

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
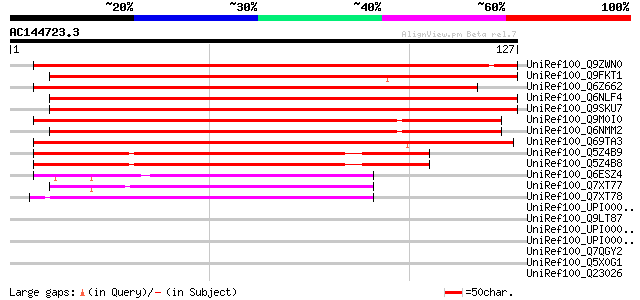
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZWN0 GPI-anchored protein [Phaseolus aureus] 218 3e-56
UniRef100_Q9FKT1 Similarity to GPI-anchored protein [Arabidopsis... 169 1e-41
UniRef100_Q6Z662 Hypothetical protein P0654B04.18 [Oryza sativa] 154 6e-37
UniRef100_Q6NLF4 At2g20700 [Arabidopsis thaliana] 145 2e-34
UniRef100_Q9SKU7 Hypothetical protein At2g20700 [Arabidopsis tha... 145 2e-34
UniRef100_Q9M0I0 Putative GPI-anchored protein [Arabidopsis thal... 142 2e-33
UniRef100_Q6NMM2 At4g28280 [Arabidopsis thaliana] 140 8e-33
UniRef100_Q69TA3 Putative GPI-anchored protein [Oryza sativa] 125 2e-28
UniRef100_Q5Z4B9 Putative GPI-anchored protein [Oryza sativa] 93 2e-18
UniRef100_Q5Z4B8 Putative GPI-anchored protein [Oryza sativa] 93 2e-18
UniRef100_Q6ESZ4 Hypothetical protein P0472F10.10 [Oryza sativa] 77 7e-14
UniRef100_Q7XT77 OSJNBa0029H02.12 protein [Oryza sativa] 67 9e-11
UniRef100_Q7XT78 OSJNBa0029H02.11 protein [Oryza sativa] 66 2e-10
UniRef100_UPI00003AE287 UPI00003AE287 UniRef100 entry 35 0.50
UniRef100_Q9LT87 Similarity to serine/threonine kinase [Arabidop... 35 0.50
UniRef100_UPI0000361097 UPI0000361097 UniRef100 entry 33 1.4
UniRef100_UPI00003384D7 UPI00003384D7 UniRef100 entry 33 1.4
UniRef100_Q7QGY2 ENSANGP00000012454 [Anopheles gambiae str. PEST] 32 2.5
UniRef100_Q5X0G1 SdbB protein [Legionella pneumophila str. Lens] 32 4.2
UniRef100_Q23026 Hypothetical protein R09F10.3 [Caenorhabditis e... 31 5.5
>UniRef100_Q9ZWN0 GPI-anchored protein [Phaseolus aureus]
Length = 169
Score = 218 bits (554), Expect = 3e-56
Identities = 100/121 (82%), Positives = 110/121 (90%), Gaps = 1/121 (0%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GCSVNFEFLNYTIITSKCKGP+YPPKECCG+FKEFACPYADV+NDLTN+CASTMFSY
Sbjct: 50 AKKGCSVNFEFLNYTIITSKCKGPQYPPKECCGAFKEFACPYADVLNDLTNECASTMFSY 109
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL 126
INLYG+YPPGLFASEC + K GL C ALPPSVSADDT+NQ +H PSL+L+LT C FLILL
Sbjct: 110 INLYGKYPPGLFASECHDTKRGLECPALPPSVSADDTSNQFLHCPSLLLLLTTC-FLILL 168
Query: 127 F 127
F
Sbjct: 169 F 169
>UniRef100_Q9FKT1 Similarity to GPI-anchored protein [Arabidopsis thaliana]
Length = 168
Score = 169 bits (428), Expect = 1e-41
Identities = 77/120 (64%), Positives = 93/120 (77%), Gaps = 3/120 (2%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C VNFEF+NYTIITSKCKGPKYPPKECCG+FK+FACPY D +NDL++DCA+TMFSYINLY
Sbjct: 49 CPVNFEFMNYTIITSKCKGPKYPPKECCGAFKDFACPYTDQLNDLSSDCATTMFSYINLY 108
Query: 71 GRYPPGLFASECREGKEGLACDA---LPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
G+YPPGLFA++C+EGKEGL C A LPP SA+ A + + V A + + LF
Sbjct: 109 GKYPPGLFANQCKEGKEGLECPAGSQLPPETSAEVNAATTSSSRLWLTVSAALLVFVKLF 168
>UniRef100_Q6Z662 Hypothetical protein P0654B04.18 [Oryza sativa]
Length = 167
Score = 154 bits (388), Expect = 6e-37
Identities = 68/111 (61%), Positives = 84/111 (75%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A C VNFEF NYTIITSKCKGP++P K+CC +FKEFACP+ + IND +NDCASTMFSY
Sbjct: 46 AKKNCPVNFEFQNYTIITSKCKGPRFPAKQCCDAFKEFACPFNEYINDESNDCASTMFSY 105
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVL 117
INLYG+YPPGLFA+ECREGK GL+C+ + S +A Q + L ++
Sbjct: 106 INLYGKYPPGLFANECREGKLGLSCEGVSQKDSVVSSAGQQAQSSLLAFIM 156
>UniRef100_Q6NLF4 At2g20700 [Arabidopsis thaliana]
Length = 163
Score = 145 bits (366), Expect = 2e-34
Identities = 67/117 (57%), Positives = 83/117 (70%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C +F NYTIITS+CKGP YP CC +FK+FACP+A+V+ND NDCASTMFSYINLY
Sbjct: 44 CKEDFANKNYTIITSRCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYINLY 103
Query: 71 GRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
GRYPPG+FA+ C+EGKEGL C + S SA + T + + VL+ + L LLF
Sbjct: 104 GRYPPGIFANMCKEGKEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 160
>UniRef100_Q9SKU7 Hypothetical protein At2g20700 [Arabidopsis thaliana]
Length = 181
Score = 145 bits (366), Expect = 2e-34
Identities = 67/117 (57%), Positives = 83/117 (70%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C +F NYTIITS+CKGP YP CC +FK+FACP+A+V+ND NDCASTMFSYINLY
Sbjct: 62 CKEDFANKNYTIITSRCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYINLY 121
Query: 71 GRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
GRYPPG+FA+ C+EGKEGL C + S SA + T + + VL+ + L LLF
Sbjct: 122 GRYPPGIFANMCKEGKEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 178
>UniRef100_Q9M0I0 Putative GPI-anchored protein [Arabidopsis thaliana]
Length = 160
Score = 142 bits (357), Expect = 2e-33
Identities = 66/117 (56%), Positives = 84/117 (71%), Gaps = 1/117 (0%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A A C +F NYTIITSKCKGP YP K CC +FK+FACP+A+V+ND DCASTMFSY
Sbjct: 44 AKATCKEDFAAKNYTIITSKCKGPNYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSY 103
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFL 123
INLYGRYPPG+FA+ C+EGKEGL C + P+ S+ + +V T L++ ++ L
Sbjct: 104 INLYGRYPPGIFANMCKEGKEGLDCTDVTPT-SSSHASIPLVSTHVLLITVSILFHL 159
>UniRef100_Q6NMM2 At4g28280 [Arabidopsis thaliana]
Length = 134
Score = 140 bits (352), Expect = 8e-33
Identities = 64/113 (56%), Positives = 82/113 (71%), Gaps = 1/113 (0%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C +F NYTIITSKCKGP YP K CC +FK+FACP+A+V+ND DCASTMFSYINLY
Sbjct: 22 CKEDFAAKNYTIITSKCKGPNYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSYINLY 81
Query: 71 GRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFL 123
GRYPPG+FA+ C+EGKEGL C + P+ S+ + +V T L++ ++ L
Sbjct: 82 GRYPPGIFANMCKEGKEGLDCTDVTPT-SSSHASIPLVSTHVLLITVSILFHL 133
>UniRef100_Q69TA3 Putative GPI-anchored protein [Oryza sativa]
Length = 174
Score = 125 bits (314), Expect = 2e-28
Identities = 60/125 (48%), Positives = 77/125 (61%), Gaps = 5/125 (4%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A C VNFE NYT+ITS+CKGP YPP CC + K+ ACP+ IND CA++MFSY
Sbjct: 48 AKKDCPVNFEEANYTVITSRCKGPMYPPALCCQALKDLACPFTAYINDAQTTCAASMFSY 107
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSV-----SADDTANQIVHTPSLVLVLTACI 121
INLYG+YPPGLFA+ C+EG GL C P + A +A IV + ++
Sbjct: 108 INLYGKYPPGLFANTCKEGANGLECPEDTPQMKPGEDKAASSAAAIVAAVARPVLAAVSA 167
Query: 122 FLILL 126
FL+L+
Sbjct: 168 FLMLI 172
>UniRef100_Q5Z4B9 Putative GPI-anchored protein [Oryza sativa]
Length = 163
Score = 92.8 bits (229), Expect = 2e-18
Identities = 45/99 (45%), Positives = 62/99 (62%), Gaps = 5/99 (5%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GC V+FE NYT ITSKCK P +P CC + EFAC ++ IND + +CA +M+ Y
Sbjct: 39 AKGGCPVSFENQNYTTITSKCKSP-WPADLCCPALNEFACNFSQYINDESTNCAESMWVY 97
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTAN 105
+N +G YP GLF++EC L C+ ++S + TAN
Sbjct: 98 LNAHGSYPAGLFSNECAV----LDCNGNNSTISTNQTAN 132
>UniRef100_Q5Z4B8 Putative GPI-anchored protein [Oryza sativa]
Length = 137
Score = 92.8 bits (229), Expect = 2e-18
Identities = 45/99 (45%), Positives = 62/99 (62%), Gaps = 5/99 (5%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GC V+FE NYT ITSKCK P +P CC + EFAC ++ IND + +CA +M+ Y
Sbjct: 13 AKGGCPVSFENQNYTTITSKCKSP-WPADLCCPALNEFACNFSQYINDESTNCAESMWVY 71
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTAN 105
+N +G YP GLF++EC L C+ ++S + TAN
Sbjct: 72 LNAHGSYPAGLFSNECAV----LDCNGNNSTISTNQTAN 106
>UniRef100_Q6ESZ4 Hypothetical protein P0472F10.10 [Oryza sativa]
Length = 147
Score = 77.4 bits (189), Expect = 7e-14
Identities = 38/88 (43%), Positives = 50/88 (56%), Gaps = 5/88 (5%)
Query: 7 APAG-CSVNFEFLN--YTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTM 63
AP G C V F+ + +T + KCK ECC +FKE ACP+ ++NDL N C M
Sbjct: 62 APLGMCPVRFDEMKGPFTELGKKCKAASVT--ECCDAFKEIACPHNTLLNDLNNGCGDDM 119
Query: 64 FSYINLYGRYPPGLFASECREGKEGLAC 91
F +I+ YGR PPG +C EG G+ C
Sbjct: 120 FYFIHTYGRLPPGTIFKKCVEGPYGMKC 147
>UniRef100_Q7XT77 OSJNBa0029H02.12 protein [Oryza sativa]
Length = 150
Score = 67.0 bits (162), Expect = 9e-11
Identities = 33/82 (40%), Positives = 45/82 (54%), Gaps = 2/82 (2%)
Query: 11 CSVNFEFLN-YTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINL 69
C V FE + + + +KC K KECC FK+ ACPY ++ND+TN CA+ F I+
Sbjct: 70 CPVEFEQVKGFGELGAKCND-KQTMKECCELFKKIACPYNHLLNDITNVCANEFFYLIHT 128
Query: 70 YGRYPPGLFASECREGKEGLAC 91
G+ PG C EG G+ C
Sbjct: 129 KGKLQPGTILENCNEGPMGINC 150
>UniRef100_Q7XT78 OSJNBa0029H02.11 protein [Oryza sativa]
Length = 149
Score = 66.2 bits (160), Expect = 2e-10
Identities = 34/86 (39%), Positives = 45/86 (51%), Gaps = 1/86 (1%)
Query: 6 IAPAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFS 65
+AP C V F+ + I K K CC +FK FACP+ +IND+ N CA MF
Sbjct: 65 LAPV-CPVRFDKMKGPAIELGKKCKTTGVKVCCEAFKTFACPHNKLINDVNNGCADEMFY 123
Query: 66 YINLYGRYPPGLFASECREGKEGLAC 91
I+ YG+ PG +C EG G+ C
Sbjct: 124 TIHTYGQLLPGTIFKKCLEGPHGMKC 149
>UniRef100_UPI00003AE287 UPI00003AE287 UniRef100 entry
Length = 365
Score = 34.7 bits (78), Expect = 0.50
Identities = 29/101 (28%), Positives = 41/101 (39%), Gaps = 11/101 (10%)
Query: 1 MTLFHIAPAGC--SVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTND 58
+T + +PAG S N + N + + GP P K P+A V N L +
Sbjct: 75 ITAQYNSPAGLYSSENIQTFNSAVESKTSPGPPEPSKLRLSPMSAQQAPFASVYNPLQSP 134
Query: 59 CASTMFSYINLYGRYPPGLFASECREGKEGL--ACDALPPS 97
+ GR+ PG +S+ R G E L A A P S
Sbjct: 135 S-------VEARGRHSPGALSSQVRVGPEQLKHAAQACPKS 168
>UniRef100_Q9LT87 Similarity to serine/threonine kinase [Arabidopsis thaliana]
Length = 663
Score = 34.7 bits (78), Expect = 0.50
Identities = 23/76 (30%), Positives = 34/76 (44%), Gaps = 11/76 (14%)
Query: 6 IAPAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEF----ACPYAD------VINDL 55
+ AGC ++F N+T++ S C K CC F YA+ V +DL
Sbjct: 25 LTEAGCPLDFTSSNFTLVASVCSNNTERAK-CCRYMNAFVAISVARYANYTADLGVTSDL 83
Query: 56 TNDCASTMFSYINLYG 71
T C +T+ + LYG
Sbjct: 84 TEICITTISRTMELYG 99
>UniRef100_UPI0000361097 UPI0000361097 UniRef100 entry
Length = 1616
Score = 33.1 bits (74), Expect = 1.4
Identities = 14/38 (36%), Positives = 23/38 (59%)
Query: 80 SECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVL 117
S+C +G + L+CDA PP A N IV +++L++
Sbjct: 1340 SDCSDGSDELSCDADPPGHGAGSGPNAIVSVIAIILLV 1377
>UniRef100_UPI00003384D7 UPI00003384D7 UniRef100 entry
Length = 1060
Score = 33.1 bits (74), Expect = 1.4
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 6/69 (8%)
Query: 63 MFSYINLYGRYPPGLFASECREGKEGLACDALPPSVSADDT----ANQIVHTPSLVLVLT 118
++ + G PG+ + + E LA LPP V+ + T Q+ HTP+L +
Sbjct: 830 LYPAAEVQGIAAPGVASGVALKRMEELARQVLPPGVAFEWTDLAHQQQLPHTPTLAIFAA 889
Query: 119 A--CIFLIL 125
A C+FL+L
Sbjct: 890 AALCVFLVL 898
>UniRef100_Q7QGY2 ENSANGP00000012454 [Anopheles gambiae str. PEST]
Length = 234
Score = 32.3 bits (72), Expect = 2.5
Identities = 13/41 (31%), Positives = 21/41 (50%)
Query: 76 GLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLV 116
G + EC G +G C+ LPPS+ T++ PS ++
Sbjct: 152 GHYRCECPSGYKGRNCEILPPSMMTSTTSSSTTEQPSTTMM 192
>UniRef100_Q5X0G1 SdbB protein [Legionella pneumophila str. Lens]
Length = 415
Score = 31.6 bits (70), Expect = 4.2
Identities = 26/83 (31%), Positives = 36/83 (43%), Gaps = 5/83 (6%)
Query: 42 KEFACPYADVINDLTNDCASTMF-SYINL---YGRYPPGLFASECREGKEGLACDALPPS 97
K+F P+AD IN N+ +T F ++ NL R P L + + E E L C+ S
Sbjct: 48 KDFLAPFADFINQQKNNPEATYFNAFQNLELTLNRVPSQLVSGDWCE-LEVLKCEPEADS 106
Query: 98 VSADDTANQIVHTPSLVLVLTAC 120
T IV+ P AC
Sbjct: 107 PQKPGTGKHIVYFPGANTYYQAC 129
>UniRef100_Q23026 Hypothetical protein R09F10.3 [Caenorhabditis elegans]
Length = 468
Score = 31.2 bits (69), Expect = 5.5
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 60 ASTMFSYINLYGRYPPGLFASECREGKEGLA-CDALPPSVSADDTANQIVH 109
A T+F +I+LYG++ LF ++CR K+ L+ + LPP + N H
Sbjct: 415 AGTLFHFISLYGKFRE-LFTTKCRSCKQVLSRKNFLPPLIFDIRNPNNAAH 464
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 225,342,479
Number of Sequences: 2790947
Number of extensions: 8550812
Number of successful extensions: 20032
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 20010
Number of HSP's gapped (non-prelim): 22
length of query: 127
length of database: 848,049,833
effective HSP length: 103
effective length of query: 24
effective length of database: 560,582,292
effective search space: 13453975008
effective search space used: 13453975008
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144723.3


