

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144930.1 - phase: 0 /pseudo
(1322 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
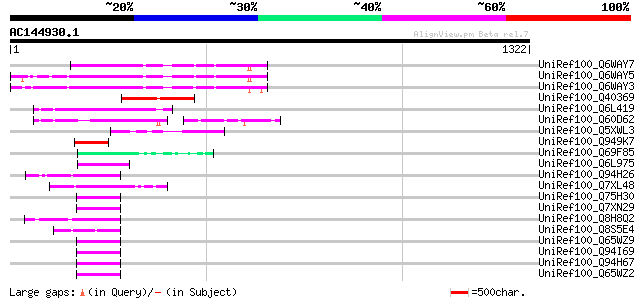
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum] 261 1e-67
UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum] 259 3e-67
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 259 3e-67
UniRef100_Q40369 RPE15 protein [Medicago sativa] 259 3e-67
UniRef100_Q6L419 Putative gag polyprotein [Solanum demissum] 149 7e-34
UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solan... 112 8e-23
UniRef100_Q5XWL3 Gag-pol polyprotein-like [Solanum tuberosum] 98 1e-18
UniRef100_Q949K7 Putative gag polyprotein [Cicer arietinum] 96 6e-18
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 71 2e-10
UniRef100_Q6L975 GAG-POL [Vitis vinifera] 69 1e-09
UniRef100_Q94H26 Putative gag-pol polyprotein [Oryza sativa] 63 5e-08
UniRef100_Q7XL48 OSJNBa0056L23.9 protein [Oryza sativa] 61 3e-07
UniRef100_Q75H30 Hypothetical protein OSJNBa0027H16.12 [Oryza sa... 60 3e-07
UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa] 60 4e-07
UniRef100_Q8H8Q2 Putative gag-pol polyprotein [Oryza sativa] 60 6e-07
UniRef100_Q8S5E4 Hypothetical protein OSJNBb0047B19.20 [Oryza sa... 60 6e-07
UniRef100_Q65WZ9 Putative polyprotein [Oryza sativa] 59 1e-06
UniRef100_Q94I69 Putative retroelement [Oryza sativa] 59 1e-06
UniRef100_Q94H67 Putative gag-pol protein [Oryza sativa] 59 1e-06
UniRef100_Q65WZ2 Putative polyprotein [Oryza sativa] 57 3e-06
>UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 261 bits (667), Expect = 1e-67
Identities = 177/528 (33%), Positives = 266/528 (49%), Gaps = 55/528 (10%)
Query: 156 NADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHC 215
N+ F ++ LV + +P KFK P FD+YNG +CP+ H+ Y RK+ Y D++ + ++
Sbjct: 148 NSLGFDVTNMGLVEGLRIPYKFKAPSFDKYNGTSCPRTHVQAYYRKISAYTDDEKMWMYF 207
Query: 216 FQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFR 275
FQDSL + +WY L ++ I + +L AF Y N + P+R L+SL QK ESF+
Sbjct: 208 FQDSLSGASLDWYMELKRDSIRCWKDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSGESFK 267
Query: 276 EYAQRWRGAAARITPALDEEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAV 335
EYAQRWR AAR+ P + E E+T F+ TL+ +++RM +FS++V G R E +
Sbjct: 268 EYAQRWRELAARVQPPMLERELTDMFIGTLQGVFMDRMGSCPFVSFSDVVICGERTESLI 327
Query: 336 RDGIIVFEKAESSVNASKRYGNGHHKKKETEIGMVSAGAGQSMATVAPINAAQMPPSYPY 395
+ G I ++ ++SK+ G +++E E V Q+ + + A +P
Sbjct: 328 KTGKI----QDAGSSSSKKPFAGAPRRREGETNAVQYRRDQNRSQRCQVAAVTIPA---- 379
Query: 396 VPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPPIPMLYA 455
P P Q Q V QQ +QQ P Q +Q R F +PM YA
Sbjct: 380 ----------------PQPRQQQQQRVQQPQQQQQQQRPYQPRQRMPDR-RFDSLPMSYA 422
Query: 456 ELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLID 515
ELLP LL G R PP LPP + ++++CDFH GA GH E C AL++ V+ LID
Sbjct: 423 ELLPELLRLGMVELRT-MAPPTVLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLID 481
Query: 516 QGKLTFENNVPHVLDNPLPNHAA--VNMIEVCE-EAPRLDVRNVATPLVPLHIKLCKASL 572
+ F VP+V++NP+P H VN IE E E ++V +V T L+ + +L +
Sbjct: 482 AKAINFA-PVPNVVNNPMPQHGGHRVNNIEGKEAEDLVVNVDDVQTSLLVVKGRLLNGGV 540
Query: 573 FSHDHAKCLGCLHNPLGCYTVQDDIQSLMNDNFL----------TVSDV----------- 611
+S CLGC + GC ++ IQ +M++ L TVS +
Sbjct: 541 YSGCDEDCLGCAESENGCDQLRTGIQGMMDEGCLQFSRAVKDRGTVSTITIYFKPSEGRG 600
Query: 612 --CVIVPVFHDPPVK-SIPLKKNAEPLVIRLPGPVPYTSDRAIPYKYN 656
V P + PV S+P+ +A P I G + RA+P+KY+
Sbjct: 601 QRVVSAPATNGTPVTISVPVTISA-PTTIAASGRRAVENSRAVPWKYD 647
>UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum]
Length = 1105
Score = 259 bits (663), Expect = 3e-67
Identities = 204/689 (29%), Positives = 320/689 (45%), Gaps = 79/689 (11%)
Query: 3 QLRTELATLREELAKANDVMTALLAAQEQ-------SATAIPIATTIPMTTSMLPTASTD 55
Q++ ELA ++ +A+ ++M + QE+ AIP P P +
Sbjct: 3 QVQAELAEMKANMAQFMNMMQGVAQGQEELRAMVQRQEAAIPPVNHAPPEGG--PVNDNN 60
Query: 56 ARFAMPAGFPYGLPPFFTPSTAAGTSGTANNVLIPATNAASINATLPQTTAAVTEPLVHT 115
A+P + + G + ++ A NA +I A A P+V
Sbjct: 61 VAAAVP------INNYDVGDELGGIRINSQPIVPDAANARTIRAP-----ARNPVPIVDR 109
Query: 116 MPQSVNINTQHGSI-PVTKTMEEMMEELAKELRHEIKANRGNADSFKTQDLCLVSKVDVP 174
+ ++ + + ++ LA+++R N+ SF ++ LV + +P
Sbjct: 110 QEDMFTLLSEDEDVLGRVDERDRKVDALAEKIR---AMECQNSLSFDVTNMGLVEGLRIP 166
Query: 175 KKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSKN 234
KFK P FD+YN +CP+ H+ Y RK+ Y D++ + ++ FQDSL + +WY L ++
Sbjct: 167 YKFKAPSFDKYNDTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKRD 226
Query: 235 DIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALDE 294
I + +L AF Y N + P+R L+SL QK ESF+EYAQRWR AAR+ P + E
Sbjct: 227 SIRCWRDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSGESFKEYAQRWRELAARVQPPMLE 286
Query: 295 EEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFEKAESSVNASKR 354
E+T F+ TL+ +++RM +FS++V G R E ++ G I + ++SK+
Sbjct: 287 RELTDMFIGTLQGVFMDRMGSCPFGSFSDVVICGERTESLIKTGKI----QDVGSSSSKK 342
Query: 355 YGNGHHKKKETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLP 414
G +++E E +V Q+ A +P P P
Sbjct: 343 PFAGAPRRREGETNVVQHRRDQNRIEYRQAAAVTIPA--------------------PQP 382
Query: 415 PGQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKP 474
Q Q V QQ +QQ P Q +Q R F +PM YAELLP LL G R
Sbjct: 383 RQQQQQRVQQPQQQQQQQRPYQPRQRMPDR-RFDSLPMSYAELLPELLRLGMVELRT-MA 440
Query: 475 PPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLP 534
PP LPP + ++++CDFH GA GH E C AL++ V+ LID + F VP+V++NP+P
Sbjct: 441 PPTVLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDAKAINFA-PVPNVVNNPMP 499
Query: 535 NHAA--VNMIEVCE-EAPRLDVRNVATPLVPLHIKLCKASLFSHDHAKCLGCLHNPLGCY 591
H VN IE E E ++V +V T L+ + +L ++S CLGC + GC
Sbjct: 500 QHGGHRVNNIEGKEAEDLVVNVDDVQTSLLVVKGRLLNGGVYSGCDEDCLGCAESENGCD 559
Query: 592 TVQDDIQSLMNDNFL----------TVSDV-------------CVIVPVFHDPPVK-SIP 627
++ IQ +M++ L TVS + V P +D PV S+P
Sbjct: 560 QLRAGIQGMMDEGCLQFSRAVKDRGTVSTITIYFKPTEGRGQRVVSAPATNDTPVTISVP 619
Query: 628 LKKNAEPLVIRLPGPVPYTSDRAIPYKYN 656
+ +A P I G + R +P+KY+
Sbjct: 620 VTISA-PTTIVASGRRAVENSRVVPWKYD 647
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 259 bits (663), Expect = 3e-67
Identities = 206/688 (29%), Positives = 316/688 (44%), Gaps = 79/688 (11%)
Query: 3 QLRTELATLREELAKANDVMTALLAAQEQSATAIPIATTIPMTTSMLPTASTDARFAMPA 62
Q++TELA +R +A+ +M + QE+ + + ++ P + A+P
Sbjct: 3 QVQTELAEMRANMAQFMHMMQGVAQGQEE------LRALVQRQEAVTPPPNQ----ALPE 52
Query: 63 GFP-YGLPPFFTPSTAAGTSGTANNVLIPATNAA--SINATLPQTTAAVTEPLVHTMPQS 119
G P + P P + + A + NA + A P+V
Sbjct: 53 GNPVHDTPAAAIPVNNYAVGEELMGIRVDGQPIAPDAANARVIHAPARNRIPIVDRQEDL 112
Query: 120 VNINTQHGSIPVTK-TMEEMMEELAKELRHEIKANRGNADSFKTQDLCLVSKVDVPKKFK 178
+ ++ IP + ++ LA+++R N+ F ++ LV + +P KFK
Sbjct: 113 FTMFSEDEDIPGRNDARDRKVDALAEKIR---AMECQNSLGFDVTNMGLVEGLRIPYKFK 169
Query: 179 IPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHT 238
P FD+YNG +CP+ H+ Y RK+ Y D++ + ++ FQDSL + +WY L + I
Sbjct: 170 APSFDKYNGTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKSDSIRC 229
Query: 239 FDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALDEEEMT 298
+ +L AF Y N + P+R L+SL QK ESF+EYAQRWR AAR+ P + E E+T
Sbjct: 230 WRDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSNESFKEYAQRWRELAARVQPPMLERELT 289
Query: 299 QTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFEKAESSVNASKRYGNG 358
F+ TL+ +++RM +FS++V G R E ++ G I + ++SK+ G
Sbjct: 290 DMFIGTLQGVFMDRMGSCPFGSFSDVVICGERTESLIKTGKI----QDVGSSSSKKPFAG 345
Query: 359 HHKKKETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQP 418
+++E E V Q+ A +P P P Q
Sbjct: 346 APRRREGETNAVQHRRDQNRIEYRQAAAVTIPA--------------------PQPRQQQ 385
Query: 419 QVPVNAIAQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRG---HCTTRQGKPP 475
Q V QQ +QQ P Q +Q R F +PM YAELLP LL G CT P
Sbjct: 386 QQRVQQ-PQQQQQQRPYQPRQRMPDR-RFDSLPMSYAELLPELLRLGLVELCT----MAP 439
Query: 476 PDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLPN 535
P LPP + ++++CDFH GA GH E C AL++ V+ LID + F VP+V++NP+P
Sbjct: 440 PTVLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDANAINFA-PVPNVVNNPMPQ 498
Query: 536 HAA--VNMIEVCEEAPRLD-VRNVATPLVPLHIKLCKASLFSHDHAKCLGCLHNPLGCYT 592
H VN IE E +D V +V T L+ + +L ++S CLGC + GC
Sbjct: 499 HGGHRVNNIEGKEAEDLVDNVDDVQTSLLVVKSRLLNEGVYSGCDEDCLGCAESENGCDQ 558
Query: 593 VQDDIQSLMNDNFLTVS-------DVCVIVPVF-----HDPPVKSIPLKKNAEPLVIRLP 640
++ IQ +M++ L S V I F H V S P N P+ I +P
Sbjct: 559 LRAGIQGMMDEGCLQFSRAVKDRGTVSTITIYFKPSEGHGQRVVSAP-ATNGTPVTIPVP 617
Query: 641 ------------GPVPYTSDRAIPYKYN 656
G + RA+P+KY+
Sbjct: 618 VTISAPTTIVASGRRAVENSRAVPWKYD 645
>UniRef100_Q40369 RPE15 protein [Medicago sativa]
Length = 219
Score = 259 bits (663), Expect = 3e-67
Identities = 129/192 (67%), Positives = 151/192 (78%), Gaps = 11/192 (5%)
Query: 284 AAARITPALDEEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFE 343
AAAR++P LDEEE+TQTFLKTLKK++ ERMI++APNNFS+MVTMGTRLEEAVR+GIIV E
Sbjct: 13 AAARVSPRLDEEELTQTFLKTLKKEHKERMIVSAPNNFSDMVTMGTRLEEAVREGIIVLE 72
Query: 344 KAESSVNASKRYGNGHHKKKET-EIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHP 402
K E S +A K+YGNGHHK+KE+ ++GMVS P+NA Q+P Y Y+ SQHP
Sbjct: 73 KGECSASAPKKYGNGHHKRKESKKVGMVSGNHD------TPVNATQLPLPYQYMQISQHP 126
Query: 403 FFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPT----FPPIPMLYAELL 458
FFPPFY QYPLPPGQP VPVNA+AQQ++ Q P QQQQ QQ P FPPIPM YAELL
Sbjct: 127 FFPPFYQQYPLPPGQPPVPVNAMAQQVQHQPPAQQQQQQQQGPRPKTYFPPIPMRYAELL 186
Query: 459 PTLLHRGHCTTR 470
PTLL +GHC TR
Sbjct: 187 PTLLAKGHCVTR 198
>UniRef100_Q6L419 Putative gag polyprotein [Solanum demissum]
Length = 421
Score = 149 bits (375), Expect = 7e-34
Identities = 104/364 (28%), Positives = 174/364 (47%), Gaps = 32/364 (8%)
Query: 61 PAGFPYGLPPFFTPSTAAGTSGTANNVLIPATNAASINATLPQTTAAVTEPLVHTMPQSV 120
P P P F P+ A T+ T+N ++P +N L E L +P S
Sbjct: 78 PLNTPMMSNPLFVPT--APTNSTSNPTVVPKSNNDPSFQVLHDHGYTPEEAL--KIPSSY 133
Query: 121 NINTQHGS-IPVTKTME----EMMEELAKELRHEIKANRG--NADSFKTQDLCLVSKVDV 173
Q+ S + KT++ E M + K L H I+ +G DLC+ V +
Sbjct: 134 PQTHQYSSPFKIEKTVKNEEHEEMTKKMKSLEHSIRDMQGLGGHKGISFSDLCMFPHVHL 193
Query: 174 PKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSK 233
P FK P+F++Y+G P H+ +Y ++ + + L++ F +SL+ A++W+
Sbjct: 194 PTGFKTPNFEKYDGHGDPIAHLKRYYNQLRGAEGKEELLMAYFGESLVGIASKWFIDQDI 253
Query: 234 NDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALD 293
+ HT+D+LA F + +N + P+R L ++ +K E+FREYA RWR AAR+ P +
Sbjct: 254 ANWHTWDDLARCFVQQFQYNIDIVPDRSSLANMRKKTTENFREYAVRWREQAARVKPPMK 313
Query: 294 EEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFEKAESSVNASK 353
E EM FL+ + DY ++ A F+E++ +G +E ++ G IV + A + +
Sbjct: 314 ELEMIDVFLQAQEPDYFHYLLSAVGKTFTEVIKVGEMVENGIKSGKIVSQAALKATTQAL 373
Query: 354 RYGNGH---HKKKETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQ 410
G+G+ K++E +VSA P +Y ++QH +FPP Q
Sbjct: 374 PNGSGNIGGKKRREDVATVVSA-----------------PRTYAQDNHTQH-YFPPQIPQ 415
Query: 411 YPLP 414
Y +P
Sbjct: 416 YLVP 419
>UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solanum demissum]
Length = 1784
Score = 112 bits (280), Expect = 8e-23
Identities = 95/367 (25%), Positives = 159/367 (42%), Gaps = 58/367 (15%)
Query: 61 PAGFPYGLPPFFTPSTAAGTSGTANNVLIPATNAASINATLPQTTAAVTEPLVHTMPQSV 120
P P P F P+ A T+ T+N ++P +N L E L +P S
Sbjct: 598 PLNTPMMSNPLFVPT--APTNSTSNPTVVPKSNNDPSFQVLHDHGYTPEEAL--KIPSSY 653
Query: 121 NINTQHGS-IPVTKTME-EMMEELAKELRHEIKANR-----GNADSFKTQDLCLVSKVDV 173
Q+ S + KT++ E EE+AK+++ ++ R G DLC+ V +
Sbjct: 654 PQTHQYSSPFKIEKTVKNEEHEEMAKKMKSLEQSIRDMQGLGGHKGISFSDLCMFPHVHL 713
Query: 174 PKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSK 233
P + + L++ F +SL+ A+EW+
Sbjct: 714 PT----------------------------GAEGKEELLMAYFGESLVGIASEWFIDQDI 745
Query: 234 NDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALD 293
+ HT+D LA F + +N + P+R L ++ +K E+FREYA RWR AAR+ ++
Sbjct: 746 ANWHTWDNLARCFVQQFQYNIDIVPDRSSLANMRKKTTENFREYAVRWREQAARVKLSMK 805
Query: 294 EEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFEKAESSVNASK 353
E EM FL+ + DY ++ A F E++ +G +E ++ G IV + A + +
Sbjct: 806 ESEMIDVFLQAQEPDYFHYLLSAVGKTFVEVIKVGEMVENGIKSGKIVSQAALKATTQAL 865
Query: 354 RYGNGH--HKKKETEIGMVSAGA-------GQSMAT----------VAPINAAQMPPSYP 394
+ G+G+ KK+ ++ V GA G+S A+ ++PI P
Sbjct: 866 QNGSGNIGGKKRREDVATVGNGAKDEFTPIGESYASLFQKLRTLNVLSPIERKMSNPPPR 925
Query: 395 YVPYSQH 401
+ YSQH
Sbjct: 926 NLDYSQH 932
Score = 67.0 bits (162), Expect = 4e-09
Identities = 69/265 (26%), Positives = 112/265 (42%), Gaps = 42/265 (15%)
Query: 443 ARPTFPPIPMLYAEL---LPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHD 499
A+ F PI YA L L TL + PPP L C + A GH+
Sbjct: 888 AKDEFTPIGESYASLFQKLRTLNVLSPIERKMSNPPPRNLD----YSQHCAYCSDAPGHN 943
Query: 500 VEGCYALKYIVKKLIDQGKLTFEN-NVPHVLDNPLPNHAAVNMIEVCEEAPRLDVRNVAT 558
+E C+ LK ++ LID ++ E+ N P++ NPLP H NM+E+ + VA
Sbjct: 944 IERCWYLKKAIQDLIDTRRIIVESPNGPNINQNPLPRHTETNMLEMVN-----NHEEVAV 998
Query: 559 PLVPLHIKL---CKASLFSHDHAKCLGCLHNPLGCYTVQD----------DIQSLMNDNF 605
P P+ +K+ ++S D K + P G + + I+ + D +
Sbjct: 999 PYKPI-LKVETGMESSTNVIDLTKIM-----PSGAESTSEKLTPSSAPILTIKGALEDVW 1052
Query: 606 LTVSDVCVIVPVFHDPPVKSIPLKKNAEPLVIRLPGPVPYTSDRAIPYKYNATIIE-NGV 664
+ + ++VP D P+ I P++IR +P T+ +A+P+ Y T++ G
Sbjct: 1053 ASQREARLVVPRGPDKPI-LIVQGAYIPPVIIRPVSQLPMTNPKAVPWNYEPTVVTYEGK 1111
Query: 665 EVPLVSLAVVNNTAEGTSAALRSGR 689
E +N + RSGR
Sbjct: 1112 E--------INEEVDEIGGMTRSGR 1128
>UniRef100_Q5XWL3 Gag-pol polyprotein-like [Solanum tuberosum]
Length = 1716
Score = 98.2 bits (243), Expect = 1e-18
Identities = 81/294 (27%), Positives = 126/294 (42%), Gaps = 72/294 (24%)
Query: 258 PNREFLRSLSQKKEESFREYAQRWRGAAARITPALDEEEMTQTFLKTLKKDYVERMIIAA 317
P+R L + QK ES+RE+A RWR AAR+ P + E+E+ + F++ K
Sbjct: 299 PDRYSLEKMKQKSTESYREFAYRWRKEAARVRPPMSEKEIVEVFVRVAK----------- 347
Query: 318 PNNFSEMVTMGTRLEEAVRDGIIVFEKA--ESSVNASKRYGNGHHKKKETEIGMVSAGAG 375
F+E+V +G +E+ +R G IV+ A ESS+ KKK ++ +S G
Sbjct: 348 ---FAEIVKIGETIEDGLRTGKIVYVAASPESSILL---------KKKRDDVSSISY-EG 394
Query: 376 QSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPV 435
+ A + + PS H F
Sbjct: 395 KKTAKKSSSCQGRSRPS--------HSSF------------------------------- 415
Query: 436 QQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGA 495
+ + AR F P+ +L L G+ G P D +R D +C +H +
Sbjct: 416 ---EKKPAR-NFTPLIKSRTKLFERLTVAGYIHP-VGPKPVDTSSKFYRPDQRCAYHSNS 470
Query: 496 LGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLPNHA--AVNMIEVCEE 547
+GHD E C LK+ ++ LI+Q ++ + P+V NPLPNH +NMIE E+
Sbjct: 471 VGHDTEDCINLKHKIQDLINQNVVSLQTVAPNVNSNPLPNHGGITINMIETDED 524
>UniRef100_Q949K7 Putative gag polyprotein [Cicer arietinum]
Length = 88
Score = 96.3 bits (238), Expect = 6e-18
Identities = 41/88 (46%), Positives = 59/88 (66%)
Query: 165 LCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDA 224
+CLV V +P+KFK+ +F++Y G TCP+NH+ Y RKM ++ ND L+IH FQDSL +
Sbjct: 1 MCLVPDVTIPRKFKVLEFEKYKGATCPKNHLTMYGRKMASHAHNDKLLIHFFQDSLSGAS 60
Query: 225 AEWYTSLSKNDIHTFDELAAAFKSHYGF 252
WY L +N IH++ +LA AF Y +
Sbjct: 61 LSWYMHLERNRIHSWKDLADAFLRQYKY 88
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 71.2 bits (173), Expect = 2e-10
Identities = 79/348 (22%), Positives = 125/348 (35%), Gaps = 76/348 (21%)
Query: 173 VPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLS 232
+P K++ + Y+G T P H+ + +M Y + ++ F SL E W++ L
Sbjct: 221 LPDKWRGLTINLYDGSTDPDEHLNIFRTQMTLYTTDRTVWCKVFPTSLREGPLGWFSDLP 280
Query: 233 KNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPAL 292
N I +FD L F + Y + + + L ++ Q++ ES R + R+ I
Sbjct: 281 PNSIASFDALELKFTTQYATSRPHRTSSMSLLNVKQERGESLRTFMNRFSKVCMNIRNLN 340
Query: 293 DEEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAVRDGIIVFEKAESSVNAS 352
E M L + E +I P N E+ T T+ F + E ++
Sbjct: 341 PEIAMHHLVSAILPGRFTESLIKRPPCNMDELRTRATK-----------FMQIEEHID-- 387
Query: 353 KRYGNGHHKKKETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQY- 411
+H+K E S G PP+ P P P +H Y
Sbjct: 388 ------YHRKTYAENTDNSKGI--------------RPPTIPTDRERHRPNRGPRFHNYT 427
Query: 412 PLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQ 471
PL VP + + L EL+PTL
Sbjct: 428 PL-----IVPRGKVLDE-----------------------ALQIELIPTL---------- 449
Query: 472 GKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQGKL 519
P PP + +C +H+ GH EGC ALK +++L+ G L
Sbjct: 450 ---RPSQTPPNADTSKRCQYHRN-YGHTTEGCQALKDKIEELVQAGHL 493
>UniRef100_Q6L975 GAG-POL [Vitis vinifera]
Length = 549
Score = 68.9 bits (167), Expect = 1e-09
Identities = 40/132 (30%), Positives = 66/132 (49%), Gaps = 1/132 (0%)
Query: 174 PKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSK 233
P+ F +P F Y+G + P +HI+ Y + M ND+L+ F SL A W+ L
Sbjct: 16 PRGFLVPKFSTYDGTSDPFDHIMHYRQLMTLDIGNDALLCKVFPASLQGQALSWFHRLPP 75
Query: 234 NDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAARITPALD 293
N + F +L+ AF Y + R K N L+++ + ES RE+ +R+ G A A
Sbjct: 76 NSVGNFRDLSEAFVGQYLCSARHKQNISTLQNIKMQDNESLREFVKRF-GQAVLQVEACS 134
Query: 294 EEEMTQTFLKTL 305
+ + Q F +++
Sbjct: 135 MDAVLQIFKRSI 146
Score = 40.4 bits (93), Expect = 0.36
Identities = 35/124 (28%), Positives = 55/124 (44%), Gaps = 19/124 (15%)
Query: 434 PVQQQQNQQARPTFPPIPML---YAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCD 490
P ++Q +RP PP+ L Y +LLP + +G + +P R RS +C
Sbjct: 211 PSDRRQEGPSRPKMPPLTPLSISYEKLLPMI--QGLSDFKWPRPIATDPSTRDRSR-RCA 267
Query: 491 FHQGALGHDVEGCYALKYIVKKLIDQGKL------------TFENNVPHVLDNPLPNHAA 538
FH+ GH E C + +Y+V++LI G L ++N P P+ A
Sbjct: 268 FHKDH-GHTTETCRSFQYLVERLIKAGHLKQYLRTDTGGRDVSQHNNPGAPRAPVATKAV 326
Query: 539 VNMI 542
+N I
Sbjct: 327 INYI 330
>UniRef100_Q94H26 Putative gag-pol polyprotein [Oryza sativa]
Length = 1538
Score = 63.2 bits (152), Expect = 5e-08
Identities = 61/246 (24%), Positives = 107/246 (42%), Gaps = 11/246 (4%)
Query: 40 TTIPMTTSMLPTASTDARFAMPAGFPYGLPPFFTPSTAAGTSGTANNVLIPATNAASINA 99
T M SM+ + + + P P G+ P + P T LIP NA S
Sbjct: 168 TLANMVKSMVDGSMAEYQATGPVYLPGGVFPNYRPII---TDNQPVVQLIPL-NAPSAQP 223
Query: 100 TLPQTTAAVTEPLVHTMPQSVNINT-QHGSIPVTKTMEEMMEELAKELRHEIKANRGNAD 158
T P +A P + Q +N Q P + +++ ++ KE + GN
Sbjct: 224 TAP-VSAHAPAPTLSAPGQVINPRLLQSQGTPQQRPWADLIADVMKEQFGLRLKDAGNL- 281
Query: 159 SFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTC--PQNHIIKYVRKMGNYKDNDSLMIHCF 216
++ +V +P +FK+PDF ++ G HI +++ + G D+L + F
Sbjct: 282 -YRQPYPEWFERVPLPNRFKVPDFSKFLGQDSISTYKHISRFLAQCGEASAVDALKVRLF 340
Query: 217 QDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFRE 276
+ SL A W++SL I+++ +L F S++ ++ + L S+ Q+ +E E
Sbjct: 341 RLSLYGSAFTWFSSLPYGSINSWADLEKQFHSYF-YSGVHEMKLSDLTSIKQRHDEPVHE 399
Query: 277 YAQRWR 282
Y QR+R
Sbjct: 400 YIQRFR 405
>UniRef100_Q7XL48 OSJNBa0056L23.9 protein [Oryza sativa]
Length = 836
Score = 60.8 bits (146), Expect = 3e-07
Identities = 72/308 (23%), Positives = 128/308 (41%), Gaps = 27/308 (8%)
Query: 102 PQTTAAVTEPLVHTMPQSVNINTQHGSIPV---TKTMEEMMEELAKELRHEIKANRGNAD 158
PQ A L+ T+ + +N + + TKTM A+ H + + + D
Sbjct: 281 PQQNAVAAITLLDTLLKENELNQSDHVVNILNQTKTMIAASRRAARGHHHSLDRHDDDLD 340
Query: 159 SFK--TQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCF 216
T DL +VD P FK ++Y+G T P++ + Y + + + +
Sbjct: 341 GVAAFTNDL---RRVDWPAGFKPTGIEKYDGSTNPESWLTVYGLAIRAAGGDSKALANYL 397
Query: 217 QDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREF-LRSLSQKKEESFR 275
+L + A W L + I ++ EL F +++ T +P +F L ++ QK ES R
Sbjct: 398 PVALADSARSWLHGLPRGTIGSWAELRDHFIANFQ-GTFERPGTQFDLYNVIQKSGESLR 456
Query: 276 EYAQRWRGAAARITPALDEEEMTQTFLKTLKKDYVERMIIAAPNNFSEMVTMGTRLEEAV 335
+Y +R+ +I+ D + + F K ++ + + P T+ E+A
Sbjct: 457 DYIRRFSEQCNKISDITD-DVIIAAFTKGIRHEELVGKFGRKPPK-----TVKLMFEKAN 510
Query: 336 RDGIIVFEKAESSVNASKRYGNGHHKKKETEIGMVSAGAGQSMATVAPINAAQ--MPPSY 393
+ KAE +V ASK+ G+ KK+T + G G + + +N M
Sbjct: 511 E-----YAKAEDAVIASKQSGSSWKPKKDTP----ATGGGGRSSKRSRVNTFDKIMNSQC 561
Query: 394 PYVPYSQH 401
P+ P S H
Sbjct: 562 PHHPKSNH 569
>UniRef100_Q75H30 Hypothetical protein OSJNBa0027H16.12 [Oryza sativa]
Length = 1782
Score = 60.5 bits (145), Expect = 3e-07
Identities = 32/115 (27%), Positives = 63/115 (53%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P +FK+PDF +++G HI +++ + G D+L + F+ SL A W
Sbjct: 464 RVPLPNRFKVPDFSKFSGQDGVSTYEHISRFLAQYGEASAVDALKVRLFKLSLSGSAFTW 523
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL I+++ +L F S++ ++ + +L ++ Q+ +E EY QR+R
Sbjct: 524 FSSLPYGSINSWADLEKQFHSYF-YSGVHEMKLSYLIAIKQRHDEPVHEYIQRFR 577
>UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa]
Length = 1563
Score = 60.1 bits (144), Expect = 4e-07
Identities = 33/115 (28%), Positives = 62/115 (53%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P +FKIPDF +++G HI +Y+ + G D+L + F+ SL A W
Sbjct: 466 RVPLPNRFKIPDFSKFSGQEGVSTYEHISRYLAQYGEASAVDALRVRMFRLSLSGSAFTW 525
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL ++++ +L F S++ ++ + L ++ Q+ +E EY QR+R
Sbjct: 526 FSSLPYGSVNSWADLEKQFHSYF-YSGVHEMKLSDLTAIKQRYDEPVHEYIQRFR 579
>UniRef100_Q8H8Q2 Putative gag-pol polyprotein [Oryza sativa]
Length = 1152
Score = 59.7 bits (143), Expect = 6e-07
Identities = 54/253 (21%), Positives = 107/253 (41%), Gaps = 23/253 (9%)
Query: 38 IATTIPMTTSMLPTASTDARFAMPAGFPYGLPPFFTPSTAAGTSGTANNVLI-----PAT 92
+ T + +T +P+ + + PA P P+T+A G N + P
Sbjct: 115 LITDVQSSTQAVPSVTPTTQPTAPASTPL-------PATSAMAPGQPMNPRLLLREHPQH 167
Query: 93 NAASINATLPQTTAAVTEPLVHTMPQSVNINTQHGSIPVTKTMEEMMEELAKELRHEIKA 152
+ N AA+ P H + Q I T ++++++ + +++
Sbjct: 168 SGQVANRPTQDLVAAMFLPPQHAVDLIQQQPIQQQPIQQTPQIQQVVQPIQQQVMQP--- 224
Query: 153 NRGNADSFKTQDLC-LVSKVDVPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDND 209
A + Q +V +P +FK+PDF +++G HI + + + G D
Sbjct: 225 ----AQPIQQQPYPEWFERVPLPNRFKVPDFSKFSGQDGVSTYEHISRLLAQCGEASAVD 280
Query: 210 SLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQK 269
+L + F+ SL A W++SL I+++ +L F S++ ++ + L ++ Q+
Sbjct: 281 ALKVRLFRLSLSGSAFTWFSSLPYGSINSWADLEKQFHSYF-YSGVHEMKLSDLTAIKQQ 339
Query: 270 KEESFREYAQRWR 282
+E EY QR+R
Sbjct: 340 HDEPVHEYIQRFR 352
>UniRef100_Q8S5E4 Hypothetical protein OSJNBb0047B19.20 [Oryza sativa]
Length = 821
Score = 59.7 bits (143), Expect = 6e-07
Identities = 40/172 (23%), Positives = 80/172 (46%), Gaps = 8/172 (4%)
Query: 113 VHTMPQSVNINTQHGSIPVTKTMEEMMEELAKELRHEIKANRGNADSFKTQDLCLVSKVD 172
V+T+ H P K +M+ ++ +E + GN ++ L +V
Sbjct: 96 VYTIDSCYKKTGVHKGAPQQKPWADMIADVMREQFGLKPKDSGNL--YRHPYLEHFERVP 153
Query: 173 VPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTS 230
+P ++K+PDF +++G H+ +++ G D+L + F A W+++
Sbjct: 154 LPNRYKVPDFSKFSGKDNVTTYEHVNRFLAHCGEVSATDALRVRLFPS---RSAFTWFSA 210
Query: 231 LSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
L N +HT+ +L F ++ ++ + L S+ QK +ES ++Y QR+R
Sbjct: 211 LPYNFVHTWADLERQFHKYF-YSGIHQMKLSDLTSIKQKHDESVQDYIQRFR 261
>UniRef100_Q65WZ9 Putative polyprotein [Oryza sativa]
Length = 1316
Score = 58.9 bits (141), Expect = 1e-06
Identities = 31/115 (26%), Positives = 63/115 (53%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P +K+PDF +++G HI +++ + G D+L + F SL A W
Sbjct: 227 RVPLPSGYKVPDFSKFSGQDNVSTYEHISRFLAQSGEASTIDALRVRLFPLSLSGSAFTW 286
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL N ++++ +L F S++ ++ + L ++ Q+ +ES ++Y QR+R
Sbjct: 287 FSSLPYNSVNSWADLEKQFHSYF-YSGIHEMKLSDLMAIRQRHDESMQDYIQRFR 340
>UniRef100_Q94I69 Putative retroelement [Oryza sativa]
Length = 2014
Score = 58.9 bits (141), Expect = 1e-06
Identities = 33/115 (28%), Positives = 61/115 (52%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYN--GLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P +FK+PDF +++ T HI +++ + G D+L + F+ SL W
Sbjct: 400 RVPLPNRFKVPDFSKFSWQDSTSTYEHISRFLAQCGEASAVDALKVRLFRLSLAGSTFTW 459
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL I+++ +L F S++ ++ + L S+ QK +E EY QR+R
Sbjct: 460 FSSLPYGSINSWADLEKQFHSYF-YSGIHEMKLSDLTSIKQKHDEPVHEYIQRFR 513
>UniRef100_Q94H67 Putative gag-pol protein [Oryza sativa]
Length = 2273
Score = 58.9 bits (141), Expect = 1e-06
Identities = 33/115 (28%), Positives = 61/115 (52%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYN--GLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P +FK+PDF +++ T HI +++ + G D+L + F+ SL W
Sbjct: 653 RVPLPNRFKVPDFSKFSWQDSTSTYEHISRFLAQCGEASAVDALKVRLFRLSLAGSTFTW 712
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL I+++ +L F S++ ++ + L S+ QK +E EY QR+R
Sbjct: 713 FSSLPYGSINSWADLEKQFHSYF-YSGIHEMKLSDLTSIKQKHDEPVHEYIQRFR 766
>UniRef100_Q65WZ2 Putative polyprotein [Oryza sativa]
Length = 1490
Score = 57.4 bits (137), Expect = 3e-06
Identities = 30/115 (26%), Positives = 64/115 (55%), Gaps = 3/115 (2%)
Query: 170 KVDVPKKFKIPDFDRYNGL--TCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEW 227
+V +P ++K+PDF +++G HI +++ + G D+L + F SL A W
Sbjct: 245 RVPLPSRYKVPDFSKFSGQDNVSTYEHISRFLAQCGEASAIDTLRVRLFPLSLSGSAFTW 304
Query: 228 YTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 282
++SL N ++++ +L F +++ ++ + L ++ Q+ +ES ++Y QR+R
Sbjct: 305 FSSLPYNSVNSWADLEKQFHNYF-YSGIHEMKLLDLTAIRQRHDESVQDYIQRFR 358
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.143 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,174,283,827
Number of Sequences: 2790947
Number of extensions: 93246459
Number of successful extensions: 336343
Number of sequences better than 10.0: 835
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 795
Number of HSP's that attempted gapping in prelim test: 331373
Number of HSP's gapped (non-prelim): 3384
length of query: 1322
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1182
effective length of database: 457,317,253
effective search space: 540548993046
effective search space used: 540548993046
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 81 (35.8 bits)
Medicago: description of AC144930.1


