
GYS SHFR G W+SGG+C+ ET P++ E T P MM+ +ESVI MKTPV Y+NI+
Sbjct: 330 VGYSSSHFRKGAWNSGGQCDGETRPIQNE-TYTGVYPWMMKVVESVISEMKTPVFYMNIT 388
Query: 421 KMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
KMT +R D HPS+YR T +QDCSHWCLPGVPD WN+ +YA LL
Sbjct: 389 KMTWYRTDGHPSVYRQPADPRGTSPAAGMYQDCSHWCLPGVPDSWNQLLYATLL 442
>UniRef100_Q7X6G0 OSJNBa0043L24.14 protein [Oryza sativa]
Length = 711
Score = 417 bits (1071), Expect = e-115
Identities = 203/391 (51%), Positives = 263/391 (66%), Gaps = 18/391 (4%)
Query: 101 PQQDKSFAPMPSPNARV-------DSKNV----IQSDEHLRNNCDIYEGSWVLDD----- 144
P + A MP P A + D++ V +QS CD++ G WV DD
Sbjct: 313 PDRRDDGAAMPLPAATIINTSTVGDNRVVWTSGVQSGLVSFAKCDVFSGRWVRDDDEGGG 372
Query: 145 SFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVF 204
++P Y GSCPHID+ FNC NGR D + ++RWQP C++PRLN L+ LRG+R++F
Sbjct: 373 AYPFYPPGSCPHIDDDFNCHKNGRADTGFLRWRWQPHGCDIPRLNPIDFLERLRGQRIIF 432
Query: 205 VGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPF 264
VGDSLNRNMW+SLVCIL + V++K R++EASGR +F+T YSF F DYNCS++F RS F
Sbjct: 433 VGDSLNRNMWESLVCILRHGVRDKRRMYEASGRNQFKTRGYYSFRFRDYNCSVDFIRSIF 492
Query: 265 LVQEWEMPGKKGSKKET-LRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEG 323
LV+E K G+ + LRLD ++ + Y+ AD+++FNTGHWWTH KT G YYQEG
Sbjct: 493 LVKEMINETKGGAVVDAKLRLDELDETTPAYRTADIVVFNTGHWWTHWKTSRGLNYYQEG 552
Query: 324 NHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESET 383
N++H L V +A +A TW+RWVD NID +T V FRGYS +HFRGG+W+SGGRC ET
Sbjct: 553 NYVHPSLEVMDAYKRALTTWARWVDKNIDSTRTQVVFRGYSLTHFRGGQWNSGGRCHRET 612
Query: 384 EPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEET 443
EP+ T L+E P MR +E V+ M+TPV+YLNIS MTD+R DAHPS+YR +EE
Sbjct: 613 EPI-FNRTHLAEYPEKMRILEQVLGRMRTPVIYLNISAMTDYRKDAHPSVYRVRYETEEE 671
Query: 444 KKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
+ + QDCSHWCLPGVPD WNE +YA LL
Sbjct: 672 RMAAVAKQDCSHWCLPGVPDSWNELLYASLL 702
>UniRef100_Q8LD24 Hypothetical protein [Arabidopsis thaliana]
Length = 485
Score = 383 bits (983), Expect = e-105
Identities = 170/357 (47%), Positives = 241/357 (66%), Gaps = 8/357 (2%)
Query: 123 IQSDEHL----RNNCDIYEGSWVLDDS-FPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFR 177
++S EH CD+Y GSWV DD +PLY+ GSCP++D+ F+C NGRRD+ Y +R
Sbjct: 127 VESVEHAVIEKMRGCDLYRGSWVKDDDEYPLYQPGSCPYVDDAFDCQRNGRRDSDYLNWR 186
Query: 178 WQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGR 237
W+P C++PR N L LRGK L+ VGDS+NRN ++S++C+L + +KSR++E G
Sbjct: 187 WKPDGCDLPRFNATDFLVKLRGKSLMLVGDSMNRNQFESMLCVLREGLSDKSRMYEVHGH 246
Query: 238 QEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNA 297
+ + F F DYNC++EF RS FLV+E +G+ TL +D +++S K+K A
Sbjct: 247 NITKGRGYFVFKFEDYNCTVEFVRSHFLVREGVRANAQGNTNPTLSIDRIDKSHAKWKRA 306
Query: 298 DVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTI 357
D+L+FNTGHWW H KT GK YY+EG++I+ + + EA ++ TW++W+D N++PKK +
Sbjct: 307 DILVFNTGHWWVHGKTARGKNYYKEGDYIYPKFDATEAYRRSLKTWAKWIDQNVNPKKQL 366
Query: 358 VFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYL 417
VF+RGYS +HFRGGEWDSGG C E EP+K S + P M+ ++ IK M+ PV+ L
Sbjct: 367 VFYRGYSSAHFRGGEWDSGGSCNGEVEPVKKGSI-IDSYPLKMKIVQEAIKEMQVPVILL 425
Query: 418 NISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
N++K+T+FR D HPS+Y N + KK + QDCSHWCLPGVPD+WN +YA LL
Sbjct: 426 NVTKLTNFRKDGHPSIYGKTN--TDGKKVSTRRQDCSHWCLPGVPDVWNHLIYASLL 480
>UniRef100_Q9LZQ1 Hypothetical protein T12C14_90 [Arabidopsis thaliana]
Length = 475
Score = 371 bits (953), Expect = e-101
Identities = 190/461 (41%), Positives = 267/461 (57%), Gaps = 17/461 (3%)
Query: 16 FSKVFSHIFNNSSSFSPPPSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKE 75
F+ S + ++SS P P Q + +S S L+ + +S +
Sbjct: 29 FTFFSSSLLKSNSSLYPTPEANFQIDLSPIAAISDSSVSPQASPILISTHFNSPE--NTS 86
Query: 76 GSGLAPKQSNQNDGQLFPQIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEHLRNNCDI 135
GS + G+ + I +PS N D K + +E CD+
Sbjct: 87 GSSKISVFEQKISGESLVKEVREIANLTSIKVIELPSNNGE-DKKTEKRIEE-----CDV 140
Query: 136 YEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLK 195
+G WV D +PLY + SCP IDE F C NGR D Y +RW+P++C+ PR N ML+
Sbjct: 141 TKGKWVYDSDYPLYTNASCPFIDEGFGCQSNGRLDLNYMNWRWEPQDCHAPRFNATKMLE 200
Query: 196 MLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNC 255
M+RGKRLVFVGDS+NRN W+S++C+L +VK+ R++E R+ + + +YSF F DY C
Sbjct: 201 MIRGKRLVFVGDSINRNQWESMLCLLFQAVKDPKRVYETHNRRITKEKGNYSFRFVDYKC 260
Query: 256 SIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTM 314
++EF+ + FLV+E GKK ++ETLR+D ++R+ ++K A++L+FNT HWW+H KT
Sbjct: 261 TVEFYVTHFLVREGRARIGKK--RRETLRIDAMDRTSSRWKGANILVFNTAHWWSHYKTK 318
Query: 315 EGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWD 374
G YYQEG+ IH +L+V A KA TWS WVD N+DPKKT VFFR +PSHF GGEW+
Sbjct: 319 SGVNYYQEGDLIHPKLDVSTAFKKALQTWSSWVDKNVDPKKTRVFFRSAAPSHFSGGEWN 378
Query: 375 SGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMY 434
SGG C P+ T +E V+K M+TPV LN+S ++ +R DAHPS+Y
Sbjct: 379 SGGHCREANMPL--NQTFKPSYSSKKSIVEDVLKQMRTPVTLLNVSGLSQYRIDAHPSIY 436
Query: 435 RNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
+ ++ QDCSHWCLPGVPD WN F+Y HLL+
Sbjct: 437 GTKPENRRSRAV----QDCSHWCLPGVPDTWNHFLYLHLLH 473
>UniRef100_Q67U25 Lustrin A-like [Oryza sativa]
Length = 463
Score = 362 bits (928), Expect = 2e-98
Identities = 170/385 (44%), Positives = 240/385 (62%), Gaps = 7/385 (1%)
Query: 93 PQIASTIKPQQDKSFAPM--PSPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDDSF-PLY 149
P T P S PM P+ ++ S + E CD+Y+G W D++ PLY
Sbjct: 72 PLAPLTTPPPPPPSTLPMAPPTRSSPAASPPPPEKGEGGGTTCDLYDGEWARDEAARPLY 131
Query: 150 KSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSL 209
G+CP++DE + C NGR D Y ++RW P+ C +PR N L LRGKRL+ VGDS+
Sbjct: 132 APGTCPYVDEAYACASNGRPDAAYTRWRWAPRRCRLPRFNATDFLATLRGKRLMLVGDSM 191
Query: 210 NRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEW 269
NRN ++SL+CIL ++ +K+R+FE G + + + F F DY+C++EF RS FLV+E
Sbjct: 192 NRNQFESLLCILREAIPDKTRMFETHGYRISKGRGYFVFKFVDYDCTVEFVRSHFLVREG 251
Query: 270 EMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQ 329
++ + L++D +++S +++ DVL+FNTGHWWTH KT G YY+EG+ ++ Q
Sbjct: 252 VRYNRQKNSNPILQIDRIDKSASRWRKVDVLVFNTGHWWTHGKTARGINYYKEGDTLYPQ 311
Query: 330 LNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTE 389
+ EA +A TW+RW+D N+DP K++VF+RGYS +HFRGGEWDSGG C E EP
Sbjct: 312 FDSTEAYRRALKTWARWIDKNMDPAKSVVFYRGYSTAHFRGGEWDSGGSCSGEREP-TLR 370
Query: 390 STDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLK 449
+ P MR +E VI M+ PV LN++K+T+FR D HPS+Y + KK +
Sbjct: 371 GAVIGSYPRKMRIVEEVIGRMRFPVRLLNVTKLTNFRRDGHPSVY---GKAAAGKKVSRR 427
Query: 450 HQDCSHWCLPGVPDLWNEFVYAHLL 474
QDCSHWCLPGVPD WNE +YA L+
Sbjct: 428 KQDCSHWCLPGVPDAWNELIYASLV 452
>UniRef100_Q9LDG2 Hypothetical protein F24F17.6 [Arabidopsis thaliana]
Length = 469
Score = 319 bits (817), Expect = 1e-85
Identities = 155/351 (44%), Positives = 217/351 (61%), Gaps = 10/351 (2%)
Query: 127 EHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMP 186
E + CD+++G WV D+S+PLY+S C +DE F C GR D Y ++RWQP++CN+P
Sbjct: 97 EESGSGCDVFDGDWVWDESYPLYQSKDCRFLDEGFRCSDFGRSDLFYTQWRWQPRHCNLP 156
Query: 187 RLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSY 246
R + K ML+ LR KRLVFVGDS+ RN W+SL+C+LS++VKN+S I+E +G + +
Sbjct: 157 RFDAKLMLEKLRDKRLVFVGDSIGRNQWESLLCLLSSAVKNESLIYEINGSPITKHKGFL 216
Query: 247 SFIFTDYNCSIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTG 305
F F +YNC++E++RSPFLV + P G G K +L+LD ++ + K+++ADVL+ NTG
Sbjct: 217 VFKFEEYNCTVEYYRSPFLVPQSRPPIGSPGKVKTSLKLDTMDWTSSKWRDADVLVLNTG 276
Query: 306 HWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSP 365
HWW KT Y+QEG + ++NV++A +A T +W+ T +D KT VFFR ++P
Sbjct: 277 HWWNEGKTTRTGCYFQEGEEVKLKMNVDDAYKRALNTVVKWIHTELDSNKTQVFFRTFAP 336
Query: 366 SHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVI-----KGMKTPVVYLNIS 420
HFRGG+W +GG C ET P S SE ++ + V+ + V LNI+
Sbjct: 337 VHFRGGDWKTGGTCHMETLPEIGTSLASSETWEQLKILRDVLSHNSNRSETVKVKLLNIT 396
Query: 421 KMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYA 471
M R D HPS+Y L QDCSHWCLPGVPD WNE YA
Sbjct: 397 AMAAQRKDGHPSLY----YLGPHGPAPLHRQDCSHWCLPGVPDTWNELFYA 443
>UniRef100_O22716 F8A5.30 protein [Arabidopsis thaliana]
Length = 664
Score = 306 bits (784), Expect = 9e-82
Identities = 149/349 (42%), Positives = 217/349 (61%), Gaps = 4/349 (1%)
Query: 34 PSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKEGSGLAPKQSNQNDGQLFP 93
PS+ + ++ TT + S + D+ + +EG+G N +
Sbjct: 90 PSFVVPDAGSKNTTLSEESKVPSFDSGQRSGETVKNSSLAEEGNGSVADDQNTLEANATT 149
Query: 94 QIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEH-LRNNCDIYEGSWVL--DDSFPLYK 150
+ ++ D + N ++ +V + +CDIY+GSWV D++ P Y
Sbjct: 150 SVGNSSSLVSDLGGRFVVPANTSKENGSVTEDRSRGSYEDCDIYDGSWVRADDETMPYYP 209
Query: 151 SGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLN 210
GSCP+ID FNC NGR D+ Y K+RWQP C++PRLNG L+ LRGK+LVFVGDS+N
Sbjct: 210 PGSCPYIDRDFNCHANGRPDDAYVKWRWQPNGCDIPRLNGTDFLEKLRGKKLVFVGDSIN 269
Query: 211 RNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWE 270
RNMW+SL+CIL +S+K+K R++E SGR+EF+ + Y+F F DYNC+++F SPF V+E
Sbjct: 270 RNMWESLICILRHSLKDKKRVYEISGRREFKKKGFYAFRFEDYNCTVDFVGSPFFVRESS 329
Query: 271 MPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQL 330
G G+ ETLRLD+++++ Y++AD+LIFNTGHWWTH+KT G+ YYQEGN ++ +L
Sbjct: 330 FKGVNGTTLETLRLDMMDKTTSMYRDADILIFNTGHWWTHDKTKLGENYYQEGNVVYPRL 389
Query: 331 NVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRG-GEWDSGGR 378
V EA +A +TW++WVD NID +T + FRGYS +HF W+ GR
Sbjct: 390 KVLEAYKRALITWAKWVDKNIDRSQTHIVFRGYSVTHFSFISAWNVTGR 438
>UniRef100_Q9ASZ0 AT5g06230/MBL20_11 [Arabidopsis thaliana]
Length = 372
Score = 284 bits (727), Expect = 4e-75
Identities = 145/367 (39%), Positives = 209/367 (56%), Gaps = 17/367 (4%)
Query: 125 SDEHLRNNCDIYEGSWVLDDSFP------LYKSGSCPHIDEPFNCFLNGRRDNKYEKFRW 178
S + + CD +G WV S L+ C +D F C +GR+D+ Y +RW
Sbjct: 13 SSQTVTKECDYSKGKWVRRASSSSSSVNGLFYGEECRFLDSGFRCHKHGRKDSGYLDWRW 72
Query: 179 QPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQ 238
QP C++PR N +L+ R R+VFVGDS+ RN W+SL+C+LS ++ NKS I+E +G
Sbjct: 73 QPHGCDLPRFNASDLLERSRNGRIVFVGDSIGRNQWESLMCMLSQAIPNKSEIYEVNGNP 132
Query: 239 EFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSK-KETLRLDLVERSCDKYKNA 297
+ + S F N ++E+ RSPFLV P K + K T+R+D ++ +
Sbjct: 133 ITKHKGFLSMRFPRENLTVEYHRSPFLVVIGRPPDKSPKEIKTTVRVDEFNWQSKRWVGS 192
Query: 298 DVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTI 357
DVL+FN+GHWW +KT+ Y++EG ++ + V EA K+ TW WV +DP K+
Sbjct: 193 DVLVFNSGHWWNEDKTVLTGCYFEEGRKVNKTMGVMEAFGKSLKTWKSWVLEKLDPDKSY 252
Query: 358 VFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK---TPV 414
VFFR YSP H+R G W++GG C++E EP +T+ L + I VI+ M+ + V
Sbjct: 253 VFFRSYSPVHYRNGTWNTGGLCDAEIEP-ETDKRKLEPDASHNEYIYKVIEEMRYRHSKV 311
Query: 415 VYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
+LNI+ +T+FR D H S YR S + QDCSHWCLPGVPD WNE +YA LL
Sbjct: 312 KFLNITYLTEFRKDGHISRYREQGTSVDVP------QDCSHWCLPGVPDTWNEILYAQLL 365
Query: 475 YIMKRNR 481
+ R +
Sbjct: 366 SMNYRTK 372
>UniRef100_Q9FFZ4 Gb|AAF00670.1 [Arabidopsis thaliana]
Length = 413
Score = 284 bits (727), Expect = 4e-75
Identities = 145/367 (39%), Positives = 209/367 (56%), Gaps = 17/367 (4%)
Query: 125 SDEHLRNNCDIYEGSWVLDDSFP------LYKSGSCPHIDEPFNCFLNGRRDNKYEKFRW 178
S + + CD +G WV S L+ C +D F C +GR+D+ Y +RW
Sbjct: 54 SSQTVTKECDYSKGKWVRRASSSSSSVNGLFYGEECRFLDSGFRCHKHGRKDSGYLDWRW 113
Query: 179 QPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQ 238
QP C++PR N +L+ R R+VFVGDS+ RN W+SL+C+LS ++ NKS I+E +G
Sbjct: 114 QPHGCDLPRFNASDLLERSRNGRIVFVGDSIGRNQWESLMCMLSQAIPNKSEIYEVNGNP 173
Query: 239 EFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSK-KETLRLDLVERSCDKYKNA 297
+ + S F N ++E+ RSPFLV P K + K T+R+D ++ +
Sbjct: 174 ITKHKGFLSMRFPRENLTVEYHRSPFLVVIGRPPDKSPKEIKTTVRVDEFNWQSKRWVGS 233
Query: 298 DVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTI 357
DVL+FN+GHWW +KT+ Y++EG ++ + V EA K+ TW WV +DP K+
Sbjct: 234 DVLVFNSGHWWNEDKTVLTGCYFEEGRKVNKTMGVMEAFGKSLKTWKSWVLEKLDPDKSY 293
Query: 358 VFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK---TPV 414
VFFR YSP H+R G W++GG C++E EP +T+ L + I VI+ M+ + V
Sbjct: 294 VFFRSYSPVHYRNGTWNTGGLCDAEIEP-ETDKRKLEPDASHNEYIYKVIEEMRYRHSKV 352
Query: 415 VYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
+LNI+ +T+FR D H S YR S + QDCSHWCLPGVPD WNE +YA LL
Sbjct: 353 KFLNITYLTEFRKDGHISRYREQGTSVDVP------QDCSHWCLPGVPDTWNEILYAQLL 406
Query: 475 YIMKRNR 481
+ R +
Sbjct: 407 SMNYRTK 413
>UniRef100_Q9CAX1 Hypothetical protein F24K9.24 [Arabidopsis thaliana]
Length = 427
Score = 284 bits (726), Expect = 5e-75
Identities = 143/356 (40%), Positives = 203/356 (56%), Gaps = 14/356 (3%)
Query: 133 CDIYEGSWVL---DDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLN 189
CD G WV D Y C +D F C NGR+D+ + ++RWQP C++PR N
Sbjct: 79 CDYSYGRWVRRRRDVDETSYYGEECRFLDPGFRCLNNGRKDSGFRQWRWQPHGCDLPRFN 138
Query: 190 GKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFI 249
L+ R R+VFVGDS+ RN W+SL+C+LS +V NKS I+E +G + + S
Sbjct: 139 ASDFLERSRNGRIVFVGDSIGRNQWESLLCMLSQAVSNKSEIYEVNGNPISKHKGFLSMR 198
Query: 250 FTDYNCSIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWW 308
F + N ++E+ R+PFLV P K T+R+D K+ +DVL+FNTGHWW
Sbjct: 199 FPEQNLTVEYHRTPFLVVVGRPPENSPVDVKMTVRVDEFNWQSKKWVGSDVLVFNTGHWW 258
Query: 309 THEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHF 368
+KT Y+QEG ++ + V E K+ TW WV +D +++ VFFR +SP H+
Sbjct: 259 NEDKTFIAGCYFQEGGKLNKTMGVMEGFEKSLKTWKSWVLERLDSERSHVFFRSFSPVHY 318
Query: 369 RGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK---TPVVYLNISKMTDF 425
R G W+ GG C+++TEP +T+ + +P I I+ M+ + V +LNI+ +T+F
Sbjct: 319 RNGTWNLGGLCDADTEP-ETDMKKMEPDPIHNNYISQAIQEMRYEHSKVKFLNITYLTEF 377
Query: 426 RHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLYIMKRNR 481
R DAHPS YR E+ QDCSHWCLPGVPD WNE +YA LL + R +
Sbjct: 378 RKDAHPSRYREPGTPEDAP------QDCSHWCLPGVPDTWNEILYAQLLAMNYRTK 427
>UniRef100_Q8H3D7 Leaf senescence related protein-like [Oryza sativa]
Length = 441
Score = 278 bits (710), Expect = 3e-73
Identities = 146/384 (38%), Positives = 222/384 (57%), Gaps = 27/384 (7%)
Query: 108 APMPSPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDDSF---PLYKSGSCPHIDEPFNCF 164
A S + VD V+ + EH CD+ +G WV DD PLY+ CP +DE F C
Sbjct: 64 ATTTSSSDAVDDVAVVAAAEH----CDVVDGEWVRDDDDERRPLYEPRRCPFVDEGFRCR 119
Query: 165 LNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNS 224
NGR D+ + K+RWQP++C +PR + K +L+ LR +RLVFVGDS+ RN W+S++C+L+ +
Sbjct: 120 ENGRPDDAFAKWRWQPRHCTLPRFDAKNLLETLRNRRLVFVGDSIGRNQWESMLCMLATA 179
Query: 225 V--------KNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGK-- 274
V + I+E +G + E + SF F DYNC++E +RSP+LV+ P +
Sbjct: 180 VAFAGDGDGDKAASIYEVNGSPITKHEGALSFRFRDYNCTVEHYRSPYLVRRGRPPPRRA 239
Query: 275 -KGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVE 333
+G+ TL+LD ++ ++K+ADV++FNTGHWW+ E+ ++ + +Q G + +++E
Sbjct: 240 HRGAACSTLQLDAMDARAHRWKDADVVVFNTGHWWSRERLLQLRCNFQVGKKLILNMSIE 299
Query: 334 EAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEP-MKTESTD 392
A +A T + WV ++P K++V FR YSP+H R S G C ET P + +
Sbjct: 300 AAYQRAMNTLTSWVHREVNPHKSLVIFRTYSPAHTRA---SSNGGCAKETTPELNSSRIS 356
Query: 393 LSENPPMMR-TIESVIKGMKTPVV-YLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKH 450
L P M+ E G + LN++ MT R D HPS++ N++ + +
Sbjct: 357 LHRWPGMVNPAFEPSRSGTAAAKLRVLNVTLMTAQRRDGHPSVF---NVAAAARSPARQR 413
Query: 451 QDCSHWCLPGVPDLWNEFVYAHLL 474
DCSHWCLPGVPD WNE +YA +L
Sbjct: 414 ADCSHWCLPGVPDAWNELLYAMIL 437
>UniRef100_Q8LJK1 P0018C10.29 protein [Oryza sativa]
Length = 434
Score = 276 bits (707), Expect = 7e-73
Identities = 133/344 (38%), Positives = 193/344 (55%), Gaps = 16/344 (4%)
Query: 132 NCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGK 191
+CD+++GSWV D +PLY S CP ++ FNC NGR+D Y K+RW+P+ C++PR + +
Sbjct: 89 SCDVFDGSWVPDRRYPLYNSSDCPFVERGFNCLANGRKDTGYLKWRWKPRGCDLPRFSAR 148
Query: 192 YMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFT 251
+L+ LRGKR+VFVGDS++R W+S +C+L V+N ++E +G Q +T F
Sbjct: 149 DVLERLRGKRVVFVGDSMSRTQWESFICMLMAGVENPKTVYEVNGNQISKTIRFLGVRFA 208
Query: 252 DYNCSIEFFRSPFLVQEWEMPGKKGSK-KETLRLDLVERSCDKYKNADVLIFNTGHWWTH 310
+N ++EFFRS FLVQ+ P + + L+LD ++ K++NADVLIFN+GHWWT
Sbjct: 209 SFNLNVEFFRSVFLVQQSPAPRSSPKRVRAILKLDKMDNISRKWENADVLIFNSGHWWTP 268
Query: 311 EKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRG 370
K + Y++ G + ++ A A TW+ WV +D K+T VFFR Y PSH
Sbjct: 269 SKLFDMGCYFEAGGLLKLGTSINSAFKMALETWASWVKEKVDLKRTHVFFRTYEPSH--- 325
Query: 371 GEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAH 430
W + E T + + V+ M P LN++ M FR DAH
Sbjct: 326 --WSGSNQKVCEVTEFPTAEAKGDDRSEFGDILAGVVVNMSVPATILNVTLMGAFRSDAH 383
Query: 431 PSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
++ + + DCSHWCLPGVPD WNE V +HLL
Sbjct: 384 IGIWSHPSTI----------LDCSHWCLPGVPDAWNELVISHLL 417
>UniRef100_Q8VY22 Hypothetical protein At1g29050 [Arabidopsis thaliana]
Length = 380
Score = 272 bits (696), Expect = 1e-71
Identities = 131/344 (38%), Positives = 194/344 (56%), Gaps = 25/344 (7%)
Query: 131 NNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNG 190
+ C++++G WV D S+P Y S CP ID F+C GR D ++ K+ WQP++C +PR +G
Sbjct: 59 SGCNLFQGRWVFDASYPFYDSSKCPFIDGEFDCLKFGRPDKQFLKYSWQPESCTIPRFDG 118
Query: 191 KYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIF 250
L+ RGKR++FVGDSL+ NMW+SL C++ SV N F + + F
Sbjct: 119 GAFLRKYRGKRVMFVGDSLSLNMWESLACMIHASVPNAKTTF-------LKRTPLSTLTF 171
Query: 251 TDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTH 310
+Y ++ +R+P++V K L L +E D +KN DVL+FN+ HWWTH
Sbjct: 172 QEYGVTLYLYRTPYIVDI-----SKERVGRVLNLGAIEGGADAWKNMDVLVFNSWHWWTH 226
Query: 311 EKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRG 370
+ +G +Y ++G+ + +N +A K TW+RWVD N+D KT VFF+G SP+H+ G
Sbjct: 227 KGQSQGWDYIRDGSSLVRDMNRLDAFYKGLSTWARWVDQNVDTAKTRVFFQGISPTHYEG 286
Query: 371 GEWDSGGR-CESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDA 429
EW+ + C + +P+ S S PP + V+ MK PV L+I+ ++ R DA
Sbjct: 287 REWNEPRKTCSGQMQPLGGSSYP-SGQPPSSGVVSKVLSSMKKPVTLLDITTLSQLRKDA 345
Query: 430 HPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHL 473
HPS Y + DCSHWCLPG+PD WN+ +YA L
Sbjct: 346 HPSSYGGDGGT-----------DCSHWCLPGLPDTWNQLLYAAL 378
>UniRef100_Q9FVP8 Hypothetical protein F27K7.9 [Arabidopsis thaliana]
Length = 485
Score = 270 bits (691), Expect = 5e-71
Identities = 134/342 (39%), Positives = 200/342 (58%), Gaps = 26/342 (7%)
Query: 133 CDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKY 192
CDI++G+WV+DD++PLY + CP +++ FNC NGR ++Y K+RW+PK+C +PR +
Sbjct: 115 CDIFDGNWVVDDNYPLYNASECPFVEKGFNCLGNGRGHDEYLKWRWKPKHCTVPRFEVRD 174
Query: 193 MLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTD 252
+LK LRGKR+VFVGDS++R W+SL+C+L +++K ++E +G + F+
Sbjct: 175 VLKRLRGKRIVFVGDSMSRTQWESLICMLMTGLEDKRSVYEVNGNNITKRIRFLGVRFSS 234
Query: 253 YNCSIEFFRSPFLVQ----EWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWW 308
YN ++EF+RS FLVQ W P + K TL+LD+++ ++ +AD LIFNTG WW
Sbjct: 235 YNFTVEFYRSVFLVQPGRLRWHAPKR---VKSTLKLDVLDVINHEWSSADFLIFNTGQWW 291
Query: 309 THEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHF 368
K E Y+Q GN + +++ A A TW+ W+++ +DP KT V FR + PSH
Sbjct: 292 VPGKLFETGCYFQVGNSLRLGMSIPAAYRVALETWASWIESTVDPNKTRVLFRTFEPSH- 350
Query: 369 RGGEWDSGGRCESETEPM-KTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRH 427
W C P TE D S M I+ V+K M PV L+++ M+ FR
Sbjct: 351 ----WSDHRSCNVTKYPAPDTEGRDKSIFSEM---IKEVVKNMTIPVSILDVTSMSAFRS 403
Query: 428 DAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFV 469
D H ++ + + DCSHWCLPGVPD+WNE +
Sbjct: 404 DGHVGLWSDNPLV----------PDCSHWCLPGVPDIWNEIL 435
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.132 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 891,315,367
Number of Sequences: 2790947
Number of extensions: 39560272
Number of successful extensions: 108201
Number of sequences better than 10.0: 212
Number of HSP's better than 10.0 without gapping: 144
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 107158
Number of HSP's gapped (non-prelim): 472
length of query: 486
length of database: 848,049,833
effective HSP length: 131
effective length of query: 355
effective length of database: 482,435,776
effective search space: 171264700480
effective search space used: 171264700480
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146329.6


