

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144929.16 + phase: 0
(143 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
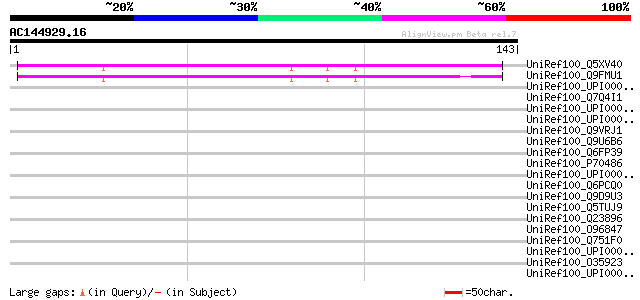
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q5XV40 Hypothetical protein [Arabidopsis thaliana] 63 1e-09
UniRef100_Q9FMU1 Arabidopsis thaliana genomic DNA, chromosome 5,... 63 1e-09
UniRef100_UPI0000364C1F UPI0000364C1F UniRef100 entry 36 0.21
UniRef100_Q7Q4I1 ENSANGP00000019222 [Anopheles gambiae str. PEST] 35 0.46
UniRef100_UPI000046C4A1 UPI000046C4A1 UniRef100 entry 34 0.78
UniRef100_UPI000023F703 UPI000023F703 UniRef100 entry 33 1.7
UniRef100_Q9VRJ1 CG4633-PA [Drosophila melanogaster] 32 2.3
UniRef100_Q9U6B6 Alanyl-tRNA synthetase [Drosophila melanogaster] 32 2.3
UniRef100_Q6FP39 Candida glabrata strain CBS138 chromosome J com... 32 2.3
UniRef100_P70486 Transcriptional regulator ATRX [Rattus norvegicus] 32 2.3
UniRef100_UPI00002554BB UPI00002554BB UniRef100 entry 32 3.0
UniRef100_Q6PCQ0 IQ motif containing E [Mus musculus] 32 3.0
UniRef100_Q9D9U3 Mus musculus adult male testis cDNA, RIKEN full... 32 3.0
UniRef100_Q5TUJ9 ENSANGP00000029120 [Anopheles gambiae str. PEST] 32 3.0
UniRef100_Q23896 Hypothetical protein [Dictyostelium discoideum] 32 3.0
UniRef100_O96847 Slime mold (D.discoideum) transposon DIRS-1, co... 32 3.0
UniRef100_Q751F0 AGL244Cp [Ashbya gossypii] 32 3.0
UniRef100_UPI00002F10CC UPI00002F10CC UniRef100 entry 32 3.9
UniRef100_O35923 Breast cancer type 2 susceptibility protein hom... 32 3.9
UniRef100_UPI000042DBDC UPI000042DBDC UniRef100 entry 31 5.1
>UniRef100_Q5XV40 Hypothetical protein [Arabidopsis thaliana]
Length = 358
Score = 63.2 bits (152), Expect = 1e-09
Identities = 50/159 (31%), Positives = 79/159 (49%), Gaps = 22/159 (13%)
Query: 3 DAAVTETQLKLISYELEKFVEAD-EECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKV 61
DA VTE LKLIS EL+KF+EA+ +E ++ SGR S + + + + +G++ E+ +
Sbjct: 123 DADVTENDLKLISNELDKFLEAEAKEGHHQPSGRNSDTNTIASTIEAIEGVDDEEDNQPM 182
Query: 62 ICPLQEYLLGSSFEIRE-KTEVRIERAS------VRETQVKQ--------------GRRS 100
PLQEY GS E+ E K + +RAS + E Q KQ +S
Sbjct: 183 KFPLQEYFFGSLIELPESKIAGKKDRASLGELFQITEVQDKQSENIYGKKKKQPNSAHKS 242
Query: 101 ALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKV 139
A H++KK+ + SS+ + D +K+ +V
Sbjct: 243 AKHLVKKVLKKIHPSSRGSVSGKPEVDSTKKKFQKMVQV 281
>UniRef100_Q9FMU1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MUA22
[Arabidopsis thaliana]
Length = 361
Score = 63.2 bits (152), Expect = 1e-09
Identities = 53/159 (33%), Positives = 80/159 (49%), Gaps = 25/159 (15%)
Query: 3 DAAVTETQLKLISYELEKFVEAD-EECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKV 61
DA VTE LKLIS EL+KF+EA+ +E ++ SGR S + + + + +G++ E+ +
Sbjct: 126 DADVTENDLKLISNELDKFLEAEAKEGHHQPSGRNSDTNTIASTIEAIEGVDDEEDNQPM 185
Query: 62 ICPLQEYLLGSSFEIRE-KTEVRIERAS------VRETQVKQ--------------GRRS 100
PLQEY GS E+ E K + +RAS + E Q KQ +S
Sbjct: 186 KFPLQEYFFGSLIELPESKIAGKKDRASLGELFQITEVQDKQSENIYGKKKKQPNSAHKS 245
Query: 101 ALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKV 139
A H++KK+ + SS+ + D ST +K KV
Sbjct: 246 AKHLVKKVLKKIHPSSRGSVSGKPEVD---STKKKFQKV 281
>UniRef100_UPI0000364C1F UPI0000364C1F UniRef100 entry
Length = 248
Score = 35.8 bits (81), Expect = 0.21
Identities = 32/129 (24%), Positives = 56/129 (42%), Gaps = 10/129 (7%)
Query: 8 ETQLKLISYELEKFVEADEECF--YESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPL 65
E +L+L+ Y+L + + EE YE + R R L R + ++E C +
Sbjct: 102 EEKLELLEYQLNEAKKIAEEADLKYEENARKLTRVEGELERAEDRAAKSESK-----CRI 156
Query: 66 QEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKS---CNTY 122
E L S F E ++E+ SV+E + ++ R+ +KK + + +S C
Sbjct: 157 LEEELKSVFSTSRSLEAQVEKYSVKEDKYEEEIRNLTDKVKKAESRAEEAERSRDNCERT 216
Query: 123 GNTADHATS 131
N + A S
Sbjct: 217 INNLEEAVS 225
>UniRef100_Q7Q4I1 ENSANGP00000019222 [Anopheles gambiae str. PEST]
Length = 243
Score = 34.7 bits (78), Expect = 0.46
Identities = 19/77 (24%), Positives = 36/77 (46%), Gaps = 2/77 (2%)
Query: 52 LETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNM 111
LET+ + C + + + SF + V+ E + +T + +G R LH I ++
Sbjct: 71 LETD--AGQYTCSVPQLGVSKSFNVVANVVVKFESTEIGKTNIVEGERLTLHCIAYGTDP 128
Query: 112 VLSSSKSCNTYGNTADH 128
++ + NTY + DH
Sbjct: 129 KITWTVGNNTYNASTDH 145
>UniRef100_UPI000046C4A1 UPI000046C4A1 UniRef100 entry
Length = 473
Score = 33.9 bits (76), Expect = 0.78
Identities = 25/97 (25%), Positives = 45/97 (45%), Gaps = 6/97 (6%)
Query: 27 ECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLL---GSSFEIREKTEVR 83
+ FYE IS N+ + + + GNK+I L+ L G+SF I +K +++
Sbjct: 180 QAFYEYENYISNNENIYICELRIYNMTCNSCGNKIINFLKNKNLIIDGNSFAIDDKIKLK 239
Query: 84 I---ERASVRETQVKQGRRSALHIIKKMSNMVLSSSK 117
I E S + + G ++ + +K N ++S K
Sbjct: 240 INIPENMSNYKNDISDGIKNKSNNVKFYVNKIMSEIK 276
>UniRef100_UPI000023F703 UPI000023F703 UniRef100 entry
Length = 7791
Score = 32.7 bits (73), Expect = 1.7
Identities = 20/57 (35%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Query: 81 EVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGN--TADHATSTNEK 135
E+ + + ETQ K+ R H+IK+++N V S S SC N A+ S N+K
Sbjct: 4511 EIHFDAKLLEETQAKRLIRQLAHVIKQLANSVPSLSLSCIDMMNPDDAEEIKSWNKK 4567
>UniRef100_Q9VRJ1 CG4633-PA [Drosophila melanogaster]
Length = 1012
Score = 32.3 bits (72), Expect = 2.3
Identities = 39/136 (28%), Positives = 64/136 (46%), Gaps = 23/136 (16%)
Query: 5 AVTETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVIC- 63
A + K ++Y++ V +D+ C E G + R +KTD EDL N+VIC
Sbjct: 664 AAIRSLFKKVTYQVSSSVSSDQ-CKLEL-GLLGKRI------QKTDVQLIEDLINRVICS 715
Query: 64 --PLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNT 121
P++ LL S+ E+ E+ ++ + E +QG R + + + LSS + C
Sbjct: 716 AAPVEVQLL-SAAEVLEQNDITMVPG---EVYPEQGLRL---VNVESPELQLSSKELC-- 766
Query: 122 YGNTADHATSTNEKLC 137
HAT+T+E C
Sbjct: 767 ---CGTHATNTSELSC 779
>UniRef100_Q9U6B6 Alanyl-tRNA synthetase [Drosophila melanogaster]
Length = 1012
Score = 32.3 bits (72), Expect = 2.3
Identities = 39/136 (28%), Positives = 64/136 (46%), Gaps = 23/136 (16%)
Query: 5 AVTETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVIC- 63
A + K ++Y++ V +D+ C E G + R +KTD EDL N+VIC
Sbjct: 664 AAIRSLFKKVTYQVSSSVSSDQ-CKLEL-GLLGKRI------QKTDVQLIEDLINRVICS 715
Query: 64 --PLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNT 121
P++ LL S+ E+ E+ ++ + E +QG R + + + LSS + C
Sbjct: 716 AAPVEVQLL-SAAEVLEQNDITMVPG---EVYPEQGLRL---VNVESPELQLSSKELC-- 766
Query: 122 YGNTADHATSTNEKLC 137
HAT+T+E C
Sbjct: 767 ---CGTHATNTSELSC 779
>UniRef100_Q6FP39 Candida glabrata strain CBS138 chromosome J complete sequence
[Candida glabrata]
Length = 512
Score = 32.3 bits (72), Expect = 2.3
Identities = 24/105 (22%), Positives = 50/105 (46%), Gaps = 4/105 (3%)
Query: 36 ISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVK 95
IS N+ +S + + + +K + +++ ++E+ + +E+ RET
Sbjct: 344 ISSSLNIKISGYRLQPIGSTSKVSKFLADKEDW---ETYELYFSSNFSLEKIFKRETDFD 400
Query: 96 QGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKVG 140
+G S L I+K++S + S+ S +T N D T +N+ + G
Sbjct: 401 KGMESLLDIVKQIS-LSFSTVVSSDTDINIQDERTRSNDSMLNGG 444
>UniRef100_P70486 Transcriptional regulator ATRX [Rattus norvegicus]
Length = 527
Score = 32.3 bits (72), Expect = 2.3
Identities = 27/123 (21%), Positives = 51/123 (40%), Gaps = 10/123 (8%)
Query: 14 ISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSS 73
+S +++ E C +SS R+ V+L KK L + G + C +
Sbjct: 161 LSDSVDRLPVKGESC--DSSEDKKTRNRVSLREKKQFSLPAKSSGKRPECSSSDTERSVK 218
Query: 74 FEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTN 133
E + T+ R++R +RE + +RS + V S S S + G++ D
Sbjct: 219 GECCDSTDKRVKRIDLRERRSSNSKRS--------TKEVKSGSSSSDAEGSSEDAKKQKK 270
Query: 134 EKL 136
+++
Sbjct: 271 QRM 273
>UniRef100_UPI00002554BB UPI00002554BB UniRef100 entry
Length = 405
Score = 32.0 bits (71), Expect = 3.0
Identities = 24/95 (25%), Positives = 49/95 (51%), Gaps = 20/95 (21%)
Query: 33 SGRISFRSNVTLSR------KKTDGLETEDLGNKVICPLQEYL--LGSSFE--------- 75
SG I+F SN+TLS+ K +D ++ N I PL++ + +GSS+E
Sbjct: 250 SGLINFSSNLTLSKGQLAAAKISDSIQMPINSNGYILPLKDEVNWIGSSYENQFQNMDVN 309
Query: 76 ---IREKTEVRIERASVRETQVKQGRRSALHIIKK 107
++E E + ++ +++ + + G ++ + +I K
Sbjct: 310 KSKLQEMIEFQCDQFNLQNAENECGSKTQIRVISK 344
>UniRef100_Q6PCQ0 IQ motif containing E [Mus musculus]
Length = 778
Score = 32.0 bits (71), Expect = 3.0
Identities = 24/98 (24%), Positives = 47/98 (47%), Gaps = 6/98 (6%)
Query: 17 ELEKFVEADEECFYESSGRISFR------SNVTLSRKKTDGLETEDLGNKVICPLQEYLL 70
ELEK V + E +S ++ SN+++ ++ EDL C QE+L
Sbjct: 345 ELEKKVSSSESPKQSTSELVNPNPLVRSPSNISVQKQPKGDQSPEDLPKVAPCEEQEHLQ 404
Query: 71 GSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKM 108
G+ +RE+ E+ ++ ++KQ +S + + K++
Sbjct: 405 GTVKSLREELGALQEQLLEKDLEMKQLLQSKIDLEKEL 442
>UniRef100_Q9D9U3 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:1700028P05 product:hypothetical IQ
calmodulin-binding motif containing protein, full insert
sequence [Mus musculus]
Length = 761
Score = 32.0 bits (71), Expect = 3.0
Identities = 24/98 (24%), Positives = 47/98 (47%), Gaps = 6/98 (6%)
Query: 17 ELEKFVEADEECFYESSGRISFR------SNVTLSRKKTDGLETEDLGNKVICPLQEYLL 70
ELEK V + E +S ++ SN+++ ++ EDL C QE+L
Sbjct: 300 ELEKKVSSSESPKQSTSELVNPNPLVRSPSNISVQKQPKGDQSPEDLPKVAPCEEQEHLQ 359
Query: 71 GSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKM 108
G+ +RE+ E+ ++ ++KQ +S + + K++
Sbjct: 360 GTVKSLREELGALQEQLLEKDLEMKQLLQSKIDLEKEL 397
>UniRef100_Q5TUJ9 ENSANGP00000029120 [Anopheles gambiae str. PEST]
Length = 1092
Score = 32.0 bits (71), Expect = 3.0
Identities = 15/55 (27%), Positives = 27/55 (48%)
Query: 77 REKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATS 131
R + +++ ++R Q RR+ H +K LS+ +S +T+ DHA S
Sbjct: 49 RNRRLALVKKRAIRLVQTFAARRAIPHYFQKRVKHTLSAKQSASTHNGATDHAPS 103
>UniRef100_Q23896 Hypothetical protein [Dictyostelium discoideum]
Length = 335
Score = 32.0 bits (71), Expect = 3.0
Identities = 22/97 (22%), Positives = 46/97 (46%)
Query: 36 ISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVK 95
++ + +LSR + + + E G++V+ P++ F+ E VR S+R+
Sbjct: 201 LAVNAQASLSRVRRNNIAKEIYGSEVLLPIKIKDTPKMFDETETERVRKLAKSIRKNNEA 260
Query: 96 QGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATST 132
+ L+ K + L +S NT GN+++ +S+
Sbjct: 261 KQSLLKLNYHSKSNVKKLVNSSGNNTTGNSSNSKSSS 297
>UniRef100_O96847 Slime mold (D.discoideum) transposon DIRS-1, complete, clone SB41
[Dictyostelium discoideum]
Length = 335
Score = 32.0 bits (71), Expect = 3.0
Identities = 22/97 (22%), Positives = 46/97 (46%)
Query: 36 ISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVK 95
++ + +LSR + + + E G++V+ P++ F+ E VR S+R+
Sbjct: 201 LAVNTQASLSRVRRNNIAKEIYGSEVLLPIKIKDTPKMFDETETERVRKLAKSIRKNNEA 260
Query: 96 QGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATST 132
+ L+ K + L +S NT GN+++ +S+
Sbjct: 261 KQSLLKLNYHSKSNVKKLVNSSGNNTTGNSSNSKSSS 297
>UniRef100_Q751F0 AGL244Cp [Ashbya gossypii]
Length = 842
Score = 32.0 bits (71), Expect = 3.0
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 14/84 (16%)
Query: 10 QLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYL 69
++ ++Y+L V+A + F ESSG + V +S + D LQ
Sbjct: 87 EMSSLAYQLNTLVDATTKIFVESSGFVLLEPFVKMSVEFAD--------------LQVLS 132
Query: 70 LGSSFEIREKTEVRIERASVRETQ 93
+ S ++IR+ E E+ + RE Q
Sbjct: 133 IVSDYDIRQTGETTFEQFNARERQ 156
>UniRef100_UPI00002F10CC UPI00002F10CC UniRef100 entry
Length = 198
Score = 31.6 bits (70), Expect = 3.9
Identities = 26/110 (23%), Positives = 51/110 (45%), Gaps = 4/110 (3%)
Query: 2 KDAAVTETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKV 61
K + E L+ + +++K + +E SSG I+ N T +KK +E + +
Sbjct: 21 KKGKLKEKWLRSLVKDIDKLRKKGKEFVIVSSGAIALGQNYTKVKKKRIKIE---MSQAL 77
Query: 62 ICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNM 111
Q YL EI +K +++I + + +Q RR A+++ + N+
Sbjct: 78 ASIGQIYLANIYREIFQKKKIKIGQILISPDDTEQ-RRRAINVKRTFENL 126
>UniRef100_O35923 Breast cancer type 2 susceptibility protein homolog [Rattus
norvegicus]
Length = 3343
Score = 31.6 bits (70), Expect = 3.9
Identities = 19/85 (22%), Positives = 43/85 (50%), Gaps = 5/85 (5%)
Query: 38 FRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQ---- 93
F+SN +L K + D ++ + PL + LG SF E+++ +V++++
Sbjct: 955 FKSNSSLYLKSDGNNDYLDKWSEFLDPLMNHKLGGSFRTASNKEIKLSEDNVKKSKMFFK 1014
Query: 94 -VKQGRRSALHIIKKMSNMVLSSSK 117
+++ ++L I +S + L++ K
Sbjct: 1015 DIEEQYPTSLDCIDTVSTLQLANKK 1039
>UniRef100_UPI000042DBDC UPI000042DBDC UniRef100 entry
Length = 1176
Score = 31.2 bits (69), Expect = 5.1
Identities = 28/117 (23%), Positives = 44/117 (36%), Gaps = 4/117 (3%)
Query: 22 VEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTE 81
V + EC SS IS S VT S + +E + + + ++ SS E
Sbjct: 606 VSSSTECISSSSEAISSSSQVTSSSTECISSSSEVISSSEVTSCSSEVVSSS----ETCI 661
Query: 82 VRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCK 138
E +S + + S + K S SS+SC + + TS+ CK
Sbjct: 662 SSKEMSSSEQISSSESTSSCSEFVSKSSEHSSLSSESCPSEETSTVSETSSETVTCK 718
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.128 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 205,651,324
Number of Sequences: 2790947
Number of extensions: 7038084
Number of successful extensions: 14392
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 27
Number of HSP's that attempted gapping in prelim test: 14375
Number of HSP's gapped (non-prelim): 40
length of query: 143
length of database: 848,049,833
effective HSP length: 119
effective length of query: 24
effective length of database: 515,927,140
effective search space: 12382251360
effective search space used: 12382251360
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144929.16


