

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
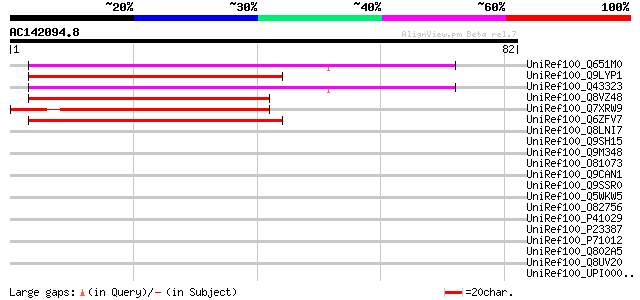
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q651M0 Putative membrane protein [Oryza sativa] 47 1e-04
UniRef100_Q9LYP1 Membrane protein [Arabidopsis thaliana] 47 1e-04
UniRef100_Q43323 Membrane protein [Saccharum hybrid cultivar H65... 45 5e-04
UniRef100_Q8VZ48 Putative membrane protein [Arabidopsis thaliana] 45 6e-04
UniRef100_Q7XRW9 OSJNBb0032E06.11 protein [Oryza sativa] 45 6e-04
UniRef100_Q6ZFV7 Putative membrane protein [Oryza sativa] 45 6e-04
UniRef100_Q8LNI7 Putative membrane protein [Oryza sativa] 44 0.001
UniRef100_Q9SH15 F28K19.7 [Arabidopsis thaliana] 44 0.001
UniRef100_Q9M348 Hypothetical protein F5K20_80 [Arabidopsis thal... 43 0.002
UniRef100_O81073 Hypothetical protein At2g29050 [Arabidopsis tha... 42 0.003
UniRef100_Q9CAN1 Membrane protein, putative; 61952-60281 [Arabid... 42 0.005
UniRef100_Q9SSR0 F6D8.20 [Arabidopsis thaliana] 40 0.015
UniRef100_Q5WKW5 PTS system, fructose specific enzyme II, C comp... 39 0.025
UniRef100_O82756 Putative membrane protein [Arabidopsis thaliana] 39 0.032
UniRef100_P41029 PTS system, fructose-specific IIBC component [B... 39 0.032
UniRef100_P23387 PTS system, fructose-specific IIBC component [R... 38 0.055
UniRef100_P71012 Phosphotransferase system (PTS) fructose-specif... 36 0.21
UniRef100_Q802A5 C16orf8-like protein [Fugu rubripes] 36 0.27
UniRef100_Q8UV20 C16ORF8 [Sphoeroides nephelus] 36 0.27
UniRef100_UPI000001838E UPI000001838E UniRef100 entry 35 0.61
>UniRef100_Q651M0 Putative membrane protein [Oryza sativa]
Length = 323
Score = 47.0 bits (110), Expect = 1e-04
Identities = 27/80 (33%), Positives = 42/80 (51%), Gaps = 11/80 (13%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVL-----------PD 52
+++ + NL IGI+P +NF IGG + GFLLGFVLL + + P
Sbjct: 209 ITLLFIIAINLAIGILPHADNFAHIGGFVTGFLLGFVLLARPQFGWMERHELPQTNQPPK 268
Query: 53 QKLHKRCLPIICFILLSTGW 72
K ++ L ++ F+LL G+
Sbjct: 269 YKAYQYVLWVVAFVLLLVGF 288
>UniRef100_Q9LYP1 Membrane protein [Arabidopsis thaliana]
Length = 346
Score = 47.0 bits (110), Expect = 1e-04
Identities = 21/41 (51%), Positives = 30/41 (72%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ V + NL IGI+P V+NF +GG + GFLLGF+LL +
Sbjct: 230 LTLLFVILINLAIGILPHVDNFAHVGGFVTGFLLGFILLAR 270
>UniRef100_Q43323 Membrane protein [Saccharum hybrid cultivar H65-7052]
Length = 325
Score = 45.1 bits (105), Expect = 5e-04
Identities = 27/80 (33%), Positives = 41/80 (50%), Gaps = 11/80 (13%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVL-----------PD 52
+++ + NL IGI+P V+NF IGG GFLLGFVLL + + P
Sbjct: 211 ITLLFIIALNLAIGILPHVDNFAHIGGFATGFLLGFVLLARPQFSWMERHELPQTNQPPK 270
Query: 53 QKLHKRCLPIICFILLSTGW 72
K ++ L ++ +LL G+
Sbjct: 271 YKAYQYILWVVALVLLLVGF 290
>UniRef100_Q8VZ48 Putative membrane protein [Arabidopsis thaliana]
Length = 307
Score = 44.7 bits (104), Expect = 6e-04
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
LS + NL IG++P V+NF IGGL+ GF LGF+LL
Sbjct: 193 LSFLFIIAINLAIGLLPWVDNFAHIGGLLTGFCLGFILL 231
>UniRef100_Q7XRW9 OSJNBb0032E06.11 protein [Oryza sativa]
Length = 342
Score = 44.7 bits (104), Expect = 6e-04
Identities = 24/42 (57%), Positives = 28/42 (66%), Gaps = 2/42 (4%)
Query: 1 MFNLSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
M NL I + NL +GI+P V+NF IGG GFLLGFVLL
Sbjct: 231 MVNLII--IAAINLALGILPRVDNFAHIGGFATGFLLGFVLL 270
>UniRef100_Q6ZFV7 Putative membrane protein [Oryza sativa]
Length = 323
Score = 44.7 bits (104), Expect = 6e-04
Identities = 20/41 (48%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ + NL IGI+P +NF IGG + GFLLGFVLL +
Sbjct: 209 ITLLFIIAINLAIGILPHADNFAHIGGFVTGFLLGFVLLAR 249
>UniRef100_Q8LNI7 Putative membrane protein [Oryza sativa]
Length = 329
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/37 (54%), Positives = 26/37 (70%)
Query: 8 MVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+V NL IGI+P V+NF IGG + GFLLGF+ L +
Sbjct: 217 IVIAINLAIGILPHVDNFAHIGGFLTGFLLGFIFLMR 253
>UniRef100_Q9SH15 F28K19.7 [Arabidopsis thaliana]
Length = 735
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/32 (62%), Positives = 23/32 (71%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
N IG +P ++NF IGG I GFLLGFVLL K
Sbjct: 241 NFLIGFLPFIDNFANIGGFISGFLLGFVLLFK 272
>UniRef100_Q9M348 Hypothetical protein F5K20_80 [Arabidopsis thaliana]
Length = 361
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ NL +G++P V+NF IGG GFLLGFVLL +
Sbjct: 202 VTLVLIVAVNLGLGVLPGVDNFAHIGGFATGFLLGFVLLIR 242
>UniRef100_O81073 Hypothetical protein At2g29050 [Arabidopsis thaliana]
Length = 372
Score = 42.4 bits (98), Expect = 0.003
Identities = 19/41 (46%), Positives = 27/41 (65%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ + NL +GI+P V+NF +GG GFLLGFV L +
Sbjct: 279 LTLIFIIAINLAVGILPHVDNFAHLGGFTSGFLLGFVFLIR 319
>UniRef100_Q9CAN1 Membrane protein, putative; 61952-60281 [Arabidopsis thaliana]
Length = 317
Score = 41.6 bits (96), Expect = 0.005
Identities = 18/41 (43%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ + NL +G++P V+NF IGG + GF LGFVLL +
Sbjct: 206 ITLLFIIAINLALGMLPRVDNFAHIGGFLTGFCLGFVLLVR 246
>UniRef100_Q9SSR0 F6D8.20 [Arabidopsis thaliana]
Length = 309
Score = 40.0 bits (92), Expect = 0.015
Identities = 17/41 (41%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ V NL++G +P V+N GG + GF LGFVLL +
Sbjct: 204 MTLILIIVLNLSVGFLPRVDNSAHFGGFLAGFFLGFVLLLR 244
>UniRef100_Q5WKW5 PTS system, fructose specific enzyme II, C component [Bacillus
clausii]
Length = 361
Score = 39.3 bits (90), Expect = 0.025
Identities = 24/75 (32%), Positives = 41/75 (54%), Gaps = 4/75 (5%)
Query: 4 LSIGMVPVF--NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLP 61
+SI P F L IG + G IGG++ GFL+G+V+L K +P + +P
Sbjct: 73 MSIADRPAFAPGLVIGFIANEIQAGFIGGILGGFLVGYVVLAIKKYVKVPKSMI--GLMP 130
Query: 62 IICFILLSTGWTGVL 76
++ L++T ++G+L
Sbjct: 131 VMIIPLIATAFSGLL 145
>UniRef100_O82756 Putative membrane protein [Arabidopsis thaliana]
Length = 313
Score = 38.9 bits (89), Expect = 0.032
Identities = 18/30 (60%), Positives = 22/30 (73%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
NL +G +P V+NF IGG GFLLGF+LL
Sbjct: 209 NLGLGTLPPVDNFAHIGGFFGGFLLGFLLL 238
>UniRef100_P41029 PTS system, fructose-specific IIBC component [Bacillus
amyloliquefaciens]
Length = 304
Score = 38.9 bits (89), Expect = 0.032
Identities = 24/60 (40%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Query: 17 GIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGWTGVL 76
G + N G +GGLI GFL G+V++ K FV Q L P++ + LL TGVL
Sbjct: 55 GFMATQANAGFLGGLIAGFLAGYVVILLKKLFVFIPQSL-DGLKPVLIYPLLGIFITGVL 113
>UniRef100_P23387 PTS system, fructose-specific IIBC component [Rhodobacter
capsulatus]
Length = 578
Score = 38.1 bits (87), Expect = 0.055
Identities = 23/70 (32%), Positives = 38/70 (53%), Gaps = 4/70 (5%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFI 66
G+ P L G++ + N G +GG++ GFL G+V +D LP + + P++
Sbjct: 313 GLTP--GLIGGMLAVNLNAGFLGGIVAGFLAGYVARWLRDAIKLP--RTLEGLKPVLILP 368
Query: 67 LLSTGWTGVL 76
LLST TG++
Sbjct: 369 LLSTAITGLI 378
>UniRef100_P71012 Phosphotransferase system (PTS) fructose-specific enzyme IIABC
component [Bacillus subtilis]
Length = 635
Score = 36.2 bits (82), Expect = 0.21
Identities = 22/60 (36%), Positives = 30/60 (49%), Gaps = 1/60 (1%)
Query: 17 GIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGWTGVL 76
G + N G +GGLI GFL G+V++ K F Q L P++ + L TGVL
Sbjct: 385 GFMATQANAGFLGGLIAGFLAGYVVILLKKVFTFIPQSL-DGLKPVLIYPLFGIFITGVL 443
>UniRef100_Q802A5 C16orf8-like protein [Fugu rubripes]
Length = 855
Score = 35.8 bits (81), Expect = 0.27
Identities = 20/69 (28%), Positives = 35/69 (49%), Gaps = 5/69 (7%)
Query: 9 VPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFV-LPDQKLHKRCLPIICFIL 67
+ F + G++P ++NFG I G + GF L F L P++ ++++ L I F+L
Sbjct: 754 ISTFLFSFGLLPWIDNFGHICGFVSGFFLSFTFL----PYISFGRSDMYRKRLQICVFLL 809
Query: 68 LSTGWTGVL 76
+ G L
Sbjct: 810 VFLGLLATL 818
>UniRef100_Q8UV20 C16ORF8 [Sphoeroides nephelus]
Length = 773
Score = 35.8 bits (81), Expect = 0.27
Identities = 21/64 (32%), Positives = 33/64 (50%), Gaps = 5/64 (7%)
Query: 9 VPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFV-LPDQKLHKRCLPIICFIL 67
V +F G++P ++NF I G I GF L F L P++ L+++ II F+L
Sbjct: 672 VVLFLFAFGLLPWIDNFAHISGFISGFFLSFAFL----PYISFGRMDLYRKRCQIIVFLL 727
Query: 68 LSTG 71
+ G
Sbjct: 728 VFVG 731
>UniRef100_UPI000001838E UPI000001838E UniRef100 entry
Length = 858
Score = 34.7 bits (78), Expect = 0.61
Identities = 20/64 (31%), Positives = 33/64 (51%), Gaps = 5/64 (7%)
Query: 9 VPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFV-LPDQKLHKRCLPIICFIL 67
V +F G++P ++NF I G I GF L F L P++ L+++ II F++
Sbjct: 757 VVLFLFAFGLLPWIDNFAHISGFISGFFLSFAFL----PYISFGRMDLYRKRCQIIVFLM 812
Query: 68 LSTG 71
+ G
Sbjct: 813 VFLG 816
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 150,132,113
Number of Sequences: 2790947
Number of extensions: 5952132
Number of successful extensions: 15648
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 15605
Number of HSP's gapped (non-prelim): 60
length of query: 82
length of database: 848,049,833
effective HSP length: 58
effective length of query: 24
effective length of database: 686,174,907
effective search space: 16468197768
effective search space used: 16468197768
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC142094.8


