

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.3 + phase: 0
(85 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
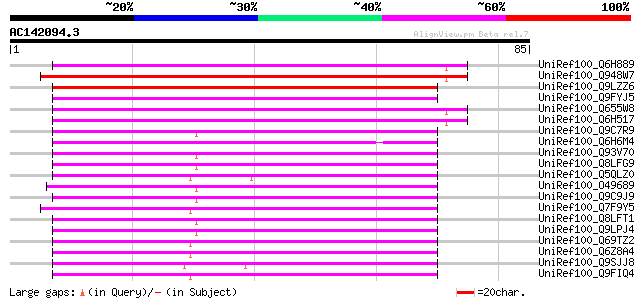
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6H889 Putative zinc-binding protein [Oryza sativa] 72 3e-12
UniRef100_Q948W7 Zinc-binding protein [Pisum sativum] 72 3e-12
UniRef100_Q9LZZ6 Hypothetical protein T4C21_80 [Arabidopsis thal... 72 4e-12
UniRef100_Q9FYJ5 F17F8.4 [Arabidopsis thaliana] 65 5e-10
UniRef100_Q655W8 Putative zinc-binding protein [Oryza sativa] 65 5e-10
UniRef100_Q6H517 Putative zinc-binding protein [Oryza sativa] 64 9e-10
UniRef100_Q9C7R9 Hypothetical protein F13A11.6 [Arabidopsis thal... 58 5e-08
UniRef100_Q6H6M4 Putative zinc-binding protein [Oryza sativa] 58 5e-08
UniRef100_Q93V70 At1g21000/F9H16_1 [Arabidopsis thaliana] 57 9e-08
UniRef100_Q8LFG9 Hypothetical protein [Arabidopsis thaliana] 57 9e-08
UniRef100_Q5QLZ0 Zinc-binding protein-like [Oryza sativa] 56 2e-07
UniRef100_O49689 Hypothetical protein AT4g17900 [Arabidopsis tha... 54 7e-07
UniRef100_Q9C9J9 Hypothetical protein F14G6.19 [Arabidopsis thal... 54 1e-06
UniRef100_Q7F9Y5 OSJNBa0086O06.23 protein [Oryza sativa] 54 1e-06
UniRef100_Q8LFT1 Hypothetical protein [Arabidopsis thaliana] 53 2e-06
UniRef100_Q9LPJ4 F6N18.8 [Arabidopsis thaliana] 52 4e-06
UniRef100_Q69TZ2 Putative zinc-binding protein [Oryza sativa] 52 4e-06
UniRef100_Q6Z8A4 Putative zinc-binding protein [Oryza sativa] 51 6e-06
UniRef100_Q9SJJ8 Hypothetical protein At2g27930 [Arabidopsis tha... 49 2e-05
UniRef100_Q9FIQ4 Arabidopsis thaliana genomic DNA, chromosome 5,... 47 9e-05
>UniRef100_Q6H889 Putative zinc-binding protein [Oryza sativa]
Length = 232
Score = 72.4 bits (176), Expect = 3e-12
Identities = 31/69 (44%), Positives = 41/69 (58%), Gaps = 1/69 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H + NEKN C+DC C HC AH +HR Q+ RY Y DV R +L+K DCS
Sbjct: 19 CSFHEHAKKNEKNICCLDCCTSICPHCVAAHRVHRLLQVRRYVYHDVVRLEDLEKLIDCS 78
Query: 68 NIQ-VTLNA 75
++Q T+N+
Sbjct: 79 SVQSYTINS 87
>UniRef100_Q948W7 Zinc-binding protein [Pisum sativum]
Length = 233
Score = 72.4 bits (176), Expect = 3e-12
Identities = 32/71 (45%), Positives = 44/71 (61%), Gaps = 1/71 (1%)
Query: 6 GYCDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFD 65
G C H++ R NEKN FC+ C + C HC +H+ H Q+ RY Y +V R +L+K D
Sbjct: 29 GDCGVHKNRRKNEKNIFCLHCCLSICPHCLSSHTSHPLLQVRRYVYHNVIRLDDLEKLID 88
Query: 66 CSNIQ-VTLNA 75
CSNIQ T+N+
Sbjct: 89 CSNIQPYTINS 99
>UniRef100_Q9LZZ6 Hypothetical protein T4C21_80 [Arabidopsis thaliana]
Length = 245
Score = 71.6 bits (174), Expect = 4e-12
Identities = 31/63 (49%), Positives = 38/63 (60%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C DH D + NEKN C+DC + C HC +H+ HR QI RY Y+DV R + K DCS
Sbjct: 22 CLDHEDDKKNEKNILCIDCCLTICPHCLSSHTSHRLLQIRRYVYRDVLRVEDGSKLMDCS 81
Query: 68 NIQ 70
IQ
Sbjct: 82 LIQ 84
>UniRef100_Q9FYJ5 F17F8.4 [Arabidopsis thaliana]
Length = 241
Score = 64.7 bits (156), Expect = 5e-10
Identities = 28/63 (44%), Positives = 36/63 (56%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H R +EKN FC+ C + C HC +H H Q+ RY Y DV R S+L+K DCS
Sbjct: 34 CGIHETRRKSEKNVFCLLCCLSVCPHCLPSHRSHPLLQVRRYVYHDVVRLSDLEKLIDCS 93
Query: 68 NIQ 70
+Q
Sbjct: 94 YVQ 96
>UniRef100_Q655W8 Putative zinc-binding protein [Oryza sativa]
Length = 261
Score = 64.7 bits (156), Expect = 5e-10
Identities = 29/69 (42%), Positives = 37/69 (53%), Gaps = 1/69 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H + NEKN FC+ C C HC +H H Q+ RY Y DV R +L K DCS
Sbjct: 20 CPAHESRKKNEKNIFCLGCCASICPHCAPSHRHHPLLQVRRYVYNDVVRLDDLDKLIDCS 79
Query: 68 NIQ-VTLNA 75
+Q T+N+
Sbjct: 80 FVQPYTINS 88
>UniRef100_Q6H517 Putative zinc-binding protein [Oryza sativa]
Length = 241
Score = 63.9 bits (154), Expect = 9e-10
Identities = 28/69 (40%), Positives = 38/69 (54%), Gaps = 1/69 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H + NEKN FC+ C C HC +H H Q+ RY Y DV R +L+K +CS
Sbjct: 20 CPAHESRKKNEKNIFCLACCTSICPHCAPSHRHHPLLQVRRYVYNDVVRLGDLEKLIECS 79
Query: 68 NIQ-VTLNA 75
+Q T+N+
Sbjct: 80 YVQPYTINS 88
>UniRef100_Q9C7R9 Hypothetical protein F13A11.6 [Arabidopsis thaliana]
Length = 216
Score = 58.2 bits (139), Expect = 5e-08
Identities = 30/64 (46%), Positives = 35/64 (53%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +E N FC+DC+ FC C H HR QI R SY +V R SE+QKH D
Sbjct: 24 CSIHSQSSKSECNLFCLDCSGNAFCSSCLAHHRTHRVIQIRRSSYHNVVRVSEIQKHIDI 83
Query: 67 SNIQ 70
S IQ
Sbjct: 84 SCIQ 87
>UniRef100_Q6H6M4 Putative zinc-binding protein [Oryza sativa]
Length = 226
Score = 58.2 bits (139), Expect = 5e-08
Identities = 29/63 (46%), Positives = 36/63 (57%), Gaps = 1/63 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H R N+KN FCVDCA CRHC + H QI++Y+ V R +L K FDC+
Sbjct: 25 CGAHPGERKNDKNHFCVDCAAALCRHCLPHDASHGVLQIWKYASCFVVRVDDL-KLFDCN 83
Query: 68 NIQ 70
IQ
Sbjct: 84 GIQ 86
>UniRef100_Q93V70 At1g21000/F9H16_1 [Arabidopsis thaliana]
Length = 246
Score = 57.4 bits (137), Expect = 9e-08
Identities = 29/64 (45%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D NE N FC+DCA FC +C H HR QI R SY +V R +E+QK D
Sbjct: 31 CSIHVDSNKNECNLFCLDCAGNAFCSYCLVKHKDHRVVQIRRSSYHNVVRVNEIQKFIDI 90
Query: 67 SNIQ 70
+ +Q
Sbjct: 91 ACVQ 94
>UniRef100_Q8LFG9 Hypothetical protein [Arabidopsis thaliana]
Length = 246
Score = 57.4 bits (137), Expect = 9e-08
Identities = 29/64 (45%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D NE N FC+DCA FC +C H HR QI R SY +V R +E+QK D
Sbjct: 31 CSIHVDSNKNECNLFCLDCAGNAFCSYCLVKHKDHRVVQIRRSSYHNVVRVNEIQKFIDI 90
Query: 67 SNIQ 70
+ +Q
Sbjct: 91 ACVQ 94
>UniRef100_Q5QLZ0 Zinc-binding protein-like [Oryza sativa]
Length = 236
Score = 56.2 bits (134), Expect = 2e-07
Identities = 29/67 (43%), Positives = 37/67 (54%), Gaps = 4/67 (5%)
Query: 8 CDDHRDLRSNEKNTFCVDCAV---RFCRHCKEAH-SIHRRFQIYRYSYQDVFRHSELQKH 63
C H + NE N FC+DC FC +C+ H S HR QI R SY DV + SEL+
Sbjct: 22 CTSHHNSPRNECNLFCIDCQAPEAAFCYYCRSCHHSSHRVIQIRRSSYHDVVKVSELEDI 81
Query: 64 FDCSNIQ 70
D S++Q
Sbjct: 82 LDISDVQ 88
>UniRef100_O49689 Hypothetical protein AT4g17900 [Arabidopsis thaliana]
Length = 254
Score = 54.3 bits (129), Expect = 7e-07
Identities = 27/65 (41%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Query: 7 YCDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFD 65
+C H D +E N +C+DC C C H HR QI R SY DV R +E+QK+ D
Sbjct: 48 HCKFHGDSHKSECNMYCLDCTNGPLCSLCLAHHKDHRTIQIRRSSYHDVIRVNEIQKYLD 107
Query: 66 CSNIQ 70
IQ
Sbjct: 108 IGGIQ 112
>UniRef100_Q9C9J9 Hypothetical protein F14G6.19 [Arabidopsis thaliana]
Length = 242
Score = 53.9 bits (128), Expect = 1e-06
Identities = 27/64 (42%), Positives = 36/64 (56%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +E N FC+DC+ FC +C H HR QI R SY +V R +E+QK+ D
Sbjct: 30 CSIHAASNKSECNMFCLDCSSEAFCSYCLLNHRNHRVLQIRRSSYHNVVRVNEIQKYIDI 89
Query: 67 SNIQ 70
S +Q
Sbjct: 90 SCVQ 93
>UniRef100_Q7F9Y5 OSJNBa0086O06.23 protein [Oryza sativa]
Length = 239
Score = 53.9 bits (128), Expect = 1e-06
Identities = 27/66 (40%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Query: 6 GYCDDHRDLRSNEKNTFCVDCAV-RFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHF 64
G C H D +E N +C+DC C C H H QI R SY DV R SE+QK
Sbjct: 37 GQCKLHADSHKSECNMYCLDCMNGALCSQCLSYHRDHHAIQIRRSSYHDVIRVSEIQKVL 96
Query: 65 DCSNIQ 70
D + +Q
Sbjct: 97 DITGVQ 102
>UniRef100_Q8LFT1 Hypothetical protein [Arabidopsis thaliana]
Length = 241
Score = 52.8 bits (125), Expect = 2e-06
Identities = 27/64 (42%), Positives = 35/64 (54%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +E N FC+DC+ FC +C H HR QI R SY +V R +E+QK D
Sbjct: 29 CSIHAASNKSECNMFCLDCSSEAFCSYCLLNHRNHRVLQIRRSSYHNVVRVNEIQKFIDI 88
Query: 67 SNIQ 70
S +Q
Sbjct: 89 SCVQ 92
>UniRef100_Q9LPJ4 F6N18.8 [Arabidopsis thaliana]
Length = 270
Score = 52.0 bits (123), Expect = 4e-06
Identities = 26/64 (40%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H H QI R SY DV R SE+QK D
Sbjct: 84 CKLHADSHKSECNMYCLDCTNGPLCSLCLSFHKDHHAIQIRRSSYHDVIRVSEIQKFLDI 143
Query: 67 SNIQ 70
+ +Q
Sbjct: 144 TGVQ 147
>UniRef100_Q69TZ2 Putative zinc-binding protein [Oryza sativa]
Length = 253
Score = 52.0 bits (123), Expect = 4e-06
Identities = 27/64 (42%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAV-RFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +L NE N FC+ C C +C AH H QI R SY +V R SE+ K D
Sbjct: 45 CASHPELSKNECNLFCLGCTGDALCAYCLPAHRDHHVVQIRRSSYHNVIRVSEVGKLIDI 104
Query: 67 SNIQ 70
S++Q
Sbjct: 105 SHVQ 108
>UniRef100_Q6Z8A4 Putative zinc-binding protein [Oryza sativa]
Length = 257
Score = 51.2 bits (121), Expect = 6e-06
Identities = 26/64 (40%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAV-RFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H H QI R SY DV R SE+QK D
Sbjct: 48 CRIHADAHKSECNMYCLDCMNGALCSLCLSHHRDHHAIQIRRSSYHDVIRVSEIQKVLDI 107
Query: 67 SNIQ 70
+ +Q
Sbjct: 108 TGVQ 111
>UniRef100_Q9SJJ8 Hypothetical protein At2g27930 [Arabidopsis thaliana]
Length = 135
Score = 49.3 bits (116), Expect = 2e-05
Identities = 26/65 (40%), Positives = 34/65 (52%), Gaps = 2/65 (3%)
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEA-HSIHRRFQIYRYSYQDVFRHSELQKHFD 65
C HR+ NE N FC+ C FC +C+ + H H QI R SY DV R SE++ D
Sbjct: 19 CPRHRETPRNECNMFCLSCQNAAFCFYCRSSFHIDHPVLQIRRSSYHDVVRVSEIENALD 78
Query: 66 CSNIQ 70
+Q
Sbjct: 79 IRGVQ 83
>UniRef100_Q9FIQ4 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MZA15
[Arabidopsis thaliana]
Length = 226
Score = 47.4 bits (111), Expect = 9e-05
Identities = 25/64 (39%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAV-RFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H L E +C+DC FC C H HR QI SY +V + E+QK+ D
Sbjct: 43 CKFHGHLPRTECKMYCLDCTNDSFCSLCLSEHENHRTIQIRISSYHNVTKVDEIQKYLDI 102
Query: 67 SNIQ 70
S+IQ
Sbjct: 103 SSIQ 106
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.329 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 132,136,441
Number of Sequences: 2790947
Number of extensions: 4337051
Number of successful extensions: 14964
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 14913
Number of HSP's gapped (non-prelim): 66
length of query: 85
length of database: 848,049,833
effective HSP length: 61
effective length of query: 24
effective length of database: 677,802,066
effective search space: 16267249584
effective search space used: 16267249584
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC142094.3


